Abstract
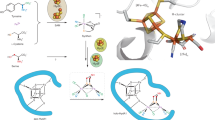
The trace element molybdenum is essential for nearly all organisms and forms the catalytic centre of a large variety of enzymes such as nitrogenase, nitrate reductases, sulphite oxidase and xanthine oxidoreductases. Nature has developed two scaffolds holding molybdenum in place, the iron–molybdenum cofactor and pterin-based molybdenum cofactors. Despite the different structures and functions of molybdenum-dependent enzymes, there are important similarities, which we highlight here. The biosynthetic pathways leading to both types of cofactor have common mechanistic aspects relating to scaffold formation, metal activation and cofactor insertion into apoenzymes, and have served as an evolutionary 'toolbox' to mediate additional cellular functions in eukaryotic metabolism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Dos Santos, P. C., Dean, D. R., Hu, Y. & Ribbe, M. W. Formation and insertion of the nitrogenase iron-molybdenum cofactor. Chem. Rev. 104, 1159–1173 (2004).
Schwarz, G. Molybdenum cofactor biosynthesis and deficiency. Cell. Mol. Life Sci. 62, 2792–2810 (2005).
Hille, R. Molybdenum and tungsten in biology. Trends Biochem. Sci. 27, 360–367 (2002).
Zhang, Y. & Gladyshev, V. N. Molybdoproteomes and evolution of molybdenum utilization. J. Mol. Biol. 379, 881–899 (2008).
Bevers, L. E., Hagedoorn, P.-L. & Hagen, W. R. The bioinorganic chemistry of tungsten. Coord. Chem. Rev. 253, 269–290 (2009).
Burgess, B. K. & Lowe, D. J. Mechanism of molybdenum nitrogenase. Chem. Rev. 96, 2983–3012 (1996).
Einsle, O. et al. Nitrogenase MoFe-protein at 1.16 Å resolution: a central ligand in the FeMo-cofactor. Science 297, 1696–1700 (2002).
Rajagopalan, K. V. & Johnson, J. L. The pterin molybdenum cofactors. J. Biol. Chem. 267, 10199–10202 (1992).
Eady, R. R. Structure–function relationships of alternative nitrogenases. Chem. Rev. 96, 3013–3030 (1996).
Schindelin, H. et al. Structure of ADP·AIF4 −-stabilized nitrogenase complex and its implications for signal transduction. Nature 387, 370–376 (1997).
Rothery, R. A., Workun, G. J. & Weiner, J. H. The prokaryotic complex iron-sulfur molybdoenzyme family. Biochim. Biophys. Acta 1778, 1897–1929 (2008). This paper is a systematic and comprehensive overview of prokaryotic molybdenum-containing enzymes of the DMSOR family.
Hille, R. Molybdenum-containing hydroxylases. Arch. Biochem. Biophys. 433, 107–116 (2005). This paper is a comprehensive overview of structure and reaction mechanisms of molybdenum-containing hydroxylases of the xanthine oxidase family.
Feng, C., Tollin, G. & Enemark, J. H. Sulfite oxidizing enzymes. Biochim. Biophys. Acta 1774, 527–539 (2007).
Moura, J. J., Brondino, C. D., Trincao, J. & Romao, M. J. Mo and W bis-MGD enzymes: nitrate reductases and formate dehydrogenases. J. Biol. Inorg. Chem. 9, 791–799 (2004).
Bertero, M. G. et al. Insights into the respiratory electron transfer pathway from the structure of nitrate reductase A. Nature Struct. Biol. 10, 681–687 (2003).
Kisker, C. et al. Molecular basis of sulfite oxidase deficiency from the structure of sulfite oxidase. Cell 91, 973–983 (1997).
Fischer, K. et al. Crystal structure of the yeast nitrate reductase molybdenum domain provides insight into eukaryotic nitrate assimilation. Plant Cell 17, 1167–1179 (2005).
Eilers, T. et al. Identification and biochemical characterization of Arabidopsis thaliana sulfite oxidase. A new player in plant sulfur metabolism. J. Biol. Chem. 276, 46989–46994 (2001).
Schrader, N. et al. The crystal structure of plant sulfite oxidase provides insights into sulfite oxidation in plants and animals. Structure 11, 1251–1263 (2003).
Kappler, U. & Bailey, S. Molecular basis of intramolecular electron transfer in sulfite-oxidizing enzymes is revealed by high resolution structure of a heterodimeric complex of the catalytic molybdopterin subunit and a c-type cytochrome subunit. J. Biol. Chem. 280, 24999–25007 (2005).
Hansch, R. et al. Plant sulfite oxidase as novel producer of H2O2: combination of enzyme catalysis with a subsequent non-enzymatic reaction step. J. Biol. Chem. 281, 6884–6888 (2006).
Campbell, W. H. Nitrate reductase structure, function and regulation: bridging the gap between biochemistry and physiology. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 277–303 (1999).
Mendel, R. R. & Bittner, F. Cell biology of molybdenum. Biochim. Biophys. Acta 1763, 621–635 (2006).
Enroth, C. et al. Crystal structures of bovine milk xanthine dehydrogenase and xanthine oxidase: structure-based mechanism of conversion. Proc. Natl Acad. Sci. USA 97, 10723–10728 (2000).
Truglio, J. J. et al. Crystal structures of the active and alloxanthine-inhibited forms of xanthine dehydrogenase from Rhodobacter capsulatus . Structure 10, 115–125 (2002).
Garattini, E. et al. Mammalian molybdo-flavoenzymes, an expanding family of proteins: structure, genetics, regulation, function and pathophysiology. Biochem. J. 372, 15–32 (2003).
Seo, M. et al. The Arabidopsis aldehyde oxidase 3 (AAO3) gene product catalyzes the final step in abscisic acid biosynthesis in leaves. Proc. Natl Acad. Sci. USA 97, 12908–12913 (2000).
Neumann, M. et al. A periplasmic aldehyde oxidoreductase represents the first molybdopterin cytosine dinucleotide cofactor containing molybdo-flavoenzyme from Escherichia coli. FEBS J. 276, 2762–2774 (2009).
Dobbek, H. et al. Catalysis at a dinuclear [CuSMo(=O)OH] cluster in a CO dehydrogenase resolved at 1.1-Å resolution. Proc. Natl Acad. Sci. USA 99, 15971–15976 (2002).
Harrison, R. Structure and function of xanthine oxidoreductase: where are we now? Free Radic. Biol. Med. 33, 774–797 (2002).
Havemeyer, A. et al. Identification of the missing component in the mitochondrial benzamidoxime prodrug-converting system as a novel molybdenum enzyme. J. Biol. Chem. 281, 34796–34802 (2006).
Groysman, S. & Holm, R. Biomimetic chemistry of iron, nickel, molybdenum, and tungsten in sulfur-ligated protein sites. Biochemistry 48, 2310–2320 (2009). This paper is a comprehensive review of the chemical synthesis of analogues of pterin cofactors of molybdenum enzymes and the FeMo-co and P-cluster of nitrogenase MoFe-protein.
Hu, Y. et al. Assembly of nitrogenase MoFe protein. Biochemistry 47, 3973–3981 (2008). This paper reports a recent development in the biosynthesis of the FeMo-co and P-cluster of nitrogenase MoFe-protein.
Smith, A. D. et al. NifS-mediated assembly of [4Fe-4S] clusters in the N- and C-terminal domains of the NifU scaffold protein. Biochemistry 44, 12955–12969 (2005).
Johnson, D. C., Dean, D. R., Smith, A. D. & Johnson, M. K. Structure, function, and formation of biological iron-sulfur clusters. Annu. Rev. Biochem. 74, 247–281 (2005).
Allen, R. M., Chatterjee, R., Ludden, P. W. & Shah, V. K. Incorporation of iron and sulfur from NifB cofactor into the iron-molybdenum cofactor of dinitrogenase. J. Biol. Chem. 270, 26890–26896 (1995).
Schmid, B. et al. Structure of a cofactor-deficient nitrogenase MoFe protein. Science 296, 352–356 (2002).
Curatti, L., Ludden, P. W. & Rubio, L. M. NifB-dependent in vitro synthesis of the iron–molybdenum cofactor of nitrogenase. Proc. Natl Acad. Sci. USA 103, 5297–5301 (2006).
Hu, Y., Fay, A. W. & Ribbe, M. W. Identification of a nitrogenase FeMo cofactor precursor on NifEN complex. Proc. Natl Acad. Sci. USA 102, 3236–3241 (2005).
Corbett, M. C. et al. Structural insights into a protein-bound iron-molybdenum cofactor precursor. Proc. Natl Acad. Sci. USA 103, 1238–1243 (2006). This paper is the first X-ray absorption spectroscopy/extended X-ray absorption fine structure (XAS/EXAFS)-based structural analysis of the FeMo-co precursor on NifEN.
Hu, Y. et al. FeMo cofactor maturation on NifEN. Proc. Natl Acad. Sci. USA 103, 17119–17124 (2006).
Zheng, L., White, R. H. & Dean, D. R. Purification of the Azotobacter vinelandii nifV-encoded homocitrate synthase. J. Bacteriol. 179, 5963–5966 (1997).
Imperial, J., Ugalde, R. A., Shah, V. K. & Brill, W. J. Role of the nifQ gene product in the incorporation of molybdenum into nitrogenase in Klebsiella pneumoniae . J. Bacteriol. 158, 187–194 (1984).
Hernandez, J. A. et al. Metal trafficking for nitrogen fixation: NifQ donates molybdenum to NifEN/NifH for the biosynthesis of the nitrogenase FeMo-cofactor. Proc. Natl Acad. Sci. USA 105, 11679–11684 (2008).
Hu, Y. et al. Nitrogenase Fe protein: a molybdate/homocitrate insertase. Proc. Natl Acad. Sci. USA 103, 17125–17130 (2006).
Georgiadis, M. M. et al. Crystallographic structure of the nitrogenase iron protein from Azotobacter vinelandii . Science 257, 1653–1659 (1992).
Rubio, L. M. et al. Cloning and mutational analysis of the γ gene from Azotobacter vinelandii defines a new family of proteins capable of metallocluster binding and protein stabilization. J. Biol. Chem. 277, 14299–14305 (2002).
Hu, Y. et al. Molecular insights into nitrogenase FeMoco insertion: TRP-444 of MoFe protein α-subunit locks FeMoco in its binding site. J. Biol. Chem. 281, 30534–30541 (2006).
Hu, Y., Fay, A. W. & Ribbe, M. W. Molecular insights into nitrogenase FeMo cofactor insertion: the role of His 362 of the MoFe protein α subunit in FeMo cofactor incorporation. J. Biol. Inorg. Chem. 12, 449–460 (2007).
Fay, A. W., Hu, Y., Schmid, B. & Ribbe, M. W. Molecular insights into nitrogenase FeMoco insertion — the role of His 274 and His 451 of MoFe protein α subunit. J. Inorg. Biochem. 101, 1630–1641 (2007).
Schwarz, G. & Mendel, R. R. Molybdenum cofactor biosynthesis and molybdenum enzymes. Annu. Rev. Plant Biol. 57, 623–647 (2006).
Schwarz, G., Hagedoorn, P. L. & Fischer, K. in Molecular Microbiology of Heavy Metals (eds Nies, D. H. & Silver, S.) 421–451 (Springer, 2007).
Reiss, J. & Johnson, J. L. Mutations in the molybdenum cofactor biosynthetic genes MOCS1, MOCS2, and GEPH . Hum. Mutat. 21, 569–576 (2003).
Wuebbens, M. M. & Rajagopalan, K. V. Structural characterization of a molybdopterin precursor. J. Biol. Chem. 268, 13493–13498 (1993).
Hanzelmann, P. & Schindelin, H. Binding of 5'-GTP to the C-terminal FeS cluster of the radical S-adenosylmethionine enzyme MoaA provides insights into its mechanism. Proc. Natl Acad. Sci. USA 103, 6829–6834 (2006). This paper describes the structural basis of radical SAM-based conversion of GTP into cPMP.
Wuebbens, M. M. & Rajagopalan, K. V. Investigation of the early steps of molybdopterin biosynthesis in Escherichia coli through the use of in vivo labeling studies. J. Biol. Chem. 270, 1082–1087 (1995).
Santamaria-Araujo, J. A. et al. The tetrahydropyranopterin structure of the sulfur-free and metal-free molybdenum cofactor precursor. J. Biol. Chem. 279, 15994–15999 (2004).
Daniels, J. N., Wuebbens, M. M., Rajagopalan, K. V. & Schindelin, H. Crystal structure of a molybdopterin synthase-precursor Z complex: insight into its sulfur transfer mechanism and its role in molybdenum cofactor deficiency. Biochemistry 47, 615–626 (2008).
Hänzelmann, P., Schwarz, G. & Mendel, R. R. Functionality of alternative splice forms of the first enzymes involved in human molybdenum cofactor biosynthesis. J. Biol. Chem. 277, 18303–18312 (2002).
Gutzke, G., Fischer, B., Mendel, R. R. & Schwarz, G. Thiocarboxylation of molybdopterin synthase provides evidence for the mechanism of dithiolene formation in metal-binding pterins. J. Biol. Chem. 276, 36268–36274 (2001).
Rudolph, M. J., Wuebbens, M. M., Rajagopalan, K. V. & Schindelin, H. Crystal structure of molybdopterin synthase and its evolutionary relationship to ubiquitin activation. Nature Struct. Biol. 8, 42–46 (2001).
Wuebbens, M. M. & Rajagopalan, K. V. Mechanistic and mutational studies of Escherichia coli molybdopterin synthase clarify the final step of molybdopterin biosynthesis. J. Biol. Chem. 278, 14523–14532 (2003).
Stallmeyer, B. et al. Human molybdopterin synthase gene: identification of a bicistronic transcript with overlapping reading frames. Am. J. Hum. Genet. 64, 698–705 (1999).
Lake, M. W., Wuebbens, M. M., Rajagopalan, K. V. & Schindelin, H. Mechanism of ubiquitin activation revealed by the structure of a bacterial MoeB–MoaD complex. Nature 414, 325–329 (2001).
Lee, I. & Schindelin, H. Structural insights into E1-catalyzed ubiquitin activation and transfer to conjugating enzymes. Cell 134, 268–278 (2008).
Leimkühler, S. & Rajagopalan, K. V. A sulfurtransferase is required in the transfer of cysteine sulfur in the in vitro synthesis of molybdopterin from precursor Z in Escherichia coli . J. Biol. Chem. 276, 22024–22031 (2001).
Forlani, F. et al. The cysteine-desulfurase IscS promotes the production of the rhodanese RhdA in the persulfurated form. FEBS Lett. 579, 6786–6790 (2005).
Matthies, A., Rajagopalan, K. V., Mendel, R. R. & Leimkuhler, S. Evidence for the physiological role of a rhodanese-like protein for the biosynthesis of the molybdenum cofactor in humans. Proc. Natl Acad. Sci. USA 101, 5946–5951 (2004).
Kuper, J. et al. Structure of the molybdopterin-bound Cnx1G domain links molybdenum and copper metabolism. Nature 430, 803–806 (2004). This paper reports the first protein-complex structures of Moco intermediates and the identification of a novel reaction intermediate essential for metal transfer.
Bevers, L. E. et al. Function of MoaB proteins in the biosynthesis of the molybdenum and tungsten cofactors. Biochemistry 47, 949–956 (2008).
Llamas, A., Mendel, R. R. & Schwarz, G. Synthesis of adenylated molybdopterin: an essential step for molybdenum insertion. J. Biol. Chem. 279, 55241–55246 (2004).
Llamas, A. et al. The mechanism of nucleotide-assisted molybdenum insertion into molybdopterin. A novel route toward metal cofactor assembly. J. Biol. Chem. 281, 18343–18350 (2006).
Bevers, L. E., Hagedoorn, P. L., Krijger, G. C. & Hagen, W. R. Tungsten transport protein A (WtpA) in Pyrococcus furiosus: the first member of a new class of tungstate and molybdate transporters. J. Bacteriol. 188, 6498–6505 (2006).
Stevenson, C. E. M. et al. Crystal structure of the molybdenum cofactor biosynthesis protein MobA from Escherichia coli at near-atomic resolution. Structure 8, 1115–1125 (2000).
Vergnes, A. et al. Involvement of the molybdenum cofactor biosynthetic machinery in the maturation of the Escherichia coli nitrate reductase A. J. Biol. Chem. 279, 41398–41403 (2004).
Bittner, F., Oreb, M. & Mendel, R. R. ABA3 is a molybdenum cofactor sulfurase required for activation of aldehyde oxidase and xanthine dehydrogenase in Arabidopsis thaliana . J. Biol. Chem. 276, 40381–40384 (2001).
Heidenreich, T., Wollers, S., Mendel, R. R. & Bittner, F. Characterization of the NifS-like domain of ABA3 from Arabidopsis thaliana provides insight into the mechanism of molybdenum cofactor sulfuration. J. Biol. Chem. 280, 4213–4218 (2005).
Wollers, S. et al. Binding of sulfurated molybdenum cofactor to the C-terminal domain of ABA3 from Arabidopsis thaliana provides insight into the mechanism of molybdenum cofactor sulfuration. J. Biol. Chem. 283, 9642–9650 (2008). This paper describes the mechanism and platform principle of Moco sulphuration in eukaryotes.
Neumann, M. et al. Rhodobacter capsulatus XdhC is involved in molybdenum cofactor binding and insertion into xanthine dehydrogenase. J. Biol. Chem. 281, 15701–15708 (2006).
Pau, R. N. & Lawson, D. M. Transport, homeostasis, regulation, and binding of molybdate and tungstate to proteins. Met. Ions Biol. Syst. 39, 31–74 (2002).
Ataya, F. S. et al. Mcp1 encodes the molybdenum cofactor carrier protein in Chlamydomonas reinhardtii and participates in protection, binding, and storage functions of the cofactor. J. Biol. Chem. 278, 10885–10890 (2003).
Fischer, K. et al. Function and structure of the molybdenum cofactor carrier protein from Chlamydomonas reinhardtii . J. Biol. Chem. 281, 30186–30194 (2006).
Sargent, F. Constructing the wonders of the bacterial world: biosynthesis of complex enzymes. Microbiology 153, 633–651 (2007).
Hollenstein, K., Frei, D. C. & Locher, K. P. Structure of an ABC transporter in complex with its binding protein. Nature 446, 213–216 (2007). This paper provides the first structural insight into the family of molybdate and tungstate ABC transporters
Hollenstein, K. et al. Distorted octahedral coordination of tungstate in a subfamily of specific binding proteins. J. Biol. Inorg. Chem. 14, 663–672 (2009).
Tomatsu, H. et al. An Arabidopsis thaliana high-affinity molybdate transporter required for efficient uptake of molybdate from soil. Proc. Natl Acad. Sci. USA 104, 18807–18812 (2007).
Tejada-Jimenez, M. et al. A high-affinity molybdate transporter in eukaryotes. Proc. Natl Acad. Sci. USA 104, 20126–20130 (2007).
Baxter, I. et al. Variation in molybdenum content across broadly distributed populations of Arabidopsis thaliana is controlled by a mitochondrial molybdenum transporter (MOT1). PLoS Genet. 4, e1000004 (2008).
Chen, S. et al. Functional characterization of AtATM1, AtATM2, and AtATM3, a subfamily of Arabidopsis half-molecule ATP-binding cassette transporters implicated in iron homeostasis. J. Biol. Chem. 282, 21561–21571 (2007).
Suttle, N. F. The interactions between copper, molybdenum, and sulphur in ruminant nutrition. Annu. Rev. Nutr. 11, 121–140 (1991).
Mason, J. Thiomolybdates: mediators of molybdenum toxicity and enzyme inhibitors. Toxicology 42, 99–109 (1986).
Mercer, J. F. The molecular basis of copper-transport diseases. Trends Mol. Med. 7, 64–69 (2001).
Brewer, G. J. Anticopper therapy against cancer and diseases of inflammation and fibrosis. Drug Discov. Today 10, 1103–1109 (2005).
Johnson, J. L. & Duran, M. in The Metabolic and Molecular Bases of Inherited Disease (eds Scriver, C. R., Beaudet, A. L., Sly, W. S. & Valle, D.) 3163–3177 (McGraw-Hill, 2001).
Zhang, X., Vincent, A. S., Halliwell, B. & Wong, K. P. A mechanism of sulfite neurotoxicity: direct inhibition of glutamate dehydrogenase. J. Biol. Chem. 279, 43035–43045 (2004).
Tan, W. H. et al. Isolated sulfite oxidase deficiency: a case report with a novel mutation and review of the literature. Pediatrics 116, 757–766 (2005).
Lee, H.-J. et al. Molybdenum cofactor-deficient mice resemble the phenotype of human patients. Hum. Mol. Genet. 11, 3309–3317 (2002).
Schwarz, G. et al. Rescue of lethal molybdenum cofactor deficiency by a biosynthetic precursor from Escherichia coli . Hum. Mol. Genet. 13, 1249–1255 (2004). This paper describes the successful therapeutic use of a Moco intermediate (cPMP) to cure Moco deficiency.
Stallmeyer, B. et al. The neurotransmitter receptor-anchoring protein gephyrin reconstitutes molybdenum cofactor biosynthesis in bacteria, plants, and mammalian cells. Proc. Natl Acad. Sci. USA 96, 1333–1338 (1999).
Reiss, J. et al. A mutation in the gene for the neurotransmitter receptor-clustering protein gephyrin causes a novel form of molybdenum cofactor deficiency. Am. J. Hum. Genet. 68, 208–213 (2001).
Acknowledgements
Support from the Deutsche Forschungsgemeinschaft (G.S., R.R.M.), the Bundeministerium für Bildung und Forschung (G.S.), the Fonds der Chemischen Industrie (G.S.), the European Union (R.R.M.) and the US National Institutes of Health (grant GM-67626; M.W.R.) is gratefully acknowledged, as is the contribution of all co-workers, especially graduate students and post-docs, during the past ten years.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reprints and permissions information is available at http://www.nature.com/reprints.
Correspondence should be addressed to G.S. (gschwarz@uni-koeln.de).
Rights and permissions
About this article
Cite this article
Schwarz, G., Mendel, R. & Ribbe, M. Molybdenum cofactors, enzymes and pathways. Nature 460, 839–847 (2009). https://doi.org/10.1038/nature08302
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08302
This article is cited by
-
Catalytic hydrogenation of olefins by a multifunctional molybdenum-sulfur complex
Nature Communications (2024)
-
A State-of-the-Science Review on Metal Biomarkers
Current Environmental Health Reports (2023)
-
A 3D-printed molybdenum-containing scaffold exerts dual pro-osteogenic and anti-osteoclastogenic effects to facilitate alveolar bone repair
International Journal of Oral Science (2022)
-
Nationwide genomic atlas of soil-dwelling Listeria reveals effects of selection and population ecology on pangenome evolution
Nature Microbiology (2021)
-
Molybdenum derived from nanomaterials incorporates into molybdenum enzymes and affects their activities in vivo
Nature Nanotechnology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.