Abstract
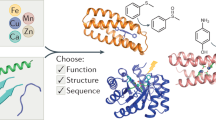
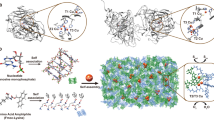
Protein design provides a rigorous test of our knowledge about proteins and allows the creation of novel enzymes for biotechnological applications. Whereas progress has been made in designing proteins that mimic native proteins structurally1,2,3, it is more difficult to design functional proteins4,5,6,7,8. In comparison to recent successes in designing non-metalloproteins4,6,7,9,10, it is even more challenging to rationally design metalloproteins that reproduce both the structure and function of native metalloenzymes5,8,11,12,13,14,15,16,17,18,19,20. This is because protein metal-binding sites are much more varied than non-metal-containing sites, in terms of different metal ion oxidation states, preferred geometry and metal ion ligand donor sets. Because of their variability, it has been difficult to predict metal-binding site properties in silico, as many of the parameters, such as force fields, are ill-defined. Therefore, the successful design of a structural and functional metalloprotein would greatly advance the field of protein design and our understanding of enzymes. Here we report a successful, rational design of a structural and functional model of a metalloprotein, nitric oxide reductase (NOR), by introducing three histidines and one glutamate, predicted as ligands in the active site of NOR, into the distal pocket of myoglobin. A crystal structure of the designed protein confirms that the minimized computer model contains a haem/non-haem FeB centre that is remarkably similar to that in the crystal structure. This designed protein also exhibits NO reduction activity, and so models both the structure and function of NOR, offering insight that the active site glutamate is required for both iron binding and activity. These results show that structural and functional metalloproteins can be rationally designed in silico.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Regan, L. & DeGrado, W. F. Characterization of a helical protein designed from first principles. Science 241, 976–978 (1988)
Hecht, M. H., Richardson, J. S., Richardson, D. C. & Ogden, R. C. De novo design, expression, and characterization of Felix: a four-helix bundle protein of native-like sequence. Science 249, 884–891 (1990)
Kuhlman, B. et al. Design of a novel globular protein fold with atomic-level accuracy. Science 302, 1364–1368 (2003)
Bolon, D. N. & Mayo, S. L. Enzyme-like proteins by computational design. Proc. Natl Acad. Sci. USA 98, 14274–14279 (2001)
Kaplan, J. & DeGrado, W. F. De novo design of catalytic proteins. Proc. Natl Acad. Sci. USA 101, 11566–11570 (2004)
Jiang, L. et al. De novo computational design of retro-aldol enzymes. Science 319, 1387–1391 (2008)
Röthlisberger, D. et al. Kemp elimination catalysts by computational enzyme design. Nature 453, 190–195 (2008)
Lu, Y., Yeung, N., Sieracki, N. & Marshall, N. M. Design of functional metalloproteins. Nature 460, 855–862 (2009)
Shifman, J. M. & Mayo, S. L. Exploring the origins of binding specificity through the computational redesign of calmodulin. Proc. Natl Acad. Sci. USA 100, 13274–13279 (2003)
Kang, S.-g. & Saven, J. G. Computational protein design: structure, function and combinatorial diversity. Curr. Opin. Chem. Biol. 11, 329–334 (2007)
Robertson, D. E. et al. Design and synthesis of multi-haem proteins. Nature 368, 425–432 (1994)
Hay, M., Richards, J. H. & Lu, Y. Construction and characterization of an azurin analog for the purple copper site in cytochrome c oxidase. Proc. Natl Acad. Sci. USA 93, 461–464 (1996)
Yeung, B. K., Wang, X., Sigman, J. A., Petillo, P. A. & Lu, Y. Construction and characterization of a manganese-binding site in cytochrome c peroxidase: towards a novel manganese peroxidase. Chem. Biol. 4, 215–221 (1997)
Sigman, J. A., Kwok, B. C. & Lu, Y. From myoglobin to heme-copper oxidase: design and engineering of a CuB center into sperm whale myoglobin. J. Am. Chem. Soc. 122, 8192–8196 (2000)
Case, M. A. & McLendon, G. L. Metal-assembled modular proteins: toward functional protein design. Acc. Chem. Res. 37, 754–762 (2004)
Cochran, F. V. et al. Computational de novo design and characterization of a four-helix bundle protein that selectively binds a nonbiological cofactor. J. Am. Chem. Soc. 127, 1346–1347 (2005)
Watanabe, Y. & Hayashi, T. Functionalization of myoglobin. Prog. Inorg. Chem. 54, 449–493 (2005)
Ghosh, D. & Pecoraro, V. L. Probing metal-protein interactions using a de novo design approach. Curr. Opin. Chem. Biol. 9, 97–103 (2005)
Petros, A. K., Reddi, A. R., Kennedy, M. L., Hyslop, A. G. & Gibney, B. R. Femtomolar Zn(II) affinity in a peptide-based ligand designed to model thiolate-rich metalloprotein active sites. Inorg. Chem. 45, 9941–9958 (2006)
Koder, R. L. et al. Design and engineering of an O2 transport protein. Nature 458, 305–309 (2009)
Wasser, I. M., de Vries, S., Moënne-Loccoz, P., Schröder, I. & Karlin, K. D. Nitric oxide in biological denitrification: Fe/Cu metalloenzyme and metal complex NOx redox chemistry. Chem. Rev. 102, 1201–1234 (2002)
van der Oost, J. et al. The heme-copper oxidase family consists of three distinct types of terminal oxidases and is related to nitric oxide reductase. FEMS Microbiol. Lett. 121, 1–9 (1994)
Girsch, P. & de Vries, S. Purification and initial kinetic and spectroscopic characterization of NO reductase from Paracoccus denitrificans . Biochim. Biophys. Acta 1318, 202–216 (1997)
Hendriks, J., Gohlke, U. & Saraste, M. From NO to OO: nitric oxide and dioxygen in bacterial respiration. J. Bioenerg. Biomembr. 30, 15–24 (1998)
Watmough, N. J. et al. Nitric oxide in bacteria: synthesis and consumption. Biochim. Biophys. Acta 1411, 456–474 (1999)
Wasser, I. M., Huang, H.-w., Moënne-Loccoz, P. & Karlin, K. D. Heme/non-heme diiron(II) complexes and O2, CO, and NO adducts as reduced and substrate-bound models for the active site of bacterial nitric oxide reductase. J. Am. Chem. Soc. 127, 3310–3320 (2005)
Collman, J. P. et al. A functional nitric oxide reductase model. Proc. Natl Acad. Sci. USA 105, 15660–15665 (2008)
Moënne-Loccoz, P. et al. Nitric oxide reductase from Paracoccus denitrificans contains an oxo-bridged heme/non-heme diiron center. J. Am. Chem. Soc. 122, 9344–9345 (2000)
Zumft, W. G. Nitric oxide reductases of prokaryotes with emphasis on the respiratory, heme-copper oxidase type. J. Inorg. Biochem. 99, 194–215 (2005)
Zhao, X., Yeung, N., Wang, Z., Guo, Z. & Lu, Y. Effects of metal ions in the CuB center on the redox properties of heme in heme-copper oxidases: spectroelectrochemical studies of an engineered heme-copper center in myoglobin. Biochemistry 44, 1210–1214 (2005)
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996)
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005)
Zhao, X., Yeung, N., Russell, B. S., Garner, D. K. & Lu, Y. Catalytic reduction of NO to N2O by a designed heme-copper center in myoglobin: implications for the role of metal ions. J. Am. Chem. Soc. 128, 6766–6767 (2006)
Taboy, C. H., Bonaventura, C. & Crumbliss, A. L. Anaerobic oxidations of myoglobin and hemoglobin by spectroelectrochemistry. Methods Enzymol. 353, 187–209 (2002)
Bonner, F. T. Nitric oxide gas. Methods Enzymol. 268, 50–57 (1996)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Sheldrick, G. M. & Schneider, T. R. SHELXL: high-resolution refinement. Methods Enzymol. 277, 319–343 (1997)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Giuffrè, A. et al. The heme-copper oxidases of Thermus thermophilus catalyze the reduction of nitric oxide: evolutionary implications. Proc. Natl Acad. Sci. USA 96, 14718–14723 (1999)
Acknowledgements
We thank M. J. Nilges for help with EPR analysis, S. L. Mullen and F. Sun for aiding in GC/MS data collection, E. Lee for help with computational modelling, N. M. Marshall for providing Azurin protein, J. R. Askim for help in FeBMb expression and purification, and T. Hayashi and P. Moënne-Loccoz for suggestions regarding N2O detection in solution. This work was supported by the US National Institutes of Health (GM062211).
Author Contributions N.Y. and Y.-W.L. performed most of the experimentation and wrote most of the manuscript. B.S.R. helped with the initial design of mutants and experiments. X.Z. and L.L. assisted in experimentation. K.D.M. performed computational modelling. Y.-G.G. guided crystallization and refined the crystal structure. H.R. collected crystal diffraction data. Y.L. designed, guided the project and edited the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains supplementary Methods and Notes, Supplementary Figures S1-S8 with Legends, Supplementary Table S1 and Supplementary References. (PDF 934 kb)
Rights and permissions
About this article
Cite this article
Yeung, N., Lin, YW., Gao, YG. et al. Rational design of a structural and functional nitric oxide reductase . Nature 462, 1079–1082 (2009). https://doi.org/10.1038/nature08620
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08620
This article is cited by
-
Overcoming universal restrictions on metal selectivity by protein design
Nature (2022)
-
A base measure of precision for protein stability predictors: structural sensitivity
BMC Bioinformatics (2021)
-
Proteins-Based Nanocatalysts for Energy Conversion Reactions
Topics in Current Chemistry (2020)
-
An efficient, step-economical strategy for the design of functional metalloproteins
Nature Chemistry (2019)
-
Reversing nitrogen fixation
Nature Reviews Chemistry (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.