Abstract
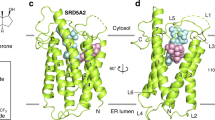
Aromatase cytochrome P450 is the only enzyme in vertebrates known to catalyse the biosynthesis of all oestrogens from androgens1,2,3. Aromatase inhibitors therefore constitute a frontline therapy for oestrogen-dependent breast cancer3,4. In a three-step process, each step requiring 1 mol of O2, 1 mol of NADPH, and coupling with its redox partner cytochrome P450 reductase, aromatase converts androstenedione, testosterone and 16α-hydroxytestosterone to oestrone, 17β-oestradiol and 17β,16α-oestriol, respectively1,2,3. The first two steps are C19-methyl hydroxylation steps, and the third involves the aromatization of the steroid A-ring, unique to aromatase. Whereas most P450s are not highly substrate selective, it is the hallmark androgenic specificity that sets aromatase apart. The structure of this enzyme of the endoplasmic reticulum membrane has remained unknown for decades, hindering elucidation of the biochemical mechanism. Here we present the crystal structure of human placental aromatase, the only natural mammalian, full-length P450 and P450 in hormone biosynthetic pathways to be crystallized so far. Unlike the active sites of many microsomal P450s that metabolize drugs and xenobiotics, aromatase has an androgen-specific cleft that binds the androstenedione molecule snugly. Hydrophobic and polar residues exquisitely complement the steroid backbone. The locations of catalytically important residues shed light on the reaction mechanism. The relative juxtaposition of the hydrophobic amino-terminal region and the opening to the catalytic cleft shows why membrane anchoring is necessary for the lipophilic substrates to gain access to the active site. The molecular basis for the enzyme’s androgenic specificity and unique catalytic mechanism can be used for developing next-generation aromatase inhibitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Thompson, E. A. & Siiteri, P. K. Utilization of oxygen and reduced nicotinamide adenine dinucleotide phosphate by human placental microsomes during aromatization of androstenedione. J. Biol. Chem. 249, 5364–5372 (1974)
Simpson, E. R. et al. Aromatase cytochrome P450, the enzyme responsible for estrogen biosynthesis. Endocr. Rev. 15, 342–355 (1994)
O’Neal Johnston, J. Aromatase inhibitors. Crit. Rev. Biochem. Mol. Biol. 33, 375–405 (1998)
Eisen, A., Trudeau, M., Shelley, W., Messersmith, H. & Pritchard, K. I. Aromatase inhibitors in adjuvant therapy for hormone receptor positive breast cancer: A systematic review. Cancer Treat. Rev. 34, 157–174 (2008)
Akhtar, M., Calder, D. L., Corina, D. L. & Wright, J. N. Mechanistic studies on C19-demethylation in oestrogen biosynthesis. Biochem. J. 201, 569–580 (1982)
Akhtar, M., Njar, V. C. & Wright, J. N. Mechanistic studies on aromatase and related C–C bond cleaving P-450 enzymes. J. Steroid Biochem. Mol. Biol. 44, 375–387 (1993)
Nakajin, S., Shinoda, M. & Hall, P. F. Purification to homogeneity of aromatase from human placenta. Biochem. Biophys. Res. Commun. 134, 704–710 (1986)
Kellis, J. T. & Vickery, L. E. Purification and characterization of human placental aromatase cytochrome P-450. J. Biol. Chem. 262, 4413–4420 (1987)
Zhou, D., Pompon, D. & Chen, S. Structure–function studies of human aromatase by site-directed mutagenesis: kinetic properties of mutants Pro-308–Phe, Tyr-361–Phe, Tyr-361–Leu, and Phe-406–Arg. Proc. Natl Acad. Sci. USA 88, 410–414 (1991)
Kadohama, N., Yarborough, C., Zhou, D., Chen, S. & Osawa, Y. Kinetic properties of aromatase mutants Pro 308Phe, Asp 309Asn, and Asp 309Ala and their interactions with aromatase inhibitors. J. Steroid Biochem. Mol. Biol. 43, 693–701 (1992)
Chen, S. et al. Structure–function studies of human aromatase. J. Steroid Biochem. Mol. Biol. 44, 347–356 (1993)
Laughton, C. A., Zvelebil, M. J. & Neidle, S. A detailed molecular model for human aromatase. J. Steroid Biochem. Mol. Biol. 44, 399–407 (1993)
Oh, S. S. & Robinson, C. H. Mechanism of human placental aromatase: a new active site model. J. Steroid Biochem. Mol. Biol. 44, 389–397 (1993)
Amarneh, B. & Simpson, E. R. Expression of a recombinant derivative of human aromatase P450 in insect cells utilizing the baculovirus vector system. Mol. Cell. Endocrinol. 109, R1–R5 (1995)
Graham-Lorence, S., Amarneh, B., White, R. E., Peterson, J. A. & Simpson, E. R. A three-dimensional model of aromatase cytochrome P450. Protein Sci. 4, 1065–1080 (1995)
Kao, Y. C., Korzekwa, K. R., Laughton, C. A. & Chen, S. Evaluation of the mechanism of aromatase cytochrome P450. A site-directed mutagenesis study. Eur. J. Biochem. 268, 243–251 (2001)
Chen, S. et al. Structure–function studies of aromatase and its inhibitors: a progress report. J. Steroid Biochem. Mol. Biol. 86, 231–237 (2003)
Hong, Y., Cho, M., Yuan, Y. C. & Chen, S. Molecular basis for the interaction of four different classes of substrates and inhibitors with human aromatase. Biochem. Pharmacol. 75, 1161–1169 (2008)
Lala, P. et al. Suppression of human cytochrome P450 aromatase activity by monoclonal and recombinant antibody fragments and identification of their stable antigenic complex. J. Steroid Biochem. Mol. Biol. 88, 235–245 (2004)
Nagano, S. & Poulos, T. L. Crystallographic study on the dioxygen complex of wild-type and mutant cytochrome P450cam. Implications for the dioxygen activation mechanism. J. Biol. Chem. 280, 31659–31663 (2005)
Nagano, S., Cupp-Vickery, J. R. & Poulos, T. L. Crystal structures of the ferrous dioxygen complex of wild-type cytochrome P450eryF and its mutants, A245S and A245T: investigation of the proton transfer system in P450eryF. J. Biol. Chem. 280, 22102–22107 (2005)
Williams, P. A. et al. Crystal structures of human cytochrome P450 3A4 bound to metyrapone and progesterone. Science 305, 683–686 (2004)
Rowland, P. et al. Crystal structure of human cytochrome P450 2D6. J. Biol. Chem. 281, 7614–7622 (2006)
Sansen, S., Hsu, M. H., Stout, C. D. & Johnson, E. F. Structural insight into the altered substrate specificity of human cytochrome P450 2A6 mutants. Arch. Biochem. Biophys. 464, 197–206 (2007)
Guallar, V., Baik, M. H., Lippard, S. J. & Friesner, R. A. Peripheral heme substituents control the hydrogen-atom abstraction chemistry in cytochromes P450. Proc. Natl Acad. Sci. USA 100, 6998–7002 (2003)
Podust, L. M., Poulos, T. L. & Waterman, M. R. Crystal structure of cytochrome P450 14α-sterol demethylase (CYP51) from Mycobacterium tuberculosis in complex with azole inhibitors. Proc. Natl Acad. Sci. USA 98, 3068–3073 (2001)
Hackett, J. C., Brueggemeier, R. W. & Hadad, C. M. The final catalytic step of cytochrome p450 aromatase: a density functional theory study. J. Am. Chem. Soc. 127, 5224–5237 (2005)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Cojocaru, V., Winn, P. J. & Wade, R. C. The ins and outs of cytochrome P450s. Biochim. Biophys. Acta 1770, 390–401 (2007)
Shimozawa, O. et al. Core glycosylation of cytochrome P-450(arom). Evidence for localization of N terminus of microsomal cytochrome P-450 in the lumen. J. Biol. Chem. 268, 21399–21402 (1993)
Yoshida, N. & Osawa, Y. Purification of human placental aromatase cytochrome P-450 with monoclonal antibody and its characterization. Biochemistry 30, 3003–3010 (1991)
Sato, R. & Omura, T. Cytochrome P-450 (Kodansha/Academic, 1978)
Otninowski, Z. & Minor, W. HKL Manual (Yale University, 1995)
Collaborative Computational Project Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Emsley, P. & Cowtan, K. Coot: model building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Acknowledgements
We thank Y. Osawa, who pioneered aromatase research and its purification from human placenta at the institute, for many discussions and encouragement; past graduate students, postdoctoral scientists and research associates for contributions to the initial purification and crystallization efforts; H. Davies for discussions; D. Gewirth, V. Cody, G. DeTitta and J. Griffin for critically reading the manuscript; staff at the Women’s and Children’s Hospital of Buffalo for providing the placenta used in this work; and staffs of the Cornell High Energy Synchrotron Source and the Advanced Photon Source, Argonne National Laboratory, for helping with the synchrotron X-ray data collection. The research is supported in part by grants GM62794 and GM59450 (to D.G.) from the National Institutes of Health.
Author Contributions J.G. and M.E. performed the purification and crystallization of aromatase. W.P. and J.G. contributed to diffraction data collection. D.G. was involved in diffraction data collection and processing. D.G. solved the structure, wrote the manuscript and was responsible for overall planning and supervision of the project.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Table 1 and supplementary Figures 1-5 with Legends (PDF 5702 kb)
Rights and permissions
About this article
Cite this article
Ghosh, D., Griswold, J., Erman, M. et al. Structural basis for androgen specificity and oestrogen synthesis in human aromatase. Nature 457, 219–223 (2009). https://doi.org/10.1038/nature07614
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature07614
This article is cited by
-
Design, Synthesis, and Characterization of Cinnamic Acid Derivatives with Two Novel Acrylohydrazones on HeLa and CHO-1 Cancer Cell Lines: The Experimental and Computational Perspective
Chemistry Africa (2024)
-
Developing multitarget coumarin based anti-breast cancer agents: synthesis and molecular modeling study
Scientific Reports (2023)
-
Scorpion Venom Peptide Smp24 Revealed Apoptotic and Antiangiogenic Activities in Solid-Ehrlich Carcinoma Bearing Mice
International Journal of Peptide Research and Therapeutics (2023)
-
Polymorphism on human aromatase affects protein dynamics and substrate binding: spectroscopic evidence
Biology Direct (2021)
-
Identification of the anti-breast cancer targets of triterpenoids in Liquidambaris Fructus and the hints for its traditional applications
BMC Complementary Medicine and Therapies (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.