Abstract
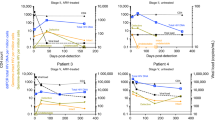
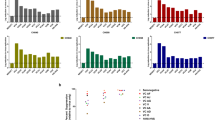
The rapid and extensive spread of the human immunodeficiency virus (HIV) epidemic provides a rare opportunity to witness host–pathogen co-evolution involving humans. A focal point is the interaction between genes encoding human leukocyte antigen (HLA) and those encoding HIV proteins. HLA molecules present fragments (epitopes) of HIV proteins on the surface of infected cells to enable immune recognition and killing by CD8+ T cells; particular HLA molecules, such as HLA-B*57, HLA-B*27 and HLA-B*51, are more likely to mediate successful control of HIV infection1. Mutation within these epitopes can allow viral escape from CD8+ T-cell recognition. Here we analysed viral sequences and HLA alleles from >2,800 subjects, drawn from 9 distinct study cohorts spanning 5 continents. Initial analysis of the HLA-B*51-restricted epitope, TAFTIPSI (reverse transcriptase residues 128–135), showed a strong correlation between the frequency of the escape mutation I135X and HLA-B*51 prevalence in the 9 study cohorts (P = 0.0001). Extending these analyses to incorporate other well-defined CD8+ T-cell epitopes, including those restricted by HLA-B*57 and HLA-B*27, showed that the frequency of these epitope variants (n = 14) was consistently correlated with the prevalence of the restricting HLA allele in the different cohorts (together, P < 0.0001), demonstrating strong evidence of HIV adaptation to HLA at a population level. This process of viral adaptation may dismantle the well-established HLA associations with control of HIV infection that are linked to the availability of key epitopes, and highlights the challenge for a vaccine to keep pace with the changing immunological landscape presented by HIV.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
Accession numbers for newly determined viral sequences are included in Supplementary Information.
References
Goulder, P. J. R. & Watkins, D. I. Impact of MHC class I diversity on immune control of immunodeficiency virus replication. Nature Rev. Immunol. 8, 619–630 (2008)
Goulder, P. J. R. et al. Evolution and transmission of stable CTL escape mutations in HIV infection. Nature 412, 334–338 (2001)
Moore, C. B. et al. Evidence of HIV-1 adaptation to HLA-restricted immune responses at a population level. Science 296, 1439–1443 (2002)
Draenert, R. et al. Immune selection for altered antigen processing leads to cytotoxic T lymphocyte escape in chronic HIV-1 infection. J. Exp. Med. 199, 905–915 (2004)
Leslie, A. J. et al. Transmission and accumulation of CTL escape variants drive negative associations between HIV polymorphisms and HLA. J. Exp. Med. 201, 891–902 (2005)
Bhattacharya, T. et al. Founder effects in the assessment of HIV polymorphisms and HLA allele associations. Science 315, 1583–1586 (2007)
Matthews, P. et al. Central role of reverting mutations in HLA associations with viral setpoint. J. Virol. 82, 8548–8559 (2008)
Tomiyama, H. et al. Identification of multiple HIV-1 CTL epitopes presented by HLA-B*5101 molecules. Hum. Immunol. 60, 177–186 (1999)
Brumme, Z. et al. Human leukocyte antigen-specific polymorphisms in HIV-1 Gag and their association with viral load in chronic untreated infection. AIDS 22, 1277–1286 (2008)
Itoh, Y. et al. High throughput DNA tying of HLA-A, -B, -C, and -DRB1 loci by a PCR-SSOP-Luminex method in the Japanese population. Immunogenetics 57, 717–729 (2005)
Kaslow, R. A. et al. Influence of combinations of human major histocompatibility complex genes on the course of HIV-1 infection. Nature Med. 2, 405–411 (1996)
O’Brien, S. J., Gao, X. & Carrington, M. HLA and AIDS: a cautionary tale. Trends Mol. Med. 7, 379–381 (2001)
Kiepiela, P. et al. CD8+ T-cell responses to different HIV proteins have discordant associations with viral load. Nature Med. 13, 46–53 (2007)
Leslie, A. J. et al. HIV evolution: CTL escape mutation and reversion after transmission. Nature Med. 10, 282–289 (2004)
Goulder, P. J. R. et al. Late escape from an immunodominant cytotoxic T-lymphocyte response associated with progression to AIDS. Nature Med. 3, 212–217 (1997)
Feeney, M. E. et al. Immune escape precedes breakthrough HIV-1 viremia and broadening of the CTL response in a HLA-B27-positive long-term nonprogressing child. J. Virol. 78, 8927–8930 (2004)
Martinez-Picado, J. et al. Fitness cost of escape mutations in p24 Gag in association with control of human immunodeficiency virus type 1. J. Virol. 80, 3617–3623 (2006)
Crawford, H. et al. Compensatory mutation partially restores fitness and delays reversion of escape mutation within the immunodominant HLA-B*5703-restricted Gag epitope in chronic human immunodeficiency virus type 1 infection. J. Virol. 81, 8346–8351 (2007)
Schneidewind, A. et al. Escape from a dominant Gag-specific CTL response in HLA-B27+ subjects is associated with a dramatic reduction in HIV-1 replication. J. Virol. 81, 12382–12393 (2007)
Brumme, Z. et al. Marked epitope- and allele-specific differences in rates of mutation in human immunodeficiency type 1 (HIV-1) Gag, Pol, and Nef cytotoxic T-lymphocyte epitopes in acute/early HIV-1 infection. J. Virol. 82, 9216–9227 (2008)
Goepfert, P. et al. Transmission of Gag immune escape mutations is associated with reduced viral load in linked recipients. J. Exp. Med. 205, 1009–1017 (2008)
Seki, S. et al. Transmission of SIV carrying multiple cytotoxic T lymphocyte escape mutations with diminished replicative capacity can result in AIDS progression in Rhesus macaques. J. Virol. 82, 5093–5098 (2008)
Gallimore, A., Dumrese, T., Hengartner, H., Zinkernagel, R. M. & Rammensee, H. G. Protective immunity does not correlate with the hierarchy of virus-specific cytotoxic T cell responses to naturally processed peptides. J. Exp. Med. 187, 1647–1657 (1998)
Holtappels, R. et al. Subdominant CD8 T-cell epitopes account for protection against cytomegalovirus independent of immunodomination. J. Virol. 82, 5781–5796 (2008)
McKiernan, S. M. et al. Distinct MHC class I and II alleles are associated with hepatitis C viral clearance, originating from a single source. Hepatology 40, 108–114 (2004)
Kiepiela, P. et al. Dominant influence of HLA-B in mediating the potential co-evolution of HIV and HLA. Nature 432, 769–775 (2004)
Tang, J. et al. Favorable and unfavorable HLA class I alleles and haplotypes in Zambians predominantly infected with clade C human immunodeficiency virus type 1. J. Virol. 76, 8276–8284 (2002)
Agresti, A. & Coull, B. Approximate is better than ‘exact’ for interval estimation of binomial proportions. Am. Stat. 52, 119–126 (1998)
Gatanaga, H., Hachiya, A., Kimura, S. & Oka, S. Mutations other than 103N in human immunodeficiency virus type 1 reverse transcriptase (RT) emerge from K103R polymorphism under non-nucleoside RT inhibitor pressure. Virology 344, 354–362 (2006)
Tomiyama, H., Akari, H., Adachi, A. & Takiguchi, M. Different effects of Nef-mediated HLA class I down-regulation on HIV-1-specific CD8+ T cell cytokine activity and cytokine production. J. Virol. 76, 7535–7543 (2002)
Acknowledgements
This work is funded by grants from the National Institutes of Health (RO1AI46995 (P.G.), 1 R01 AI067073 (B.D.W.), R01AI64060 (E.H.)), the Wellcome Trust (P.G., P.K.), the UK Medical Research Council (J.F., A.P. and P.M.), and the Mark and Lisa Schwartz Foundation, the Ministry of Health, Labour and Welfare (Health and Labour HIV/AIDS Research Grants 012), the NIHR Biomedical Research Centre Programme and the Ministry of Education, Science, Sports and Culture (number 18390141), Japan (M.T.). P.G. is an Elizabeth Glaser Pediatric AIDS Foundation Scientist; J.G.P. is a Marie Curie Fellow (contract number IEF-041811). The authors are also grateful to A. McLean and H. Fryer for discussions of the manuscript.
Author Contributions Y.K., K.P., J.F. and P. M. undertook much of the experimental work and data analysis, and contributed equally. M.T. and P.G. undertook much of the project conception, planning, supervision, analysis and writing of the manuscript, and contributed equally.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-7 with Legends, Supplementary Tables 1-3 and Supplementary Notes (PDF 421 kb)
Rights and permissions
About this article
Cite this article
Kawashima, Y., Pfafferott, K., Frater, J. et al. Adaptation of HIV-1 to human leukocyte antigen class I. Nature 458, 641–645 (2009). https://doi.org/10.1038/nature07746
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07746
This article is cited by
-
The chemokine receptor CCR5: multi-faceted hook for HIV-1
Retrovirology (2024)
-
Employing computational tools to design a multi-epitope vaccine targeting human immunodeficiency virus-1 (HIV-1)
BMC Genomics (2023)
-
Genomic characteristics of two most widely used BCG vaccine strains: Danish 1331 and Pasteur 1173P2
BMC Genomics (2022)
-
Accumulation of mutations in antibody and CD8 T cell epitopes in a B cell depleted lymphoma patient with chronic SARS-CoV-2 infection
Nature Communications (2022)
-
Subtype-specific differences in Gag-protease replication capacity of HIV-1 isolates from East and West Africa
Retrovirology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.