Abstract
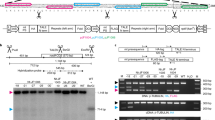
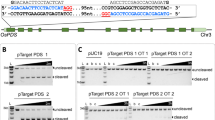
An efficient method for making directed DNA sequence modifications to plant genes (gene targeting) is at present lacking, thereby frustrating efforts to dissect plant gene function and engineer crop plants that better meet the world’s burgeoning need for food, fibre and fuel. Zinc-finger nucleases (ZFNs)—enzymes engineered to create DNA double-strand breaks at specific loci—are potent stimulators of gene targeting1,2; for example, they can be used to precisely modify engineered reporter genes in plants3,4. Here we demonstrate high-frequency ZFN-stimulated gene targeting at endogenous plant genes, namely the tobacco acetolactate synthase genes (ALS SuRA and SuRB), for which specific mutations are known to confer resistance to imidazolinone and sulphonylurea herbicides5. Herbicide-resistance mutations were introduced into SuR loci by ZFN-mediated gene targeting at frequencies exceeding 2% of transformed cells for mutations as far as 1.3 kilobases from the ZFN cleavage site. More than 40% of recombinant plants had modifications in multiple SuR alleles. The observed high frequency of gene targeting indicates that it is now possible to efficiently make targeted sequence changes in endogenous plant genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Bibikova, M., Beumer, K., Trautman, J. K. & Carroll, D. Enhancing gene targeting with designed zinc finger nucleases. Science 300, 764 (2003)
Porteus, M. H. & Baltimore, D. Chimeric nucleases stimulate gene targeting in human cells. Science 300, 763 (2003)
Lloyd, A., Plaisier, C. L., Carroll, D. & Drews, G. N. Targeted mutagenesis using zinc-finger nucleases in Arabidopsis . Proc. Natl Acad. Sci. USA 102, 2232–2237 (2005)
Wright, D. A. et al. High-frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J. 44, 693–705 (2005)
Lee, K. Y. et al. The molecular basis of sulfonylurea herbicide resistance in tobacco. EMBO J. 7, 1241–1248 (1988)
Maeder, M. L. et al. Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol. Cell 31, 294–301 (2008)
Wright, D. A. et al. Standardized reagents and protocols for engineering zinc finger nucleases by modular assembly. Nature Protocols 1, 1637–1652 (2006)
Ramirez, C. L. et al. Unexpected failure rates for modular assembly of engineered zinc fingers. Nature Methods 5, 374–375 (2008)
Segal, D. J., Dreier, B., Beerli, R. R. & Barbas, C. F. Toward controlling gene expression at will: Selection and design of zinc finger domains recognizing each of the 5′-GNN-3′ DNA target sequences. Proc. Natl Acad. Sci. USA 96, 2758–2763 (1999)
Pruett-Miller, S. M., Connelly, J. P., Maeder, M. L., Joung, J. K. & Porteus, M. H. Comparison of zinc finger nucleases for use in gene targeting in mammalian cells. Mol. Ther. 16, 707–717 (2008)
Cornu, T. I. et al. DNA-binding specificity is a major determinant of the activity and toxicity of zinc-finger nucleases. Mol. Ther. 16, 352–358 (2008)
Ronaghi, M., Uhlen, M. & Nyren, P. A sequencing method based on real-time pyrophosphate. Science 281, 363–365 (1998)
Tranel, P. & Wright, T. Resistance of weeds to ALS-inhibiting herbicides: What have we learned? Weed Sci. 50, 700–712 (2002)
Kaeppler, S. M., Kaeppler, H. F. & Rhee, Y. Epigenetic aspects of somoclonal variation in plants. Plant Mol. Biol. 43, 179–188 (2000)
Szczepek, M. et al. Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases. Nature Biotechnol. 25, 786–793 (2007)
Miller, J. C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nature Biotechnol. 25, 778–785 (2007)
Epinat, J. C. et al. A novel engineered meganuclease induces homologous recombination in yeast and mammalian cells. Nucleic Acids Res. 31, 2952–2962 (2003)
Doyon, Y. et al. Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nature Biotechnol. 26, 702–708 (2008)
Acknowledgements
We thank M. Eichtinger for help in making ZFA reagents. This work was supported by grants to D.F.V. from the National Science Foundation and to J.K.J. from the National Institutes of Health and the Massachusetts General Hospital Department of Pathology.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-2 with Legends and Supplementary Tables 1-7. (PDF 574 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Townsend, J., Wright, D., Winfrey, R. et al. High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 459, 442–445 (2009). https://doi.org/10.1038/nature07845
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07845
This article is cited by
-
Intragenic Agrobacterium-mediated gene transfer mimics micro-translocations without foreign DNA
Planta (2024)
-
Genome editing in cotton: challenges and opportunities
Journal of Cotton Research (2023)
-
Regulation of gene-edited plants in Europe: from the valley of tears into the shining sun?
aBIOTECH (2023)
-
Salinity Stress Tolerance in Solanaceous Crops: Current Understanding and Its Prospects in Genome Editing
Journal of Plant Growth Regulation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.