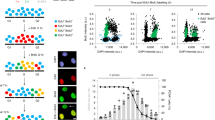
Abstract
The repair of DNA double-strand breaks (DSBs) is tightly regulated during the cell cycle. In G1 phase, the absence of a sister chromatid means that repair of DSBs occurs through non-homologous end-joining or microhomology-mediated end-joining (MMEJ)1. These pathways often involve loss of DNA sequences at the break site and are therefore error-prone. In late S and G2 phases, even though DNA end-joining pathways remain functional2, there is an increase in repair of DSBs by homologous recombination, which is mostly error-free3,4. Consequently, the relative contribution of these different pathways to DSB repair in the cell cycle has a large influence on the maintenance of genetic integrity. It has remained unknown how DSBs are directed for repair by different, potentially competing, repair pathways. Here we identify a role for CtIP (also known as RBBP8) in this process in the avian B-cell line DT40. We establish that CtIP is required not only for repair of DSBs by homologous recombination in S/G2 phase but also for MMEJ in G1. The function of CtIP in homologous recombination, but not MMEJ, is dependent on the phosphorylation of serine residue 327 and recruitment of BRCA1. Cells expressing CtIP protein that cannot be phosphorylated at serine 327 are specifically defective in homologous recombination and have a decreased level of single-stranded DNA after DNA damage, whereas MMEJ remains unaffected. Our data support a model in which phosphorylation of serine 327 of CtIP as cells enter S phase and the recruitment of BRCA1 functions as a molecular switch to shift the balance of DSB repair from error-prone DNA end-joining to error-free homologous recombination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ma, J. L., Kim, E. M., Haber, J. E. & Lee, S. E. Yeast Mre11 and Rad1 proteins define a Ku-independent mechanism to repair double-strand breaks lacking overlapping end sequences. Mol. Cell. Biol. 23, 8820–8828 (2003)
Kim, J. S. et al. Independent and sequential recruitment of NHEJ and HR factors to DNA damage sites in mammalian cells. J. Cell Biol. 170, 341–347 (2005)
Takata, M. et al. Homologous recombination and non-homologous end-joining pathways of DNA double-strand break repair have overlapping roles in the maintenance of chromosomal integrity in vertebrate cells. EMBO J. 17, 5497–5508 (1998)
Pâques, F. & Haber, J. E. Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae . Microbiol. Mol. Biol. Rev. 63, 349–404 (1999)
Sartori, A. A. et al. Human CtIP promotes DNA end resection. Nature 450, 509–514 (2007)
Baumann, P. & West, S. C. Role of the human RAD51 protein in homologous recombination and double-stranded-break repair. Trends Biochem. Sci. 23, 247–251 (1998)
Chen, P. L. et al. Inactivation of CtIP leads to early embryonic lethality mediated by G1 restraint and to tumorigenesis by haploid insufficiency. Mol. Cell. Biol. 25, 3535–3542 (2005)
Sonoda, E., Morrison, C., Yamashita, Y. M., Takata, M. & Takeda, S. Reverse genetic studies of homologous DNA recombination using the chicken B-lymphocyte line, DT40. Phil. Trans. R. Soc. Lond. B 356, 111–117 (2001)
Tauchi, H., Matsuura, S., Kobayashi, J., Sakamoto, S. & Komatsu, K. Nijmegen breakage syndrome gene, NBS1, and molecular links to factors for genome stability. Oncogene 21, 8967–8980 (2002)
Simpson, L. J. & Sale, J. E. Rev1 is essential for DNA damage tolerance and non-templated immunoglobulin gene mutation in a vertebrate cell line. EMBO J. 22, 1654–1664 (2003)
Bridge, W. L., Vandenberg, C. J., Franklin, R. J. & Hiom, K. The BRIP1 helicase functions independently of BRCA1 in the Fanconi anemia pathway for DNA crosslink repair. Nature Genet. 37, 953–957 (2005)
Yu, X. & Chen, J. DNA damage-induced cell cycle checkpoint control requires CtIP, a phosphorylation-dependent binding partner of BRCA1 C-terminal domains. Mol. Cell. Biol. 24, 9478–9486 (2004)
Pierce, A. J., Johnson, R. D., Thompson, L. H. & Jasin, M. XRCC3 promotes homology-directed repair of DNA damage in mammalian cells. Genes Dev. 13, 2633–2638 (1999)
Stark, J. M., Pierce, A. J., Oh, J., Pastink, A. & Jasin, M. Genetic steps of mammalian homologous repair with distinct mutagenic consequences. Mol. Cell. Biol. 24, 9305–9316 (2004)
Bennardo, N., Cheng, A., Huang, N. & Stark, J. M. Alternative-NHEJ is a mechanistically distinct pathway of mammalian chromosome break repair. PLoS Genet. 4, e1000110 (2008)
Yu, X., Wu, L. C., Bowcock, A. M., Aronheim, A. & Baer, R. The C-terminal (BRCT) domains of BRCA1 interact in vivo with CtIP, a protein implicated in the CtBP pathway of transcriptional repression. J. Biol. Chem. 273, 25388–25392 (1998)
Greenberg, R. A. et al. Multifactorial contributions to an acute DNA damage response by BRCA1/BARD1-containing complexes. Genes Dev. 20, 34–46 (2006)
Schlegel, B. P., Jodelka, F. M. & Nunez, R. BRCA1 promotes induction of ssDNA by ionizing radiation. Cancer Res. 66, 5181–5189 (2006)
Endicott, J. A., Noble, M. E. & Tucker, J. A. Cyclin-dependent kinases: inhibition and substrate recognition. Curr. Opin. Struct. Biol. 9, 738–744 (1999)
Ira, G. et al. DNA end resection, homologous recombination and DNA damage checkpoint activation require CDK1. Nature 431, 1011–1017 (2004)
Huertas, P., Cortes-Ledesma, F., Sartori, A. A., Aguilera, A. & Jackson, S. P. CDK targets Sae2 to control DNA-end resection and homologous recombination. Nature 455, 689–692 (2008)
Acknowledgements
The authors would like to thank M. Jasin for gifts of DR–GFP and pHPRT-SSA-GFP, S. Takeda for the gift of KU70 DT40, and J. Di Noia for DT40 DTDR-17. We would also like to thank our colleagues C. Rada and J. Sale for comments and suggestions during the preparation of this manuscript. M.H.Y. is a Milstein Student of the Darwin Trust, Edinburgh, Scotland.
Author Contributions All experiments were performed by M.H.Y. and were conceived by M.H.Y. and K.H. K.H. and M.H.Y. wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figures
This file contains Supplementary Figures S1-S11 with Legends (PDF 6933 kb)
Rights and permissions
About this article
Cite this article
Yun, M., Hiom, K. CtIP-BRCA1 modulates the choice of DNA double-strand-break repair pathway throughout the cell cycle. Nature 459, 460–463 (2009). https://doi.org/10.1038/nature07955
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07955
This article is cited by
-
Recurrence mutation in RBBP8 gene causing non-syndromic autosomal recessive primary microcephaly; geometric simulation approach for insight into predicted computational models
Journal of Human Genetics (2023)
-
The oncogenic fusion protein TAZ::CAMTA1 promotes genomic instability and senescence through hypertranscription
Communications Biology (2023)
-
Prognostic Significance of mRNA Expression RBBP8 or Its Methylation in Gliomas
Cellular and Molecular Neurobiology (2023)
-
Harnessing DSB repair to promote efficient homology-dependent and -independent prime editing
Nature Communications (2022)
-
DHX9-dependent recruitment of BRCA1 to RNA promotes DNA end resection in homologous recombination
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.