Abstract
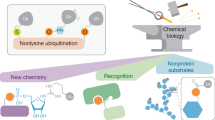
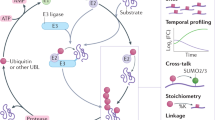
Eukaryotic proteins can be modified through attachment to various small molecules and proteins. One such modification is conjugation to ubiquitin and ubiquitin-like proteins (UBLs), which controls an enormous range of physiological processes. Bound UBLs mainly regulate the interactions of proteins with other macromolecules, for example binding to the proteasome or recruitment to chromatin. The various UBL systems use related enzymes to attach specific UBLs to proteins (or other molecules), and most of these attachments are transient. There is increasing evidence suggesting that such UBL–protein modification evolved from prokaryotic sulphurtransferase systems or related enzymes. Moreover, proteins similar to UBL-conjugating enzymes and UBL-deconjugating enzymes seem to have already been widespread at the time of the last common ancestor of eukaryotes, suggesting that UBL–protein conjugation did not first evolve in eukaryotes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hochstrasser, M. Evolution and function of ubiquitin-like protein-conjugation systems. Nature Cell Biol. 2, E153–E157 (2000).
Pickart, C. M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001).
Xu, P. & Peng, J. Dissecting the ubiquitin pathway by mass spectrometry. Biochim. Biophys. Acta 1764, 1940–1947 (2006).
Kerscher, O., Felberbaum, R. & Hochstrasser, M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu. Rev. Cell Dev. Biol. 22, 159–180 (2006).
Sharp, P. M. & Li, W.-H. Molecular evolution of ubiquitin genes. Trends Ecol. Evol. 2, 328–332 (1987).
Iyer, L. M., Burroughs, A. M. & Aravind, L. The prokaryotic antecedents of the ubiquitin-signaling system and the early evolution of ubiquitin-like β-grasp domains. Genome Biol. 7, R60 (2006). Highly sensitive sequence comparisons reveal a plethora of prokaryotic UBL/β-grasp proteins and potential UBL-modification pathways.
Begley, T. P. Cofactor biosynthesis: an organic chemist's treasure trove. Nat. Prod. Rep. 23, 15–25 (2006).
Mukhopadhyay, D. & Riezman, H. Proteasome-independent functions of ubiquitin in endocytosis and signaling. Science 315, 201–205 (2007).
Schwartz, A. L. & Ciechanover, A. Targeting proteins for destruction by the ubiquitin system: implications for human pathobiology. Annu. Rev. Pharmacol. Toxicol. 49, 73–96 (2009).
Volker, C. & Lupas, A. N. Molecular evolution of proteasomes. Curr. Top. Microbiol. Immunol. 268, 1–22 (2002).
Cavalier-Smith, T. Rooting the tree of life by transition analyses. Biol. Direct 1, 19 (2006).
Haas, A. L., Ahrens, P., Bright, P. M. & Ankel, H. Interferon induces a 15-kilodalton protein exhibiting marked homology to ubiquitin. J. Biol. Chem. 262, 11315–11323 (1987).
Loeb, K. R. & Haas, A. L. The interferon-inducible 15-kDa ubiquitin homolog conjugates to intracellular proteins. J. Biol. Chem. 267, 7806–7813 (1992).
Yuan, W. & Krug, R. M. Influenza B virus NS1 protein inhibits conjugation of the interferon (IFN)-induced ubiquitin-like ISG15 protein. EMBO J. 20, 362–371 (2001).
Zhao, C. et al. The UbcH8 ubiquitin E2 enzyme is also the E2 enzyme for ISG15, an IFN-α/β-induced ubiquitin-like protein. Proc. Natl Acad. Sci. USA 101, 7578–7582 (2004).
Kim, K. I., Giannakopoulos, N. V., Virgin, H. W. & Zhang, D. E. Interferon-inducible ubiquitin E2, Ubc8, is a conjugating enzyme for protein ISGylation. Mol. Cell. Biol. 24, 9592–9600 (2004).
Durfee, L. A., Kelley, M. L. & Huibregtse, J. M. The basis for selective E1–E2 interactions in the ISG15 conjugation system. J. Biol. Chem. 283, 23895–23902 (2008).
Lenschow, D. J. et al. IFN-stimulated gene 15 functions as a critical antiviral molecule against influenza, herpes, and Sindbis viruses. Proc. Natl Acad. Sci. USA 104, 1371–1376 (2007).
Okumura, A., Pitha, P. M. & Harty, R. N. ISG15 inhibits Ebola VP40 VLP budding in an L-domain-dependent manner by blocking Nedd4 ligase activity. Proc. Natl Acad. Sci. USA 105, 3974–3979 (2008).
Malakhova, O. A. & Zhang, D. E. ISG15 inhibits Nedd4 ubiquitin E3 activity and enhances the innate antiviral response. J. Biol. Chem. 283, 8783–8787 (2008).
Peng, J. et al. A proteomics approach to understanding protein ubiquitination. Nature Biotechnol. 21, 921–926 (2003).
Ohsumi, Y. Molecular dissection of autophagy: two ubiquitin-like systems. Nature Rev. Mol. Cell Biol. 2, 211–216 (2001).
Liu, Y., Fallon, L., Lashuel, H. A., Liu, Z. & Lansbury, P. T. Jr . The UCH-L1 gene encodes two opposing enzymatic activities that affect α-synuclein degradation and Parkinson's disease susceptibility. Cell 111, 209–218 (2002).
Dassa, B., Yanai, I. & Pietrokovski, S. New type of polyubiquitin-like genes with intein-like autoprocessing domains. Trends Genet. 20, 538–542 (2004).
Huang, T. T. et al. Regulation of monoubiquitinated PCNA by DUB autocleavage. Nature Cell Biol. 8, 339–347 (2006).
Burroughs, A. M., Balaji, S., Iyer, L. M. & Aravind, L. Small but versatile: the extraordinary functional and structural diversity of the β-grasp fold. Biol. Direct 2, 18 (2007).
Hershko, A. & Ciechanover, A. The ubiquitin system for protein degradation. Annu. Rev. Biochem. 61, 761–807 (1992).
Goldberg, A. L. & Rock, K. L. Proteolysis, proteasomes and antigen presentation. Nature 357, 375–379 (1992).
Hochstrasser, M. Ubiquitin-dependent protein degradation. Annu. Rev. Genet. 30, 405–439 (1996).
Thrower, J. S., Hoffman, L., Rechsteiner, M. & Pickart, C. M. Recognition of the polyubiquitin proteolytic signal. EMBO J. 19, 94–102 (2000).
Deveraux, Q., Ustrell, V., Pickart, C. & Rechsteiner, M. A 26 S protease subunit that binds ubiquitin conjugates. J. Biol. Chem. 269, 7059–7061 (1994).
Lam, Y. A., Lawson, T. G., Velayutham, M., Zweier, J. L. & Pickart, C. M. A proteasomal ATPase subunit recognizes the polyubiquitin degradation signal. Nature 416, 763–767 (2002).
Husnjak, K. et al. Proteasome subunit Rpn13 is a novel ubiquitin receptor. Nature 453, 481–488 (2008).
Chen, L. & Madura, K. Rad23 promotes the targeting of proteolytic substrates to the proteasome. Mol. Cell. Biol. 22, 4902–4913 (2002).
Funakoshi, M., Sasaki, T., Nishimoto, T. & Kobayashi, H. Budding yeast Dsk2p is a polyubiquitin-binding protein that can interact with the proteasome. Proc. Natl Acad. Sci. USA 99, 745–750 (2002).
Kim, I., Mi, K. & Rao, H. Multiple interactions of rad23 suggest a mechanism for ubiquitylated substrate delivery important in proteolysis. Mol. Biol. Cell 15, 3357–3365 (2004).
Hurley, J. H., Lee, S. & Prag, G. Ubiquitin-binding domains. Biochem. J. 399, 361–372 (2006).
Varadan, R. et al. Solution conformation of Lys63-linked di-ubiquitin chain provides clues to functional diversity of polyubiquitin signaling. J. Biol. Chem. 279, 7055–7063 (2004).
Kerscher, O. SUMO junction — what's your function? New insights through SUMO-interacting motifs. EMBO Rep. 8, 550–555 (2007).
Song, J., Zhang, Z., Hu, W. & Chen, Y. Small ubiquitin-like modifier (SUMO) recognition of a SUMO binding motif: a reversal of the bound orientation. J. Biol. Chem. 280, 40122–40129 (2005).
Reverter, D. & Lima, C. D. Insights into E3 ligase activity revealed by a SUMO–RanGAP1–Ubc9–Nup358 complex. Nature 435, 687–692 (2005).
Hochstrasser, M. in Protein Degradation: The Ubiquitin-Proteasome System (eds Mayer, R. J., Ciechanover, A. & Rechsteiner, M.) 249–278 (Wiley, 2006).
Archer, C. T. et al. Physical and functional interactions of monoubiquitylated transactivators with the proteasome. J. Biol. Chem. 283, 21789–21798 (2008).
Mahajan, R., Delphin, C., Guan, T., Gerace, L. & Melchior, F. A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell 88, 97–107 (1997).
McGrath, J. P., Jentsch, S. & Varshavsky, A. UBA1: an essential yeast gene encoding ubiquitin-activating enzyme. EMBO J. 10, 227–236 (1991).
Rajagopalan, K. V. Biosynthesis and processing of the molybdenum cofactors. Biochem. Soc. Trans. 25, 757–761 (1997).
Taylor, S. V. et al. Thiamin biosynthesis in Escherichia coli. Identification of this thiocarboxylate as the immediate sulfur donor in the thiazole formation. J. Biol. Chem. 273, 16555–16560 (1998).
Appleyard, M. V. et al. The Aspergillus nidulans cnxF gene and its involvement in molybdopterin biosynthesis. Molecular characterization and analysis of in vivo generated mutants. J. Biol. Chem. 273, 14869–14876 (1998).
Leimkuhler, S., Wuebbens, M. M. & Rajagopalan, K. V. Characterization of Escherichia coli MoeB and its involvement in the activation of molybdopterin synthase for the biosynthesis of the molybdenum cofactor. J. Biol. Chem. 276, 34695–34701 (2001).
Rudolph, M. J., Wuebbens, M. M., Rajagopalan, K. V. & Schindelin, H. Crystal structure of molybdopterin synthase and its evolutionary relationship to ubiquitin activation. Nature Struct. Biol. 8, 42–46 (2001).
Wang, C., Xi, J., Begley, T. P. & Nicholson, L. K. Solution structure of ThiS and implications for the evolutionary roots of ubiquitin. Nature Struct. Biol. 8, 47–51 (2001).
Huang, D. T., Walden, H., Duda, D. & Schulman, B. A. Ubiquitin-like protein activation. Oncogene 23, 1958–1971 (2004).
Duda, D. M., Walden, H., Sfondouris, J. & Schulman, B. A. Structural analysis of Escherichia coli ThiF. J. Mol. Biol. 349, 774–786 (2005).
Furukawa, K., Mizushima, N., Noda, T. & Ohsumi, Y. A protein conjugation system in yeast with homology to biosynthetic enzyme reaction of prokaryotes. J. Biol. Chem. 275, 7462–7465 (2000).
Goehring, A. S., Rivers, D. M. & Sprague, G. F. Jr . Attachment of the ubiquitin-related protein Urm1p to the antioxidant protein Ahp1p. Eukaryot. Cell 2, 930–936 (2003).
Schmitz, J. et al. The sulfurtransferase activity of Uba4 presents a link between ubiquitin-like protein conjugation and activation of sulfur carrier proteins. Biochemistry 47, 6479–6489 (2008). Identifies a persulphide in Uba4 and formation of an Urm1 thiocarboxylate, suggesting a dual function in protein conjugation and sulphur transfer.
Mueller, E. G. Trafficking in persulfides: delivering sulfur in biosynthetic pathways. Nature Chem. Biol. 2, 185–194 (2006).
Nakai, Y., Nakai, M. & Hayashi, H. Thio-modification of yeast cytosolic tRNA requires a ubiquitin-related system that resembles bacterial sulfur transfer systems. J. Biol. Chem. 283, 27469–27476 (2008).
Huang, B., Lu, J. & Bystrom, A. S. A genome-wide screen identifies genes required for formation of the wobble nucleoside 5-methoxycarbonylmethyl-2-thiouridine in Saccharomyces cerevisiae . RNA 14, 2183–2194 (2008). This study and ref. 58 implicate the Urm1 thiocarboxylate as a potential sulphur carrier in selective tRNA thiolation.
Burroughs, A. M., Jaffee, M., Iyer, L. M. & Aravind, L. Anatomy of the E2 ligase fold: implications for enzymology and evolution of ubiquitin/Ub-like protein conjugation. J. Struct. Biol. 162, 205–218 (2008).
Amerik, A. Y. & Hochstrasser, M. Mechanism and function of deubiquitinating enzymes. Biochim. Biophys. Acta 1695, 189–207 (2004).
Burns, K. E. et al. Reconstitution of a new cysteine biosynthetic pathway in Mycobacterium tuberculosis . J. Am. Chem. Soc. 127, 11602–11603 (2005).
Godert, A. M., Jin, M., McLafferty, F. W. & Begley, T. P. Biosynthesis of the thioquinolobactin siderophore: an interesting variation on sulfur transfer. J. Bacteriol. 189, 2941–2944 (2007).
Roush, R. F., Nolan, E. M., Lohr, F. & Walsh, C. T. Maturation of an Escherichia coli ribosomal peptide antibiotic by ATP-consuming N–P bond formation in microcin C7. J. Am. Chem. Soc. 130, 3603–3609 (2008). Describes the role of an E1-like adenylating enzyme in the activation of a non-β-grasp peptide.
Pearce, M. J., Mintseris, J., Ferreyra, J., Gygi, S. P. & Darwin, K. H. Ubiquitin-like protein involved in the proteasome pathway of Mycobacterium tuberculosis. Science 322, 1104–1107 (2008). Identifies a non-β-grasp protein in M. tuberculosis that modifies selected protein substrates and targets them for proteasomal degradation.
Iyer, L. M., Burroughs, A. M. & Aravind, L. Unraveling the biochemistry and provenance of pupylation: a prokaryotic analog of ubiquitination. Biol. Direct 3, 45 (2008).
Steinacher, R. & Schar, P. Functionality of human thymine DNA glycosylase requires SUMO-regulated changes in protein conformation. Curr. Biol. 15, 616–623 (2005).
Ulrich, H. D. How to activate a damage-tolerant polymerase: consequences of PCNA modifications by ubiquitin and SUMO. Cell Cycle 3, 15–18 (2004).
Palacios, S. et al. Quantitative SUMO-1 modification of a vaccinia virus protein is required for its specific localization and prevents its self-association. Mol. Biol. Cell 16, 2822–2835 (2005).
Ciechanover, A. & Ben-Saadon, R. N-terminal ubiquitination: more protein substrates join in. Trends Cell Biol. 14, 103–106 (2004).
Cadwell, K. & Coscoy, L. Ubiquitination on nonlysine residues by a viral E3 ubiquitin ligase. Science 309, 127–130 (2005).
Ravid, T. & Hochstrasser, M. Autoregulation of an E2 enzyme by ubiquitin-chain assembly on its catalytic residue. Nature Cell Biol. 9, 422–427 (2007).
Wang, X. et al. Ubiquitination of serine, threonine, or lysine residues on the cytoplasmic tail can induce ERAD of MHC-I by viral E3 ligase mK3. J. Cell Biol. 177, 613–624 (2007).
Lee, I. & Schindelin, H. Structural insights into E1-catalyzed ubiquitin activation and transfer to conjugating enzymes. Cell 134, 268–278 (2008).
Nijman, S. M. et al. A genomic and functional inventory of deubiquitinating enzymes. Cell 123, 773–786 (2005).
Acknowledgements
I thank V. J. Rubenstein, A. Kusmierczyk and J. Bloom for comments on the manuscript. Work in my laboratory is funded by grants from the National Institutes of Health.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Additional information
Reprints and permissions information is available at http://www.nature.com/reprints.
Correspondence should be addressed to the author (mark.hochstrasser@yale.edu).
Rights and permissions
About this article
Cite this article
Hochstrasser, M. Origin and function of ubiquitin-like proteins. Nature 458, 422–429 (2009). https://doi.org/10.1038/nature07958
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07958
This article is cited by
-
Ubiquitin specific peptidase 38 epigenetically regulates KLF transcription factor 5 to augment malignant progression of lung adenocarcinoma
Oncogene (2024)
-
N-terminal α-amino SUMOylation of cofilin-1 is critical for its regulation of actin depolymerization
Nature Communications (2023)
-
pTINCR microprotein promotes epithelial differentiation and suppresses tumor growth through CDC42 SUMOylation and activation
Nature Communications (2022)
-
Emerging roles of non-histone protein crotonylation in biomedicine
Cell & Bioscience (2021)
-
USP7 facilitates SMAD3 autoregulation to repress cancer progression in p53-deficient lung cancer
Cell Death & Disease (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.