Abstract
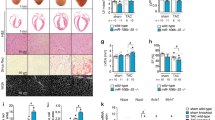
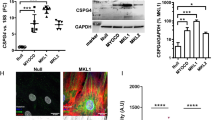
MicroRNAs (miRNAs) are regulators of myriad cellular events, but evidence for a single miRNA that can efficiently differentiate multipotent stem cells into a specific lineage or regulate direct reprogramming of cells into an alternative cell fate has been elusive. Here we show that miR-145 and miR-143 are co-transcribed in multipotent murine cardiac progenitors before becoming localized to smooth muscle cells, including neural crest stem-cell-derived vascular smooth muscle cells. miR-145 and miR-143 were direct transcriptional targets of serum response factor, myocardin and Nkx2-5 (NK2 transcription factor related, locus 5) and were downregulated in injured or atherosclerotic vessels containing proliferating, less differentiated smooth muscle cells. miR-145 was necessary for myocardin-induced reprogramming of adult fibroblasts into smooth muscle cells and sufficient to induce differentiation of multipotent neural crest stem cells into vascular smooth muscle. Furthermore, miR-145 and miR-143 cooperatively targeted a network of transcription factors, including Klf4 (Kruppel-like factor 4), myocardin and Elk-1 (ELK1, member of ETS oncogene family), to promote differentiation and repress proliferation of smooth muscle cells. These findings demonstrate that miR-145 can direct the smooth muscle fate and that miR-145 and miR-143 function to regulate the quiescent versus proliferative phenotype of smooth muscle cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kloosterman, W. P. & Plasterk, R. H. The diverse functions of microRNAs in animal development and disease. Dev. Cell 11, 441–450 (2006)
Calin, G. A. & Croce, C. M. MicroRNA signatures in human cancers. Nature Rev. Cancer 6, 857–866 (2006)
Zhao, Y. & Srivastava, D. A developmental view of microRNA function. Trends Biochem. Sci. 32, 189–197 (2007)
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009)
Rajewsky, N. MicroRNA target predictions in animals. Nature Genet. 38 (Suppl). S8–S13 (2006)
Vasudevan, S., Tong, Y. & Steitz, J. A. Switching from repression to activation: microRNAs can up-regulate translation. Science 318, 1931–1934 (2007)
Zhao, Y., Samal, E. & Srivastava, D. Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis. Nature 436, 214–220 (2005)
Kwon, C., Han, Z., Olson, E. N. & Srivastava, D. MicroRNA1 influences cardiac differentiation in Drosophila and regulates Notch signaling. Proc. Natl Acad. Sci. USA 102, 18986–18991 (2005)
Chen, J. F. et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nature Genet. 38, 228–233 (2006)
Zhao, Y. et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1–2. Cell 129, 303–317 (2007)
Ivey, K. N. et al. MicroRNA regulation of cell lineages in mouse and human embryonic stem cells. Cell Stem Cell 2, 219–229 (2008)
Srivastava, D. Making or breaking the heart: from lineage determination to morphogenesis. Cell 126, 1037–1048 (2006)
Kattman, S. J., Huber, T. L. & Keller, G. M. Multipotent flk-1+ cardiovascular progenitor cells give rise to the cardiomyocyte, endothelial, and vascular smooth muscle lineages. Dev. Cell 11, 723–732 (2006)
Le Douarin, N. M., Creuzet, S., Couly, G. & Dupin, E. Neural crest cell plasticity and its limits. Development 131, 4637–4650 (2004)
Ross, R. The pathogenesis of atherosclerosis: A perspective for the 1990s. Nature 362, 801–809 (1993)
Owens, G. K., Kumar, M. S. & Wamhoff, B. R. Molecular regulation of vascular smooth muscle cell differentiation in development and disease. Physiol. Rev. 84, 767–801 (2004)
Yoshida, T. & Owens, G. K. Molecular determinants of vascular smooth muscle cell diversity. Circ. Res. 96, 280–291 (2005)
Wang, Y., Medvid, R., Melton, C., Jaenisch, R. & Blelloch, R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nature Genet. 39, 380–385 (2007)
Cai, C. L. et al. Isl1 identifies a cardiac progenitor population that proliferates prior to differentiation and contributes a majority of cells to the heart. Dev. Cell 5, 877–889 (2003)
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001)
Moretti, A. et al. Multipotent embryonic isl1+ progenitor cells lead to cardiac, smooth muscle, and endothelial cell diversification. Cell 127, 1151–1165 (2006)
Wang, Z. et al. Myocardin and ternary complex factors compete for SRF to control smooth muscle gene expression. Nature 428, 185–189 (2004)
Wang, D. et al. Activation of cardiac gene expression by myocardin, a transcriptional cofactor for serum response factor. Cell 105, 851–862 (2001)
Chen, J., Kitchen, C. M., Streb, J. W. & Miano, J. M. Myocardin: a component of a molecular switch for smooth muscle differentiation. J. Mol. Cell. Cardiol. 34, 1345–1356 (2002)
Long, X., Bell, R. D., Gerthoffer, W. T., Zlokovic, B. V. & Miano, J. M. Myocardin is sufficient for a smooth muscle-like contractile phenotype. Arterioscler. Thromb. Vasc. Biol. 28, 1505–1510 (2008)
Chen, C. Y. & Schwartz, R. J. Recruitment of the tinman homolog Nkx-2.5 by serum response factor activates cardiac alpha-actin gene transcription. Mol. Cell. Biol. 16, 6372–6384 (1996)
Ji, R. et al. MicroRNA expression signature and antisense-mediated depletion reveal an essential role of MicroRNA in vascular neointimal lesion formation. Circ. Res. 100, 1579–1588 (2007)
Krutzfeldt, J. et al. Silencing of microRNAs in vivo with 'antagomirs'. Nature 438, 685–689 (2005)
Maurer, J. et al. Establishment and controlled differentiation of neural crest stem cell lines using conditional transgenesis. Differentiation 75, 580–591 (2007)
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007)
Liu, Y. et al. Kruppel-like factor 4 abrogates myocardin-induced activation of smooth muscle gene expression. J. Biol. Chem. 280, 9719–9727 (2005)
House, S. J. & Singer, H. A. CaMKII-delta isoform regulation of neointima formation after vascular injury. Arterioscler. Thromb. Vasc. Biol. 28, 441–447 (2008)
Mishra-Gorur, K., Singer, H. A. & Castellot, J. J. Heparin inhibits phosphorylation and autonomous activity of Ca2+/calmodulin-dependent protein kinase II in vascular smooth muscle cells. Am. J. Pathol. 161, 1893–1901 (2002)
Xu, N., Papagiannakopoulos, T., Pan, G., Thomson, J. A. & Kosik, K. S. MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 137, 647–658 (2009)
Yamagishi, H. et al. Tbx1 is regulated by tissue-specific forkhead proteins through a common Sonic hedgehog-responsive enhancer. Genes Dev. 17, 269–281 (2003)
Wang, Z., Wang, D. Z., Pipes, G. C. & Olson, E. N. Myocardin is a master regulator of smooth muscle gene expression. Proc. Natl Acad. Sci. USA 100, 7129–7134 (2003)
Yamamoto, M. et al. The roles of protein kinase C beta I and beta II in vascular smooth muscle cell proliferation. Exp. Cell Res. 240, 349–358 (1998)
Sinha, S. et al. Assessment of contractility of purified smooth muscle cells derived from embryonic stem cells. Stem Cells 24, 1678–1688 (2006)
Obernosterer, G., Martinez, J. & Alenius, M. Locked nucleic acid-based in situ detection of microRNAs in mouse tissue sections. Nature Protocols 2, 1508–1514 (2007)
Kruger, J. & Rehmsmeier, M. RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic Acids Res. 34, W451–W454 (2006)
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003)
Regan, C. P., Adam, P. J., Madsen, C. S. & Owens, G. K. Molecular mechanisms of decreased smooth muscle differentiation marker expression after vascular injury. J. Clin. Invest. 106, 1139–1147 (2000)
Acknowledgements
We thank R. Blelloch for DGCR8-null EBs; R.J. Schwartz for SRF-null ES cells; I. Charo and N. Saederup for RNA from atherosclerotic tissue; J. Maurer for JoMa neural crest cell line; L. Qian and Y. Huang for providing mouse cardiac infarct RNA; C. Tsou for help with calcium flux assays; E. N. Olson for the myocardin expression plasmid; P. Swinton for generation of transgenic mice; J. Fish and C. Miller for histopathology support; S. Ordway and G. Howard for scientific editing; B. Taylor for manuscript preparation. We also thank members of the Srivastava laboratory for discussions. J.M.M. was supported by HL62572 and HL091168 from NHLBI/NIH. D.S. was supported by grants from the NHLBI/NIH and the California Institute for Regenerative Medicine (CIRM) and was an Established Investigator of the American Heart Association. This work was also supported by NIH/NCRR grant C06 RR018928 to the Gladstone Institutes.
Author Contributions K.R.C. and D.S. designed the study and K.R.C. executed or oversaw execution of all experiments; N.T.S. and E.C.B. performed the NCC studies; M.P.W. and K.N.I. performed some expression and stem cell studies and K.N.I. helped supervise the project; A.N.M. provided technical support; T.-H.L. and J.M.M. performed carotid artery ligation studies; S.U.M. isolated YFP+ progenitor cells and performed some expression studies; J.M.M. assisted K.R.C and D.S. in editing the manuscript; K.R.C. and D.S. wrote the manuscript and D.S. supervised all aspects of the project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
D.S. serves on the Scientific Advisory Board of iZumi Bio.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-7 with Legends. Supplementary Fig. 3b was corrected on 18 August 2009. (PDF 1337 kb)
Rights and permissions
About this article
Cite this article
Cordes, K., Sheehy, N., White, M. et al. miR-145 and miR-143 regulate smooth muscle cell fate and plasticity. Nature 460, 705–710 (2009). https://doi.org/10.1038/nature08195
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08195
This article is cited by
-
A preliminary study of the miRNA restitution effect on CNV-induced miRNA downregulation in CAKUT
BMC Genomics (2024)
-
Dysregulation of micro-RNA 143-3p as a Biomarker of Carotid Atherosclerosis and the Associated Immune Reactions During Disease Progression
Journal of Cardiovascular Translational Research (2024)
-
Multi-omics in thoracic aortic aneurysm: the complex road to the simplification
Cell & Bioscience (2023)
-
Vascular smooth muscle cells enhance immune/vascular interplay in a 3-cell model of vascular inflammation
Scientific Reports (2023)
-
Flow-induced reprogramming of endothelial cells in atherosclerosis
Nature Reviews Cardiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.