Abstract
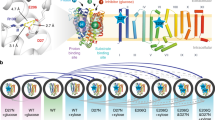
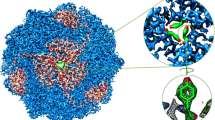
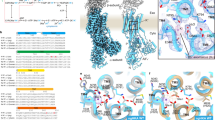
To reach the mammalian gut, enteric bacteria must pass through the stomach. Many such organisms survive exposure to the harsh gastric environment (pH 1.5–4) by mounting extreme acid-resistance responses, one of which, the arginine-dependent system of Escherichia coli, has been studied at levels of cellular physiology, molecular genetics and protein biochemistry1,2,3,4,5,6,7. This multiprotein system keeps the cytoplasm above pH 5 during acid challenge by continually pumping protons out of the cell using the free energy of arginine decarboxylation. At the heart of the process is a ‘virtual proton pump’8 in the inner membrane, called AdiC3,4, that imports l-arginine from the gastric juice and exports its decarboxylation product agmatine. AdiC belongs to the APC superfamily of membrane proteins6,7,9, which transports amino acids, polyamines and organic cations in a multitude of biological roles, including delivery of arginine for nitric oxide synthesis10, facilitation of insulin release from pancreatic β-cells11, and, when inappropriately overexpressed, provisioning of certain fast-growing neoplastic cells with amino acids12,13. High-resolution structures and detailed transport mechanisms of APC transporters are currently unknown. Here we describe a crystal structure of AdiC at 3.2 Å resolution. The protein is captured in an outward-open, substrate-free conformation with transmembrane architecture remarkably similar to that seen in four other families of apparently unrelated transport proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Audia, J. P., Webb, C. C. & Foster, J. W. Breaking through the acid barrier: an orchestrated response to proton stress by enteric bacteria. Int. J. Med. Microbiol. 291, 97–106 (2001)
Iyer, R. et al. A biological role for prokaryotic ClC chloride channels. Nature 419, 715–718 (2002)
Iyer, R., Williams, C. & Miller, C. Arginine-agmatine antiporter in extreme acid resistance in Escherichia coli . J. Bacteriol. 185, 6556–6561 (2003)
Gong, S., Richard, H. & Foster, J. W. YjdE (AdiC) is the arginine:agmatine antiporter essential for arginine-dependent acid resistance in Escherichia coli . J. Bacteriol. 185, 4402–4409 (2003)
Foster, J. W. Escherichia coli acid resistance: tales of an amateur acidophile. Nature Rev. Microbiol. 2, 898–907 (2004)
Fang, Y., Kolmakova-Partensky, L. & Miller, C. A bacterial arginine-agmatine exchange transporter involved in extreme acid resistance. J. Biol. Chem. 282, 176–182 (2007)
Casagrande, F. et al. Projection structure of a member of the amino acid/polyamine/organocation transporter superfamily. J. Biol. Chem. 283, 33240–33248 (2008)
Maloney, P. C. Bacterial transporters. Curr. Opin. Cell Biol. 6, 571–582 (1994)
Jack, D. L., Paulsen, I. T. & Saier, M. H. The amino acid/polyamine/organocation (APC) superfamily of transporters specific for amino acids, polyamines and organocations. Microbiology 146, 1797–1814 (2000)
Nicholson, B. et al. Sustained nitric oxide production in macrophages requires the arginine transporter CAT2. J. Biol. Chem. 276, 15881–15885 (2001)
Smith, P. A. et al. Electrogenic arginine transport mediates stimulus-secretion coupling in mouse pancreatic beta-cells. J. Physiol. (Lond.) 499, 625–635 (1997)
Fuchs, B. C. & Bode, B. P. Amino acid transporters ASCT2 and LAT1 in cancer: partners in crime? Semin. Cancer Biol. 15, 254–266 (2005)
Kaira, K. et al. l-type amino acid transporter 1 and CD98 expression in primary and metastatic sites of human neoplasms. Cancer Sci. 99, 2380–2386 (2008)
Kashiwagi, K. et al. Identification of the putrescine recognition site on polyamine transport protein PotE. J. Biol. Chem. 275, 36007–36012 (2000)
Kashiwagi, K. et al. Excretion and uptake of putrescine by the PotE protein in Escherichia coli . J. Biol. Chem. 272, 6318–6323 (1997)
Hu, L. A. & King, S. C. Membrane topology of the Escherichia coli γ-aminobutyrate transporter: implications on the topography and mechanism of prokaryotic and eukaryotic transporters from the APC superfamily. Biochem. J. 336, 69–76 (1998)
Fu, D. et al. Structure of a glycerol-conducting channel and the basis for its selectivity. Science 290, 481–486 (2000)
Dutzler, R. et al. X-ray structure of a ClC chloride channel at 3.0 Å reveals the molecular basis of anion selectivity. Nature 415, 287–294 (2002)
Yamashita, A. et al. Crystal structure of a bacterial homologue of Na+/Cl–-dependent neurotransmitter transporters. Nature 437, 215–223 (2005)
Faham, S. et al. The crystal structure of a sodium galactose transporter reveals mechanistic insights into Na+/sugar symport. Science 321, 810–814 (2008)
Weyand, S. et al. Structure and molecular mechanism of a nucleobase-cation-symport-1 family transporter. Science 322, 709–713 (2008)
Ressl, S. et al. Molecular basis of transport and regulation in the Na+/betaine symporter BetP. Nature 458, 47–52 (2009)
Krishnamurthy, H., Piscitelli, C. L. & Gouaux, E. Unlocking the molecular secrets of sodium-coupled transporters. Nature 459, 347–355 (2009)
Lolkema, J. & Slotboom, D.-J. The major amino acid transporter superfamily has a similar core structure as Na+-galactose and Na+-leucine transporters. Mol. Membr. Biol. 25, 567–570 (2008)
Soksawatmaekhin, W. et al. Identification of the cadaverine recognition site on the cadaverine-lysine antiporter CadB. J. Biol. Chem. 281, 29213–29220 (2006)
Guan, L. & Kaback, H. R. Lessons from lactose permease. Annu. Rev. Biophys. Biomol. Struct. 35, 67–91 (2006)
Burley, S. K. & Petsko, G. A. Amino-aromatic interactions in proteins. FEBS Lett. 203, 139–143 (1986)
Zacharias, N. & Dougherty, D. A. Cation-π interactions in ligand recognition and catalysis. Trends Pharmacol. Sci. 23, 281–287 (2002)
Gao, X. et al. Structure and mechanism of an amino acid antiporter. Science 324, 1565–1568 (2009)
Read, R. Improved Fourier coefficients for maps using phases from partial structures with errors. Acta Crystallogr. A 42, 140–149 (1986)
Cowtan, K. D. & Main, P. Phase combination and cross validation in iterated density-modification calculations. Acta Crystallogr. D 52, 43–48 (1996)
Acknowledgements
Y.F. was supported by NIH grant T32 NS 07292. We appreciate the support of the beamline scientists at the Advanced Photon Source (GM-CAT, 23-ID), Advanced Light Source (8.2.1, 8.2.2) and National Synchrotron Light Source (X-25, X-29A). We are also grateful to J. Berry for help in hybridoma sequencing, B. Bowman for crystallographic advice, E. Gouaux for sharing information about an APC homologue, and P. DeWeer, D. P. Krummel, H. H. Lim, K. Piasta and J. Robertson for suggestions on the manuscript.
Author Contributions Experiments were carried out and diffraction data collected by Y.F., H.J., T.S., L.K.-P., F.W., C.W. and C.M. Data were analysed by Y.F., H.J., Y.X. and C.M. The manuscript was prepared by Y.F., H.J., Y.X. and C.M.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Table S1 and Supplementary Figures S1-S3 with Legends. (PDF 481 kb)
Rights and permissions
About this article
Cite this article
Fang, Y., Jayaram, H., Shane, T. et al. Structure of a prokaryotic virtual proton pump at 3.2 Å resolution. Nature 460, 1040–1043 (2009). https://doi.org/10.1038/nature08201
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08201
This article is cited by
-
Cryo-EM structure of the human Asc-1 transporter complex
Nature Communications (2024)
-
A combined ligand and target-based virtual screening strategy to repurpose drugs as putrescine uptake inhibitors with trypanocidal activity
Journal of Computer-Aided Molecular Design (2023)
-
Heteromeric Amino Acid Transporters in Brain: from Physiology to Pathology
Neurochemical Research (2022)
-
High-resolution structure of the amino acid transporter AdiC reveals insights into the role of water molecules and networks in oligomerization and substrate binding
BMC Biology (2021)
-
Asc-1 Transporter (SLC7A10): Homology Models And Molecular Dynamics Insights Into The First Steps Of The Transport Mechanism
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.