Abstract
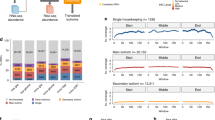
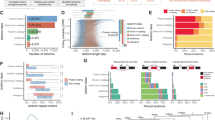
Through alternative processing of pre-messenger RNAs, individual mammalian genes often produce multiple mRNA and protein isoforms that may have related, distinct or even opposing functions. Here we report an in-depth analysis of 15 diverse human tissue and cell line transcriptomes on the basis of deep sequencing of complementary DNA fragments, yielding a digital inventory of gene and mRNA isoform expression. Analyses in which sequence reads are mapped to exon–exon junctions indicated that 92–94% of human genes undergo alternative splicing, ∼86% with a minor isoform frequency of 15% or more. Differences in isoform-specific read densities indicated that most alternative splicing and alternative cleavage and polyadenylation events vary between tissues, whereas variation between individuals was approximately twofold to threefold less common. Extreme or ‘switch-like’ regulation of splicing between tissues was associated with increased sequence conservation in regulatory regions and with generation of full-length open reading frames. Patterns of alternative splicing and alternative cleavage and polyadenylation were strongly correlated across tissues, suggesting coordinated regulation of these processes, and sequence conservation of a subset of known regulatory motifs in both alternative introns and 3′ untranslated regions suggested common involvement of specific factors in tissue-level regulation of both splicing and polyadenylation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
The reported sequence read data have been deposited to the Short Read Archive section of GEO at NCBI under accession numbers GSE12946 and SRA002355.1.
Change history
27 November 2008
The AOP version of this paper contained a typo in the legend for Figure 1. This was corrected for print on 27 November 2008.
References
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003)
Matlin, A. J., Clark, F. & Smith, C. W. Understanding alternative splicing: towards a cellular code. Nature Rev. Mol. Cell Biol. 6, 386–398 (2005)
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
Johnson, J. M. et al. Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science 302, 2141–2144 (2003)
Blencowe, B. J. Alternative splicing: new insights from global analyses. Cell 126, 37–47 (2006)
Xu, Q., Modrek, B. & Lee, C. Genome-wide detection of tissue-specific alternative splicing in the human transcriptome. Nucleic Acids Res. 30, 3754–3766 (2002)
Gupta, S., Zink, D., Korn, B., Vingron, M. & Haas, S. A. Strengths and weaknesses of EST-based prediction of tissue-specific alternative splicing. BMC Genomics 5, 72 (2004)
Yeo, G., Holste, D., Kreiman, G. & Burge, C. B. Variation in alternative splicing across human tissues. Genome Biol. 5, R74 (2004)
Sugnet, C. W. et al. Unusual intron conservation near tissue-regulated exons found by splicing microarrays. PLOS Comput. Biol. 2, e4 (2006)
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008)
Sultan, M. et al. A global view of gene activity and alternative splicing by deep sequencing of the human transcriptome. Science 321, 956–960 (2008)
Ule, J. et al. An RNA map predicting Nova-dependent splicing regulation. Nature 444, 580–586 (2006)
Wang, Z., Xiao, X., Van Nostrand, E. & Burge, C. B. General and specific functions of exonic splicing silencers in splicing control. Mol. Cell 23, 61–70 (2006)
Shi, L. et al. The MicroArray Quality Control (MAQC) project shows inter- and intraplatform reproducibility of gene expression measurements. Nature Biotechnol. 24, 1151–1161 (2006)
Mayr, J. A. et al. Mitochondrial phosphate-carrier deficiency: a novel disorder of oxidative phosphorylation. Am. J. Hum. Genet. 80, 478–484 (2007)
Graveley, B. R. The haplo-spliceo-transcriptome: common variations in alternative splicing in the human population. Trends Genet. 24, 5–7 (2008)
Nembaware, V., Wolfe, K. H., Bettoni, F., Kelso, J. & Seoighe, C. Allele-specific transcript isoforms in human. FEBS Lett. 577, 233–238 (2004)
Pan, Q. et al. Revealing global regulatory features of mammalian alternative splicing using a quantitative microarray platform. Mol. Cell 16, 929–941 (2004)
Xing, Y. & Lee, C. J. Protein modularity of alternatively spliced exons is associated with tissue-specific regulation of alternative splicing. PLoS Genet. 1, e34 (2005)
Kwan, T. et al. Genome-wide analysis of transcript isoform variation in humans. Nature Genet. 40, 225–231 (2008)
Lewis, B. P., Green, R. E. & Brenner, S. E. Evidence for the widespread coupling of alternative splicing and nonsense-mediated mRNA decay in humans. Proc. Natl Acad. Sci. USA 100, 189–192 (2003)
Underwood, J. G., Boutz, P. L., Dougherty, J. D., Stoilov, P. & Black, D. L. Homologues of the Caenorhabditis elegans Fox-1 protein are neuronal splicing regulators in mammals. Mol. Cell. Biol. 25, 10005–10016 (2005)
Auweter, S. D. et al. Molecular basis of RNA recognition by the human alternative splicing factor Fox-1. EMBO J. 25, 163–173 (2006)
Nakahata, S. & Kawamoto, S. Tissue-dependent isoforms of mammalian Fox-1 homologs are associated with tissue-specific splicing activities. Nucleic Acids Res. 33, 2078–2089 (2005)
Oberstrass, F. C. et al. Structure of PTB bound to RNA: specific binding and implications for splicing regulation. Science 309, 2054–2057 (2005)
Lewis, B. P., Burge, C. B. & Bartel, D. P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005)
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005)
Majoros, W. H. & Ohler, U. Spatial preferences of microRNA targets in 3′ untranslated regions. BMC Genomics 8, 152 (2007)
Maniatis, T. & Reed, R. An extensive network of coupling among gene expression machines. Nature 416, 499–506 (2002)
McCracken, S., Lambermon, M. & Blencowe, B. J. SRm160 splicing coactivator promotes transcript 3′-end cleavage. Mol. Cell. Biol. 22, 148–160 (2002)
Castelo-Branco, P. et al. Polypyrimidine tract binding protein modulates efficiency of polyadenylation. Mol. Cell. Biol. 24, 4174–4183 (2004)
Zhang, L., Lee, J. E., Wilusz, J. & Wilusz, C. J. The RNA-binding protein CUGBP1 regulates stability of tumor necrosis factor mRNA in muscle cells: implications for myotonic dystrophy. J. Biol. Chem. 283, 22457–22463 (2008)
Ladd, A. N. & Cooper, T. A. Finding signals that regulate alternative splicing in the post-genomic era. Genome Biol. 3, reviews0008.1–reviews0008.16 (2002)
Licatalosi, D. et al. Mechanisms of alternative mRNA processing in the brain revealed by HITS-CLIP. Nature doi: 10.1038/nature07488 (this issue)
Galarneau, A. & Richard, S. Target RNA motif and target mRNAs of the Quaking STAR protein. Nature Struct. Mol. Biol. 12, 691–698 (2005)
Kim, H. H. & Gorospe, M. GU-rich RNA: expanding CUGBP1 function, broadening mRNA turnover. Mol. Cell 29, 151–152 (2008)
Wu, J. I., Reed, R. B., Grabowski, P. J. & Artzt, K. Function of quaking in myelination: regulation of alternative splicing. Proc. Natl Acad. Sci. USA 99, 4233–4238 (2002)
Paz, R. D. et al. Increased expression of activity-dependent genes in cerebellar glutamatergic neurons of patients with schizophrenia. Am. J. Psychiatry 163, 1829–1831 (2006)
Elenbaas, B. et al. Human breast cancer cells generated by oncogenic transformation of primary mammary epithelial cells. Genes Dev. 15, 50–65 (2001)
Schroth, G. P., Luo, S. & Khrebtukova, I. Transcriptome analysis using high-throughput DNA sequencing. Methods Mol. Biol. (in the press)
Illumina, Inc. Transcriptome Analysis: mRNA-Seq. 〈http://www.illumina.com/pages.ilmn?ID=291〉 (2008)
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human genome by next generation sequencing. Nature Genet. (in the press)
Acknowledgements
We thank E. Anderson, D. Black, B. Friedman, and members of the Burge laboratory for comments on the manuscript, N. Spies for analyses, J. Mudge, G. D. May, N. A. Miller, E. Vermaas, T. Kerelska, J. Yan and V. Quijano for assistance in generating the mRNA-Seq data, and R. C. Roberts and N. Perrone-Bizzozero for supplying cerebellar cortex RNA samples. This research was supported by an NIH training grant (E.T.W.), and by grants from the Knut & Alice Wallenberg Foundation and the Swedish Foundation for Strategic Research (R.S.) and from the NIH (C.B.B.).
Author Contributions E.W. and R.S. designed and performed the computational analyses of sequencing reads, prepared figures, tables and methods and contributed to manuscript text. S.L. developed protocols and created libraries, L.Z. contributed to sequencing development, and I.K., S.L. and L.Z. did primary data analysis. G.P.S. contributed to study design and manuscript preparation. C.M. and S.F.K. provided RNA samples and contributed to manuscript preparation. C.B.B. designed the study and prepared the manuscript, with input from other authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
S.L., I.K., L.Z. and G.P.S. are employees of Illumina, Inc.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary References, Supplementary Figures S1-S9 with Legends and Supplementary Tables S1-S9. (PDF 9256 kb)
Rights and permissions
About this article
Cite this article
Wang, E., Sandberg, R., Luo, S. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008). https://doi.org/10.1038/nature07509
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07509
This article is cited by
-
A promising target for breast cancer: B7-H3
BMC Cancer (2024)
-
ClusTrast: a short read de novo transcript isoform assembler guided by clustered contigs
BMC Bioinformatics (2024)
-
Deciphering the role of alternative splicing as a potential regulator in fat-tail development of sheep: a comprehensive RNA-seq based study
Scientific Reports (2024)
-
Revisiting the development of cerebellar inhibitory interneurons in the light of single-cell genetic analyses
Histochemistry and Cell Biology (2024)
-
ASAPA: a bioinformatic pipeline based on Iso-Seq that identifies the links among alternative splicing, alternative transcription initiation and alternative polyadenylation
Functional & Integrative Genomics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.