Abstract
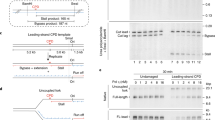
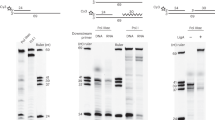
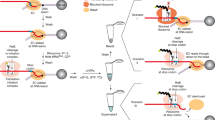
Replication forks are impeded by DNA damage and protein–nucleic acid complexes such as transcribing RNA polymerase. For example, head-on collision of the replisome with RNA polymerase results in replication fork arrest. However, co-directional collision of the replisome with RNA polymerase has little or no effect on fork progression. Here we examine co-directional collisions between a replisome and RNA polymerase in vitro. We show that the Escherichia coli replisome uses the RNA transcript as a primer to continue leading-strand synthesis after the collision with RNA polymerase that is displaced from the DNA. This action results in a discontinuity in the leading strand, yet the replisome remains intact and bound to DNA during the entire process. These findings underscore the notable plasticity by which the replisome operates to circumvent obstacles in its path and may explain why the leading strand is synthesized discontinuously in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mirkin, E. V. & Mirkin, S. M. Replication fork stalling at natural impediments. Microbiol. Mol. Biol. Rev. 71, 13–35 (2007)
Rudolph, C. J., Dhillon, P., Moore, T. & Lloyd, R. G. Avoiding and resolving conflicts between DNA replication and transcription. DNA Repair (Amst.) 6, 981–993 (2007)
Cox, M. M. et al. The importance of repairing stalled replication forks. Nature 404, 37–41 (2000)
Cox, M. M. Recombinational DNA repair of damaged replication forks in Escherichia coli: questions. Annu. Rev. Genet. 35, 53–82 (2001)
Brewer, B. J. When polymerases collide: replication and the transcriptional organization of the E. coli chromosome. Cell 53, 679–686 (1988)
Jarosz, D. F., Beuning, P. J., Cohen, S. E. & Walker, G. C. Y-family DNA polymerases in Escherichia coli. Trends Microbiol. 15, 70–77 (2007)
Goodman, M. F. Error-prone repair DNA polymerases in prokaryotes and eukaryotes. Annu. Rev. Biochem. 71, 17–50 (2002)
Tippin, B., Pham, P. & Goodman, M. F. Error-prone replication for better or worse. Trends Microbiol. 12, 288–295 (2004)
Lusetti, S. L. & Cox, M. M. The bacterial RecA protein and the recombinational DNA repair of stalled replication forks. Annu. Rev. Biochem. 71, 71–100 (2002)
Kornberg, A. & Baker, T. A. DNA Replication 2nd edn 246 (W. H. Freeman & Co, 1992)
Breier, A. M., Weier, H. U. & Cozzarelli, N. R. Independence of replisomes in Escherichia coli chromosomal replication. Proc. Natl Acad. Sci. USA 102, 3942–3947 (2005)
Blattner, F. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1474 (1997)
Rocha, E. P. & Danchin, A. Essentiality, not expressiveness, drives gene-strand bias in bacteria. Nature Genet. 34, 377–378 (2003)
Rocha, E. P. & Danchin, A. Gene essentiality determines chromosome organisation in bacteria. Nucleic Acids Res. 31, 6570–6577 (2003)
Mirkin, E. V. & Mirkin, S. M. Mechanisms of transcription-replication collisions in bacteria. Mol. Cell. Biol. 25, 888–895 (2005)
Wang, J. D., Berkmen, M. B. & Grossman, A. D. Genome-wide coorientation of replication and transcription reduces adverse effects on replication in Bacillus subtilis. Proc. Natl Acad. Sci. USA 104, 5608–5613 (2007)
Prado, F. & Aguilera, A. Impairment of replication fork progression mediates RNA polII transcription-associated recombination. EMBO J. 24, 1267–1276 (2005)
Kobayashi, T. The replication fork barrier site forms a unique structure with Fob1p and inhibits the replication fork. Mol. Cell. Biol. 23, 9178–9188 (2003)
Deshpande, A. M. & Newlon, C. S. DNA replication fork pause sites dependent on transcription. Science 272, 1030–1033 (1996)
French, S. Consequences of replication fork movement through transcription units in vivo.. Science 258, 1362–1365 (1992)
Vilette, D., Ehrlich, S. D. & Michel, B. Transcription-induced deletions in plasmid vectors: M13 DNA replication as a source of instability. Mol. Gen. Genet. 252, 398–403 (1996)
Huvet, M. et al. Human gene organization driven by the coordination of replication and transcription. Genome Res. 17, 1278–1285 (2007)
Liu, B. & Alberts, B. M. Head-on collision between a DNA replication apparatus and RNA polymerase transcription complex. Science 267, 1131–1137 (1995)
Liu, B. et al. The DNA replication fork can pass RNA polymerase without displacing the nascent transcript. Nature 366, 33–39 (1993)
Liu, B., Wong, M. L. & Alberts, B. A transcribing RNA polymerase molecule survives DNA replication without aborting its growing RNA chain. Proc. Natl Acad. Sci. USA 91, 10660–10664 (1994)
Ogawa, T. & Okazaki, T. Discontinuous DNA replication. Annu. Rev. Biochem. 49, 421–457 (1980)
Wang, T. C. Discontinuous or semi-discontinuous DNA replication in Escherichia coli? Bioessays 27, 633–636 (2005)
Johnson, A. & O’Donnell, M. Cellular DNA replicases: components and dynamics at the replication fork. Annu. Rev. Biochem. 74, 283–315 (2005)
Pomerantz, R. T. & O’Donnell, M. Replisome mechanics: insights into a twin DNA polymerase machine. Trends Microbiol. 15, 156–164 (2007)
Steitz, T. A. The structural basis of the transition from initiation to elongation phases of transcription, as well as translocation and strand separation, by T7 RNA polymerase. Curr. Opin. Struct. Biol. 14, 4–9 (2004)
Mentesana, P. E., Chin-Bow, S. T., Sousa, R. & McAllister, W. T. Characterization of halted T7 RNA polymerase elongation complexes reveals multiple factors that contribute to stability. J. Mol. Biol. 302, 1049–1062 (2000)
Jiang, M., Rong, M., Martin, C. & McAllister, W. T. Interrupting the template strand of the T7 promoter facilitates translocation of the DNA during initiation, reducing transcript slippage and the release of abortive products. J. Mol. Biol. 310, 509–522 (2001)
Uptain, S. M., Kane, C. M. & Chamberlin, M. J. Basic mechanisms of transcript elongation and its regulation. Annu. Rev. Biochem. 66, 117–172 (1997)
Trautinger, B. W., Jaktaji, R. P., Rusakova, E. & Lloyd, R. G. RNA polymerase modulators and DNA repair activities resolve conflicts between DNA replication and transcription. Mol. Cell 19, 247–258 (2005)
Stukenberg, P. T., Turner, J. & O'Donnell, M. An explanation for lagging strand replication: polymerase hopping among DNA sliding clamps. Cell 78, 877–887 (1994)
Keiler, K. C. Biology of trans-translation. Annu. Rev. Microbiol. 62, 133–151 (2008)
Okazaki, R. et al. Mechanism of DNA chain growth. I. Possible discontinuity and unusual secondary structure of newly synthesized chains. Proc. Natl Acad. Sci. USA 59, 598–605 (1968)
Sternglanz, R., Wang, H. F. & Donegan, J. J. Evidence that both growing DNA chains at a replication fork are synthesized discontinuously. Biochemistry 15, 1838–1843 (1976)
Pauling, C. & Hamm, L. Properties of a temperature-sensitive, radiation-sensitive mutant of Escherichia coli. II. DNA replication. Proc. Natl Acad. Sci. USA 64, 1195–1202 (1969)
Gottesman, M. M., Hicks, M. L. & Gellert, M. Genetics and function of DNA ligase in Escherichia coli. J. Mol. Biol. 77, 531–547 (1973)
Konrad, E. B., Modrich, P. & Lehman, I. R. Genetic and enzymatic characterization of a conditional lethal mutant of Escherichia coli K12 with a temperature-sensitive DNA ligase. J. Mol. Biol. 77, 519–529 (1973)
Okazaki, R., Arisawa, M. & Sugino, A. Slow joining of newly replicated DNA chains in DNA polymerase I-deficient Escherichia coli mutants. Proc. Natl Acad. Sci. USA 68, 2954–2957 (1971)
Olivera, R. M. & Bonhoeffer, E. Replication of Escherichia coli requires DNA polymerase I. Nature 250, 513–514 (1974)
Wang, T. C. & Smith, K. C. Discontinuous DNA replication in a lig-7 strain of Escherichia coli is not the result of mismatch repair, nucleotide-excision repair, or the base-excision repair of DNA uracil. Biochem. Biophys. Res. Commun. 165, 685–688 (1989)
Wang, T. C. & Chen, S. H. Okazaki DNA fragments contain equal amounts of lagging-strand and leading-strand sequences. Biochem. Biophys. Res. Commun. 198, 844–849 (1994)
Heller, R. C. & Marians, K. J. Replication fork reactivation downstream of a blocked nascent leading strand. Nature 439, 557–562 (2006)
Heller, R. C. & Marians, K. J. Replisome assembly and the direct restart of stalled replication forks. Nature Rev. Mol. Cell Biol. 7, 932–943 (2006)
Heller, R. C. & Marians, K. J. The disposition of nascent strands at stalled replication forks dictates the pathway of replisome loading during restart. Mol. Cell 17, 733–743 (2005)
McInerney, P. & O’Donnell, M. Functional uncoupling of twin polymerases: mechanism of polymerase dissociation from a lagging-strand block. J. Biol. Chem. 279, 21543–21551 (2004)
Severinov, K., Mooney, R., Darst, S. A. & Landick, R. Tethering of the large subunits of Escherichia coli RNA polymerase. J. Biol. Chem. 272, 24137–24140 (1997)
Severinov, K., Mooney, R., Darst, S. A. & Landick, R. Tethering of the large subunits of Escherichia coli RNA polymerase. J. Biol. Chem. 272, 24137–24140 (1997)
Acknowledgements
We are grateful to W. T. McAllister and R. Castagna for providing T7 RNAP, and to S. Darst and L. Westblade for providing E. coli RNAP proteins and plasmids. This work was supported by a grant from the National Institutes of Health (M.O.D.) and by a Marie-Josee and Henry Kravis Fellowship at the Rockefeller University (R.T.P.).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-5 with Legends. (PDF 1485 kb)
Rights and permissions
About this article
Cite this article
Pomerantz, R., O’Donnell, M. The replisome uses mRNA as a primer after colliding with RNA polymerase. Nature 456, 762–766 (2008). https://doi.org/10.1038/nature07527
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07527
This article is cited by
-
The in vivo measurement of replication fork velocity and pausing by lag-time analysis
Nature Communications (2023)
-
The human nucleoporin Tpr protects cells from RNA-mediated replication stress
Nature Communications (2021)
-
The prevention and resolution of DNA replication–transcription conflicts in eukaryotic cells
Genome Instability & Disease (2020)
-
Genomic methods for measuring DNA replication dynamics
Chromosome Research (2020)
-
DksA and DNA double-strand break repair
Current Genetics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.