Abstract
Crosstalk between the oestrogen receptor (ER) and ERBB2/HER-2 pathways has long been implicated in breast cancer aetiology and drug response1, yet no direct connection at a transcriptional level has been shown. Here we show that oestrogen–ER and tamoxifen–ER complexes directly repress ERBB2 transcription by means of a cis-regulatory element within the ERBB2 gene in human cell lines. We implicate the paired box 2 gene product (PAX2), in a previously unrecognized role, as a crucial mediator of ER repression of ERBB2 by the anti-cancer drug tamoxifen. We show that PAX2 and the ER co-activator AIB-1/SRC-3 compete for binding and regulation of ERBB2 transcription, the outcome of which determines tamoxifen response in breast cancer cells. The repression of ERBB2 by ER-PAX2 links these two breast cancer subtypes and suggests that aggressive ERBB2-positive tumours can originate from ER-positive luminal tumours by circumventing this repressive mechanism. These data provide mechanistic insight into the molecular basis of endocrine resistance in breast cancer.
Similar content being viewed by others
Main
The genomic mapping of ER-binding sites has provided insight into how ER functions in breast cancer cells, including the finding that ER rarely binds to promoter regions and that loading of ER on the chromatin requires the presence of pioneer factors, such as FoxA1 (refs 2–4). We have replicated genome-wide ER chromatin immunoprecipitation (ChIP)-on-chip analyses in ER-positive MCF-7 cells. Identification of the ER-binding sites with a false discovery rate of 5% revealed 8,525 ER sites, with high representation (86%) of the published ER binding profile2 (Supplementary Data 2). Included within the new, more extensive list was an ER-binding site within the intron of the ERBB2/HER-2 genomic region (Fig. 1a). Sequence analysis of all 8,525 ER-binding sites revealed a statistical enrichment (P < 0.0001) for the PAX transcription factor motif (GTCANGN(A/G)T) (Fig. 1b). Little is known about the function of PAX proteins in hormone signalling; however, PAX2 was shown to be expressed in a subset of breast cancers and was recently identified as a tamoxifen-regulated effector in endometrial cancer cells5,6.
a, Schematic representation of the ERBB2 gene locus and the intronic ER-binding site, as defined by ER ChIP-on-chip experiments, on chromosome 17. Both isoforms of ERBB2 are shown. b, PAX motif enriched within ER-binding sites. c, PAX2 ChIP after treatment with vehicle (blue bars), oestrogen (yellow bars) or tamoxifen (green bars). d, Changes in ERBB2 mRNA by real-time RT–PCR after treatment with vehicle (blue), oestrogen (yellow) or tamoxifen (green). All graphical results are shown as means and s.d. for three independent replicates.
Tamoxifen is one of the most successful and effective therapies in the treatment of breast cancer, but resistance to tamoxifen is common7. Tamoxifen-resistant breast tumours are characterized by elevated ERBB2 levels8, and ER-positive cell line models overexpressing ERBB2 acquire resistance to tamoxifen9. We assessed the binding of PAX2 to selected ER-binding sites adjacent to important oestrogen-regulated genes, including the newly identified binding site within the ERBB2 gene. PAX2 was generally recruited only after treatment with tamoxifen, with the exception of the ER-binding site within ERBB2 (Fig. 1c), where PAX2 was recruited to the ER-binding site after both treatment with oestrogen and treatment with tamoxifen. Given previous evidence that ERBB2 could be repressed by both oestrogen10 and tamoxifen11, we proposed that PAX2 might be functioning as a general ER-associated transcriptional repressor and that the ER-binding site within ERBB2 might be a cis-regulatory element for active repression by ER. Indeed, analysis confirmed that levels of ERBB2 messenger RNA are decreased by oestrogen and by tamoxifen in our MCF-7 cells (Fig. 1d). Co-immunoprecipitation experiments showed that ER and PAX2 form a complex after treatment with tamoxifen (Supplementary Data 3) and re-ChIP experiments (ChIP followed by release of chromatin and re-ChIP using an antibody against a different protein) confirmed that ER and PAX2 occupy the same ER-binding site within the ERBB2 gene simultaneously, after treatment with tamoxifen (Supplementary Data 3). Furthermore, we experimentally verified this ER-binding site as the cis-regulatory element for the ERBB2 gene (Supplementary Data 4). This cis-regulatory region is independent of a previously identified regulatory region10, although this previously characterized region might have an indirect function in the regulation of ERBB2 transcription.
ER-positive luminal tumours with the poorest prognosis tend to have elevated ERBB2 levels12 and up to half of ERBB2-positive tumours are also positive for ER13. We therefore proposed that the anti-proliferative effects of tamoxifen treatment require repression of ERBB2, and that breast cancers can potentially acquire tamoxifen resistance by amplifying the ERBB2 locus or by deregulating the control mechanisms that normally repress ERBB2 transcription. Unlike tamoxifen, repression of ERBB2 by oestrogen may not be a critical event, because cell proliferation by oestrogen probably results from the oestrogen-mediated upregulation of numerous oncogenes.
To investigate the possible role for PAX2 in the oestrogen-mediated and tamoxifen-mediated repression of ERBB2, we specifically silenced PAX2 with short interfering RNA (siRNA). Immunoblotting revealed efficient knockdown of PAX2 protein levels, but no significant effect on ER levels (Fig. 2a). In control transfected cells, oestrogen and tamoxifen both rapidly repressed ERBB2 mRNA (Fig. 2b), but PAX2 siRNA abrogated this inhibition and consequently ERBB2 transcription and ERBB2 protein levels were elevated in the presence of both treatment with oestrogen and treatment with tamoxifen (Figs 2a and 2b). This coincided with an accumulation of phosphorylated RNA polymerase II (PolII) at the promoter (the longer isoform) of ERBB2 after treatment with oestrogen and with tamoxifen in the presence of PAX2 siRNA (Supplementary Data 5). Relative to the control, treatment of PAX2 siRNA-transfected cells with tamoxifen resulted in an increase in cell number (Fig. 2c), reversing the growth arrest observed after treatment with tamoxifen. These experiments were reproduced with an additional PAX2 siRNA (Supplementary Data 6). Pretreatment of cells with an anti-ERBB2 antibody (Herceptin) blocked the PAX2 siRNA-mediated cell growth, confirming that the increased cell growth after PAX2 silencing was due primarily to the increase in ERBB2 levels (Fig. 2c).
a, siRNA to PAX2 was transfected into hormone-depleted MCF-7 cells and stimulated with vehicle (V), oestrogen (O) or tamoxifen (T), and total protein was immunoblotted. b, Control siRNA (siLuc; white bars) or PAX2 siRNA (grey bars) was transfected and ERBB2 mRNA levels were assessed. c, PAX2 siRNA was transfected into cells in the presence of control (black bars) or Herceptin (anti-ERBB2 antibody; white bars), after which cells were collected and the total number of viable cells was determined. The control transfection (siLuc) data are in Supplementary Fig. 7. d, Transfection of control siRNA (siLuc) or PAX2 siRNA was performed as described and cells were treated with vehicle (blue bars), oestrogen (yellow bars) or tamoxifen (green bars). Binding of AIB-1 to the ERBB2 enhancer was determined by ChIP. All graphical results are shown as means and s.d. for three independent replicates.
We recapitulated these findings in another ER-positive breast cancer cell line (ZR75-1 cells). Tamoxifen repressed ERBB2 mRNA and ERBB2 protein levels in ZR75-1 cells, and this repression was inhibited after silencing of PAX2. Similarly to MCF-7 cells, PAX2 siRNA reversed the growth inhibitory effects of tamoxifen, such that ZR75-1 cells acquired tamoxifen resistance in the absence of PAX2 (Supplementary Data 8).
As well as having elevated ERBB2 levels, tamoxifen-resistant breast cancers are also characterized by increased levels of the ER co-activator AIB-1 (amplified in breast cancer-1; also known as SRC-3 (ref. 8)). AIB-1 promotes tumorigenesis14,15 and is essential for ERBB2-driven oncogenesis in mice16. We assessed whether AIB-1 could compete with PAX2 for binding to the ERBB2 cis-regulatory element, an event that may contribute to the elevated ERBB2 levels associated with tamoxifen-resistant tumours8. ChIP showed decreased AIB-1 binding at the ERBB2 cis-regulatory element after treatment with oestrogen and treatment with tamoxifen (Fig. 2d), probably as a result of displacement by PAX2. This was proved by inhibiting PAX2 with siRNA, which consequently allowed oestrogen-mediated and tamoxifen-mediated recruitment of AIB-1 to the ERBB2 cis-regulatory element (Fig. 2d).
We subsequently showed that expression of AIB-1 competes with PAX2 for binding to the ERBB2 cis-regulatory element and that this results in an increase in ERBB2 transcription and an increase in cell proliferation in the presence of tamoxifen (Supplementary Data 9). Elevated AIB-1 levels block PAX2 binding and ERBB2 gene repression, thereby reversing the antiproliferative effects of tamoxifen. This suggests that a stoichiometric balance between the co-activator AIB-1 and the putative repressor PAX2 impinges on the binding and regulation of ERBB2, providing mechanistic insight into the function of AIB-1 in the tamoxifen response17. MCF-7 cells already have elevated AIB-1 levels as a result of a genomic amplification of the AIB-1 locus14, but they also have increased PAX2 protein levels (data not shown), potentially explaining why they retain sensitivity to treatment with tamoxifen. However, we were also able to show that AIB-1 expression could reverse the anti-proliferative effects of tamoxifen in T47D cells, a cell line that does not already have elevated AIB-1 levels14 (Supplementary Data 9). These data confirm a general role for AIB-1 in reversing tamoxifen responsiveness in ER-positive breast cancer cell lines.
The role of ERBB2 in tamoxifen resistance is demonstrated by data showing that tamoxifen-resistant breast cancer cell lines can be inhibited by treatment with anti-ERBB2 antibodies (Herceptin)18. We investigated the hypothesis that PAX2 is required for repression of ERBB2 and that tamoxifen resistance may be due to alterations in this pathway. We used an MCF-7 cell line model that had been grown in the presence of tamoxifen and had acquired resistance18. These tamoxifen-resistant (Tam-R) cells have elevated ERBB2 levels but do not have amplification of the ERBB2 locus18. In wild-type MCF-7 cells, tamoxifen repressed ERBB2 mRNA levels (Fig. 2b) and ERBB2 protein levels (by 40%) (Fig. 3a) as expected, but ERBB2 levels were elevated in Tam-R cells and did not decrease in response to treatment with tamoxifen (Fig. 3a). Western blot analysis comparing wild-type and Tam-R MCF-7 cells revealed no changes in ER protein levels, supporting clinical studies showing that changes in ER levels are not a general mechanism for tamoxifen-resistant breast cancers19,20. AIB-1 protein levels were also unaltered, but PAX2 protein levels were lower in Tam-R cells (Fig. 3a), providing a potential explanation for the elevated ERBB2 levels in these tamoxifen-resistant cells.
a, Total protein from wild-type or tamoxifen-resistant (Tam-R) MCF-7 cells was immunoblotted. V, vehicle; T, tamoxifen. b, ChIP of ER, PAX2, AIB-1 and HDAC-1 at the ER-binding site in the ERBB2 gene in wild-type and Tam-R cells after treatment with vehicle (blue bars) or tamoxifen (green bars). c, Control or PAX2-expressing plasmids were transfected into Tam-R cells, followed by treatment with vehicle (V) or tamoxifen (T). Total protein was immunoblotted. After PAX2 expression in Tam-R cells, total numbers of viable cells were determined after treatment with vehicle or tamoxifen. The data are tamoxifen treatment versus vehicle. Green, control; blue, PAX2. The immunoblots have been cropped; the original figures are in Supplementary Data 11. All graphical results are shown as means and s.d. for three independent replicates.
Tamoxifen-mediated ER recruitment to the ERBB2 cis-regulatory element was assessed in the Tam-R cells and was shown to be similar to that in wild-type MCF-7 cells (Fig. 3b). However, as suspected from the lower PAX2 levels in Tam-R cells, PAX2 binding was significantly decreased in the Tam-R cells. Similarly, binding of histone deacetylase 1 (HDAC-1) was shown to occur only in the wild-type cells and not in the Tam-R cells, confirming that active repression occurs at the ERBB2 cis-regulatory element in wild-type cells but not in the Tam-R cells. In contrast, tamoxifen-mediated AIB-1 recruitment was elevated in Tam-R cells (Fig. 3b), despite unaltered AIB-1 levels. To test the hypothesis that the decreased PAX2 levels contributed to the increased expression of ERBB2 and the altered response to tamoxifen in the Tam-R cells, we reintroduced PAX2 into these cells (Fig. 3c). After overexpression of PAX2, tamoxifen was now able to repress ERBB2 mRNA (Supplementary Data 12) and ERBB2 protein levels (Fig. 3c) in Tam-R cells. The overexpression of PAX2 resulted in decreased binding of PolII to the ERBB2 promoter and decreased binding of AIB-1 to the ER-binding site (Supplementary Data 12), strengthening the hypothesis that AIB-1 and PAX2 compete for binding and regulation of the ERBB2 gene. Active gene repression by tamoxifen was restored by PAX2 expression, as indicated by the recruitment of HDAC-1 (Supplementary Data 12). PAX2 was subsequently shown to be a critical regulator of cellular proliferation, because expression of PAX2 restored the ability of tamoxifen to inhibit cell growth in these previously resistant cells (Fig. 3c).
We recapitulated these findings in BT-474 breast cancer cells, which are ER positive but resistant to tamoxifen21, probably as a result of a genomic amplification of the ERBB2 locus22. These cells therefore represent another possible mechanism of acquired tamoxifen resistance, whereby amplification of the ERBB2 locus can overcome the growth inhibitory effects imposed by tamoxifen in ER-positive breast cancers23,24. Expression of PAX2 in BT-474 cells resulted in tamoxifen-mediated repression of ERBB2 mRNA and ERBB2 protein levels (Supplementary Data 13) and resulted in tamoxifen-dependent inhibition of cell growth (Supplementary Data 13), such that growth inhibitory effects of tamoxifen were restored, even in the presence of the amplified ERBB2 locus.
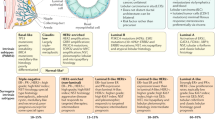
Our findings suggest that PAX2 is a key deterministic component in the tamoxifen response. To confirm these findings in primary breast cancer, we performed PAX2 immunohistochemistry on 109 ER-positive breast cancer samples25, all of which had been treated with tamoxifen. Of these 109 tumours, 68 were PAX2 positive and 41 were PAX2 negative. Tumours with positive PAX2 staining corresponded to a significantly improved recurrence-free survival in patients relative to PAX2-negative tumours (P < 0.0001) (Supplementary Data 14). Furthermore, within the PAX2-positive tumours only, those that were also positive for AIB-1 had a worse clinical outcome than the tumours that were AIB-1 negative (Fig. 4). The tumours that were PAX2 positive and AIB-1 negative had the best prognosis of all, with a recurrence rate of only 5.8% (Fig. 4). Cox regression analysis confirmed an inverse dependent relationship between PAX2 and AIB-1 levels in determining relapse (P < 0.03). The PAX2-positive, AIB-1-negative tumours also had the lowest percentage of ERBB2-positive staining (Fig. 4), supporting our hypothesis that a balance between PAX2 and AIB-1 ultimately dictates ERBB2 expression and determines tamoxifen efficacy.
Kaplan–Meier curve representing the percentage relapse-free survival in tumours based on staining for PAX2 and AIB-1 (n = 109). The percentage of ERBB2-overexpressing tumours within each category is shown.
Endocrine resistance is a significant problem in breast cancer treatment. One of the few validated clinical features of tamoxifen-resistant breast cancer is the combined elevation of the AIB-1 and ERBB2 pathways8. We now provide evidence that PAX2 is a critical tamoxifen-recruited transcriptional repressor of the ERBB2 gene and that increased AIB-1 expression can outcompete PAX2 binding, directly resulting in increased ERBB2 expression. Alterations in AIB-1–PAX2 stoichiometry dictate the efficacy of tamoxifen in breast cancers (a schematic model of these events is shown in Supplementary Data 1). These new data suggest an intrinsic transcriptional link between tumours driven by ER and those driven by ERBB2, which together account for a significant majority of all breast cancers. The role of PAX2 as a repressor is unexpected, because PAX2 is generally a transcriptional activator and was shown to be encoded by a tamoxifen-regulated gene that can induce endometrial cancer6. Given that tamoxifen has antiproliferative effects in the breast but possesses agonist properties in the endometrium26, it is possible that PAX2 may have tissue specific effects and may be one of the primary determinants for selective oestrogen receptor modulator (SERM) action in female reproductive tissue.
Methods Summary
The MCF-7, T47D, ZR75-1 and BT-474 human cell lines were grown as described previously27. Tam-R cells were derived by long-term exposure to tamoxifen18 and grown under the same conditions as wild-type MCF-7 cells. Genome-wide ER ChIP-on-chip experiments were performed in MCF-7 cells in duplicate, as described previously2. ER-binding-site analyses were determined with MAT28, with a false discovery rate of 5%. The analysis of motif enrichments was performed with the CEAS program (http://ceas.cbi.pku.edu.cn/). For mRNA experiments, cells were deprived of hormones as described previously29. Total RNA was collected and RT–PCR was performed as described previously2. ChIP experiments were performed in hormone-depleted medium as described previously3. Proliferation assays were performed in complete medium. siRNA experiments were performed as described previously2 in hormone-depleted medium. For immunohistochemistry, 109 ER-positive breast cancer sections were collected and processed as described previously25. Immunohistochemistry for PAX2 was performed on an automated BondMax Immunostainer (Leica) with anti-PAX2 antibody (ab38738; Abcam) and anti-AIB-1 antibody (611105; Transduction Laboratories). Immunohistochemistry for ERBB2 was performed as described previously25. Statistical analysis was performed with two-tailed paired t-tests, and P < 0.05 was considered statistically significant. In all figures, graphical results are shown as means and s.d. for a minimum of three independent replicates. Kaplan–Meier curve statistics were determined with a log-rank test. PAX2 and AIB-1 relationships were established by Cox regression analysis.
Online Methods
Cell lines
The MCF-7, T47D, ZR75-1 and BT-474 human cell lines were grown as described previously27. Tam-R cells were derived by long-term exposure to tamoxifen18 and grown under the same conditions as wild-type MCF-7 cells. Herceptin was added to the medium at a final concentration of 10 μM.
ChIP-on-chip experiments
Genome-wide ER ChIP-on-chip experiments were performed in duplicate, as described previously2, with the exception that the Affymetrix seven Genechip tiling array 2.0R set was used. Analysis of ER-binding sites were determined with MAT28, with a false discovery rate of 5%. Microarray data are available in the ArrayExpress database (www.ebi.ac.uk/arrayexpress) under accession number E-TABM-563.
Motif enrichment
Analysis of motif enrichments was performed with the CEAS program (http://ceas.cbi.pku.edu.cn/). The PAX motif is represented with Weblogo (http://weblogo.berkeley.edu/).
Plasmids
PAX2 expression was from p-TARGET-PAX2 (a gift from S. Buttiglieri), AIB-1 expression was from a pcDNA-AIB-1 construct (a gift from J. Eeckhoute) and SRC-1 expression was from pSG5-SRC-1.
RT–PCR
Cells were deprived of hormones as described previously29. Total RNA was collected and RT–PCR was performed as described previously2. Primer sequences are provided in Supplementary Data 15.
ChIP
ChIP experiments were performed as described previously3. Antibodies used were anti-ERα (HC-20), anti-AIB-1/RAC3 (M-397) and anti-HDAC-1 (sc-6298 and sc-6299) from Santa Cruz Biotechnologies, and anti-PolII (ab5408), anti-H3R17 dimethyl (ab8284), anti-PAX2 (ab23799), anti-SRC-1 (ab2859) and anti-N-CoR (ab24552) from Abcam. Primer sequences are provided in Supplementary Data 15.
siRNA
siRNA experiments were performed as described previously2. The sequence of the siRNAs were as follows: PAX2 siRNA (sequence 1), 5′-GAAGUCAAGUCGAGUCUAUUU-3′ (sense) and 5′-AUAGACUCGACUUGACUUCUU-3′ (antisense); PAX2 siRNA (sequence 2), 5′-CAUCAGAGCACAUCAAAUCUU-3′ (sense) and 5′-GAUUUGAUGUGCUCUGAUGUU-3′ (antisense) (Dharmacon, USA); siN-CoR Smartpool (Dharmacon), containing the sequences 5′-GAUCACAUCUGUCAAAUUAUU-3′, 5′-GAACGUGGCUCUCAAAGUUU-3′, 5′-GAAAGGAAAUCGACACUGAUU-3′and 5′-GCCCUGGGAUUUAUGAUGAUU-3′.
Western blotting
Cells were deprived of hormones as described previously29. Antibodies used were anti-ERα (Ab-10) from Neomarkers (Lab Vision, UK); anti-ERBB2 (ab16901), anti-PAX2 (ab38738), anti-SRC-1 (ab2859), anti-N-CoR (ab24552) and anti-β-actin (ab6276) from Abcam; and anti-AIB-1/RAC3 (M-397) from Santa Cruz Biotechnologies.
Cell counting
Cells were plated at equal confluence and grown in full DMEM medium. Cells were transfected as described previously2 and cells were stimulated with 100 nM oestrogen or 1 μM 4-hydroxytamoxifen for 24 h, or in time-course experiments for the periods given in the figure. Total cells were harvested for automated cell counting using the Z2 Coulter Particle Count Analyzer.
Chromosome conformation capture (CCC) assay
CCC was performed in accordance with published protocols30. The chromatin was digested with PstI, and the real-time primers used were 5′-GGAGCGGAAGTGATTCAGAG-3′ (forward) and 5′-TTGCAGAGACCTCTGGGAGT-3′ (reverse). The TaqMan probe was 6-carboxy-fluorescein-5′-AGAGCAGTTCTGCTCTTCGC-3′. A control reverse primer against another PstI site was included, namely 5′-AGAGTCACCAGCCTCTGCAT-3′.
Immunohistochemistry
One hundred and nine ER-positive breast cancer sections were collected and processed as described previously25. Immunohistochemistry for PAX2 was performed on an automated BondMax Immunostainer (Leica) with anti-PAX2 antibody (ab38738; Abcam) at a dilution of 1:100. AIB-1 immunohistochemistry was performed with anti-AIB-1 antibody (611105; Transduction Laboratories) at 1:200. Immunohistochemistry for ERBB2 was described previously25. Examples of PAX2-positive and PAX2-negative stained samples are provided in Supplementary Data 16.
Statistical analyses
Analysis was performed in SPSS V16.0 for Mac (SPSS Inc.). Kaplan–Meier plots were constructed to display the data, and analysis was conducted with Cox regression. Time to relapse was taken as the outcome; binary variables for AIB-1 and PAX2 and the interaction between AIB-1 and PAX2 were included as predictor variables.
References
Ali, S. & Coombes, R. C. Endocrine-responsive breast cancer and strategies for combating resistance. Nature Rev. 2, 101–112 (2002)
Carroll, J. S. et al. Genome-wide analysis of estrogen receptor binding sites. Nature Genet. 38, 1289–1297 (2006)
Carroll, J. S. et al. Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein FoxA1. Cell 122, 33–43 (2005)
Lupien, M. et al. FoxA1 translates epigenetic signatures into enhancer-driven lineage-specific transcription. Cell 132, 958–970 (2008)
Muratovska, A. et al. Paired-Box genes are frequently expressed in cancer and often required for cancer cell survival. Oncogene 22, 7989–7997 (2003)
Wu, H. et al. Hypomethylation-linked activation of PAX2 mediates tamoxifen-stimulated endometrial carcinogenesis. Nature 438, 981–987 (2005)
Clarke, R. et al. Cellular and molecular pharmacology of antiestrogen action and resistance. Pharmacol. Rev. 53, 25–71 (2001)
Osborne, C. K. et al. Role of the estrogen receptor coactivator AIB1 (SRC-3) and HER-2/neu in tamoxifen resistance in breast cancer. J. Natl Cancer Inst. 95, 353–361 (2003)
Benz, C. C. et al. Estrogen-dependent, tamoxifen-resistant tumorigenic growth of MCF-7 cells transfected with HER2/neu. Breast Cancer Res. Treat. 24, 85–95 (1992)
Bates, N. P. & Hurst, H. C. An intron 1 enhancer element mediates oestrogen-induced suppression of ERBB2 expression. Oncogene 15, 473–481 (1997)
Frasor, J. et al. Selective estrogen receptor modulators: discrimination of agonistic versus antagonistic activities by gene expression profiling in breast cancer cells. Cancer Res. 64, 1522–1533 (2004)
Kun, Y. et al. Classifying the estrogen receptor status of breast cancers by expression profiles reveals a poor prognosis subpopulation exhibiting high expression of the ERBB2 receptor. Hum. Mol. Genet. 12, 3245–3258 (2003)
Piccart-Gebhart, M. J. et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N. Engl. J. Med. 353, 1659–1672 (2005)
Anzick, S. L. et al. AIB1, a steroid receptor coactivator amplified in breast and ovarian cancer. Science 277, 965–968 (1997)
Torres-Arzayus, M. I. et al. High tumor incidence and activation of the PI3K/AKT pathway in transgenic mice define AIB1 as an oncogene. Cancer Cell 6, 263–274 (2004)
Fereshteh, M. P. et al. The nuclear receptor coactivator amplified in breast cancer-1 is required for Neu (ErbB2/HER2) activation, signaling, and mammary tumorigenesis in mice. Cancer Res. 68, 3697–3706 (2008)
Louie, M. C. et al. ACTR/AIB1 functions as an E2F1 coactivator to promote breast cancer cell proliferation and antiestrogen resistance. Mol. Cell. Biol. 24, 5157–5171 (2004)
Knowlden, J. M. et al. Elevated levels of epidermal growth factor receptor/c-erbB2 heterodimers mediate an autocrine growth regulatory pathway in tamoxifen-resistant MCF-7 cells. Endocrinology 144, 1032–1044 (2003)
Johnston, S. R. et al. Changes in estrogen receptor, progesterone receptor, and pS2 expression in tamoxifen-resistant human breast cancer. Cancer Res. 55, 3331–3338 (1995)
Green, K. A. & Carroll, J. S. Oestrogen-receptor-mediated transcription and the influence of co-factors and chromatin state. Nature Rev. 7, 713–722 (2007)
Zhou, Y. et al. The NFκB pathway and endocrine-resistant breast cancer. Endocr. Relat. Cancer 12 (Suppl 1). S37–S46 (2005)
Clark, J. et al. Identification of amplified and expressed genes in breast cancer by comparative hybridization onto microarrays of randomly selected cDNA clones. Genes Chromosom. Cancer 34, 104–114 (2002)
Dowsett, M. Overexpression of HER-2 as a resistance mechanism to hormonal therapy for breast cancer. Endocr. Relat. Cancer 8, 191–195 (2001)
Dowsett, M. et al. HER-2 amplification impedes the antiproliferative effects of hormone therapy in estrogen receptor-positive primary breast cancer. Cancer Res. 61, 8452–8458 (2001)
Jiang, J. et al. Phosphorylation of estrogen receptor-α at Ser167 is indicative of longer disease-free and overall survival in breast cancer patients. Clin. Cancer Res. 13, 5769–5776 (2007)
Fisher, B. et al. Tamoxifen for the prevention of breast cancer: current status of the National Surgical Adjuvant Breast and Bowel Project P-1 study. J. Natl Cancer Inst. 97, 1652–1662 (2005)
Neve, R. M. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10, 515–527 (2006)
Johnson, W. E. et al. Model-based analysis of tiling-arrays for ChIP-chip. Proc. Natl Acad. Sci. USA 103, 12457–12462 (2006)
Carroll, J. S. et al. A pure estrogen antagonist inhibits cyclin E–Cdk2 activity in MCF-7 breast cancer cells and induces accumulation of p130–E2F4 complexes characteristic of quiescence. J. Biol. Chem. 275, 38221–38229 (2000)
Hagege, H. et al. Quantitative analysis of chromosome conformation capture assays (3C-qPCR). Nature Protocols 2, 1722–1733 (2007)
Acknowledgements
We thank D. Carroll, S.-F. Chin, I. Mills, C. Massie and P. Edwards for discussions; J. Mitchell for histology work; M. Eldridge, S. Vowler and B. Adryan for bioinformatic help; S. Buttiglieri for the p-TARGET-PAX2 construct; J. Eeckhoute for the pcDNA-AIB-1 construct; and M. Iddawela for the gift of Herceptin. This work was supported by grants from the National Institute of Diabetes and Digestive and Kidney Disease (R01DK074967 to M.B.) and the National Cancer Institute (P01CA8011105, and the DF/HCC Breast Cancer SPORE Grant to M.B.). We acknowledge support from the University of Cambridge, Cancer Research UK and Hutchison Whampoa Limited.
Author Contributions A.H. and J.S.C. conceived all experiments. A.H., K.A.H. and J.S.C. performed all experiments. T.R.G. and M.B. provided essential reagents and bioinformatics support for the ChIP-on-chip experiment. I.R.H. and R.I.N. provided essential cell line reagents. J.J. and S.A. collected primary breast cancer samples and performed immunohistochemistry, with help from W.J.H. A.H. and J.S.C. wrote the manuscript with advice from all authors.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Data and Supplementary Figures 1-16 with Legends. There is a corrigendum associated with the Supplementary Information of this paper. (PDF 4765 kb)
Supplementary Information
Supplementary Data file 1 contains the text version of a schematic model of oestrogen receptor-Pax2 repression of ERBB2. (TXT 273 kb)
Supplementary Information
This file contains the same information as nature07483-s2. However this bed file with the list of ER binding sites can be loaded into UCSC genome browser (http://genome.ucsc.edu/) using the March 2006 version of the human genome. (ZIP 95 kb)
Rights and permissions
About this article
Cite this article
Hurtado, A., Holmes, K., Geistlinger, T. et al. Regulation of ERBB2 by oestrogen receptor–PAX2 determines response to tamoxifen. Nature 456, 663–666 (2008). https://doi.org/10.1038/nature07483
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07483
This article is cited by
-
Comparison of tumors with HER2 overexpression versus HER2 amplification in HER2-positive breast cancer patients
BMC Cancer (2022)
-
Endocrine resistance in breast cancer: from molecular mechanisms to therapeutic strategies
Journal of Molecular Medicine (2021)
-
Epigenetic reprogramming at estrogen-receptor binding sites alters 3D chromatin landscape in endocrine-resistant breast cancer
Nature Communications (2020)
-
Combination of mTORC1/2 inhibitor vistusertib plus fulvestrant in vitro and in vivo targets oestrogen receptor-positive endocrine-resistant breast cancer
Breast Cancer Research (2019)
-
Temporal dynamic reorganization of 3D chromatin architecture in hormone-induced breast cancer and endocrine resistance
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.