Abstract
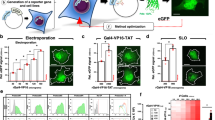
In-cell NMR is an isotope-aided multi-dimensional NMR technique that enables observations of conformations and functions of proteins in living cells at the atomic level1. This method has been successfully applied to proteins overexpressed in bacteria, providing information on protein–ligand interactions2 and conformations3,4. However, the application of in-cell NMR to eukaryotic cells has been limited to Xenopus laevis oocytes5,6,7. Wider application of the technique is hampered by inefficient delivery of isotope-labelled proteins into eukaryote somatic cells. Here we describe a method to obtain high-resolution two-dimensional (2D) heteronuclear NMR spectra of proteins inside living human cells. Proteins were delivered to the cytosol by the pyrenebutyrate-mediated action of cell-penetrating peptides8 linked covalently to the proteins. The proteins were subsequently released from cell-penetrating peptides by endogenous enzymatic activity or by autonomous reductive cleavage. The heteronuclear 2D spectra of three different proteins inside human cells demonstrate the broad application of this technique to studying interactions and protein processing. The in-cell NMR spectra of FKBP12 (also known as FKBP1A) show the formation of specific complexes between the protein and extracellularly administered immunosuppressants, demonstrating the utility of this technique in drug screening programs. Moreover, in-cell NMR spectroscopy demonstrates that ubiquitin has much higher hydrogen exchange rates in the intracellular environment, possibly due to multiple interactions with endogenous proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Serber, Z. & Dotsch, V. In-cell NMR spectroscopy. Biochemistry 40, 14317–14323 (2001)
Burz, D. S., Dutta, K., Cowburn, D. & Shekhtman, A. Mapping structural interactions using in-cell NMR spectroscopy (STINT-NMR). Nature Methods 3, 91–93 (2006)
Dedmon, M. M., Patel, C. N., Young, G. B. & Pielak, G. J. FlgM gains structure in living cells. Proc. Natl Acad. Sci. USA 99, 12681–12684 (2002)
Sakakibara, D. et al. Protein structure determination in living cells by in-cell NMR spectroscopy. Nature 10.1038/nature07814 (this issue)
Sakai, T. et al. In-cell NMR spectroscopy of proteins inside Xenopus laevis oocytes. J. Biomol. NMR 36, 179–188 (2006)
Selenko, P. et al. Quantitative NMR analysis of the protein G B1 domain in Xenopus laevis egg extracts and intact oocytes. Proc. Natl Acad. Sci. USA 103, 11904–11909 (2006)
Selenko, P. et al. In situ observation of protein phosphorylation by high-resolution NMR spectroscopy. Nature Struct. Mol. Biol. 15, 321–329 (2008)
Takeuchi, T. et al. Direct and rapid cytosolic delivery using cell-penetrating peptides mediated by pyrenebutyrate. ACS Chem. Biol. 1, 299–303 (2006)
Futaki, S. Oligoarginine vectors for intracellular delivery: design and cellular-uptake mechanisms. Biopolymers 84, 241–249 (2006)
Nakase, I., Takeuchi, T., Tanaka, G. & Futaki, S. Methodological and cellular aspects that govern the internalization mechanisms of arginine-rich cell-penetrating peptides. Adv. Drug Deliv. Rev. 60, 598–607 (2008)
Wender, P. A. et al. The design of guanidinium-rich transporters and their internalization mechanisms. Adv. Drug Deliv. Rev. 60, 452–472 (2008)
Schwarze, S. R., Ho, A., Vocero-Akbani, A. & Dowdy, S. F. In vivo protein transduction: delivery of a biologically active protein into the mouse. Science 285, 1569–1572 (1999)
Loison, F. et al. A ubiquitin-based assay for the cytosolic uptake of protein transduction domains. Mol. Ther. 11, 205–214 (2005)
Carlson, N. & Rechsteiner, M. Microinjection of ubiquitin - intracellular-distribution and metabolism in Hela-cells maintained under normal physiological conditions. J. Cell Biol. 104, 537–546 (1987)
O’Brien, R. & Gottlieb-Rosenkrantz, P. An automatic method for viability assay of cultured cells. J. Histochem. Cytochem. 18, 581–589 (1970)
Giriat, I. & Muir, T. W. Protein semi-synthesis in living cells. J. Am. Chem. Soc. 125, 7180–7181 (2003)
Hicke, L., Schubert, H. L. & Hill, C. P. Ubiquitin-binding domains. Nature Rev. Mol. Cell Biol. 6, 610–621 (2005)
Itoh, S. & Navia, M. A. Structure comparison of native and mutant human recombinant FKBP12 complexes with the immunosuppressant drug FK506 (tacrolimus). Protein Sci. 4, 2261–2268 (1995)
Ellis, R. J. Macromolecular crowding: obvious but underappreciated. Trends Biochem. Sci. 26, 597–604 (2001)
Bai, Y. et al. Thermodynamic parameters from hydrogen exchange measurements. Methods Enzymol. 259, 344–356 (1995)
Lange, O. F. et al. Recognition dynamics up to microseconds revealed from an RDC-derived ubiquitin ensemble in solution. Science 320, 1471–1475 (2008)
Delom, F., Fessart, D., Caruso, M. E. & Chevet, E. Tat-mediated protein delivery in living Caenorhabditis elegans . Biochem. Biophys. Res. Commun. 352, 587–591 (2007)
Schanda, P. & Brutscher, B. Very fast two-dimensional NMR spectroscopy for real-time investigation of dynamic events in proteins on the time scale of seconds. J. Am. Chem. Soc. 127, 8014–8015 (2005)
Laue, E. D., Mayger, M. R., Skilling, J. & Staunton, J. Reconstruction of phase-sensitive two-dimensional NMR-spectra by maximum-entropy. J. Magn. Reson. 68, 14–29 (1986)
Tenno, T. et al. Structural basis for distinct roles of Lys63- and Lys48-linked polyubiquitin chains. Genes Cells 9, 865–875 (2004)
Mezo, G., Mihala, N., Andreu, D. & Hudecz, F. Conjugation of epitope peptides with SH group to branched chain polymeric polypeptides via Cys(Npys). Bioconjug. Chem. 11, 484–491 (2000)
Rosen, M. K., Michnick, S. W., Karplus, M. & Schreiber, S. L. Proton and nitrogen sequential assignments and secondary structure determination of the human FK506 and rapamycin binding protein. Biochemistry 30, 4774–4789 (1991)
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995)
Goddard, T. D. & Kneller, D. G. SPARKY 3 (University of California, 1999)
Acknowledgements
We thank M. Waelchli, A. Kidera and H. Akutsu for discussion, T. Kokubo for monkey COS-7 cells, H. Ohnishi for the plasmid for production of FKBP12 and M. Imanishi for taking gel fluorimaging. This work was supported by grants to M.S. from Japan Science and Technology Agency and the Ministry of Education, Culture, Sports, Science and Technology-Japan (MEXT), and also in part by the Global COE Program ‘International Center for Integrated Research and Advanced Education in Materials Science’ (No. B-09) of MEXT, administered by the Japan Society for the Promotion of Science. This work was partly supported by the Innovative Techno-Hub for Integrated Medical Bio-imaging Project of the Special Coordination Funds for Promoting Science and Technology, from MEXT to A.O. and M.S., and by grants from MEXT to S.F. and H.T.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Figures
This file contains Supplementary Figures S1-S5 with Legends (PDF 358 kb)
Rights and permissions
About this article
Cite this article
Inomata, K., Ohno, A., Tochio, H. et al. High-resolution multi-dimensional NMR spectroscopy of proteins in human cells. Nature 458, 106–109 (2009). https://doi.org/10.1038/nature07839
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature07839
This article is cited by
-
Real-time monitoring of the reaction of KRAS G12C mutant specific covalent inhibitor by in vitro and in-cell NMR spectroscopy
Scientific Reports (2023)
-
In-cell NMR: recent progresses and future challenges
Rendiconti Lincei. Scienze Fisiche e Naturali (2023)
-
A thermosensitive gel matrix for bioreactor-assisted in-cell NMR of nucleic acids and proteins
Journal of Biomolecular NMR (2023)
-
Microfluidics delivery of DARPP-32 into HeLa cells maintains viability for in-cell NMR spectroscopy
Communications Biology (2022)
-
Protein delivery to living cells by thermal stimulation for biophysical investigation
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.