Abstract
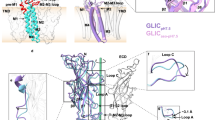
Pentameric ligand-gated ion channels from the Cys-loop family mediate fast chemo-electrical transduction1,2,3, but the mechanisms of ion permeation and gating of these membrane proteins remain elusive. Here we present the X-ray structure at 2.9 Å resolution of the bacterial Gloeobacter violaceus pentameric ligand-gated ion channel homologue4 (GLIC) at pH 4.6 in an apparently open conformation. This cationic channel is known to be permanently activated by protons5. The structure is arranged as a funnel-shaped transmembrane pore widely open on the outer side and lined by hydrophobic residues. On the inner side, a 5 Å constriction matches with rings of hydrophilic residues that are likely to contribute to the ionic selectivity6,7,8,9. Structural comparison with ELIC, a bacterial homologue from Erwinia chrysanthemi solved in a presumed closed conformation10, shows a wider pore where the narrow hydrophobic constriction found in ELIC is removed. Comparative analysis of GLIC and ELIC reveals, in concert, a rotation of each extracellular β-sandwich domain as a rigid body, interface rearrangements, and a reorganization of the transmembrane domain, involving a tilt of the M2 and M3 α-helices away from the pore axis. These data are consistent with a model of pore opening based on both quaternary twist and tertiary deformation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Corringer, P. J. & Changeux, J. P. Nicotinic acetylcholine receptors. Scholarpedia 3, 3468 (2008)
Lester, H. A., Dibas, M. I., Dahan, D. S., Leite, J. F. & Dougherty, D. A. Cys-loop receptors: New twists and turns. Trends Neurosci. 6, 329–336 (2004)
Sine, S. M. & Engel, A. G. Recent advances in Cys-loop receptor structure and function. Nature 440, 448–455 (2006)
Tasneem, A., Iyer, L. M., Jakobsson, E. & Aravind, L. Identification of the prokaryotic ligand-gated ion channels and their implications for the mechanisms and origins of animal Cys-loop ion channels. Genome Biol. 6, R4 (2005)
Bocquet, N. et al. A prokaryotic proton-gated ion channel from the nicotinic acetylcholine receptor family. Nature 445, 116–119 (2007)
Imoto, K. et al. Rings of negatively charged amino acids determine the acetylcholine receptor channel conductance. Nature 335, 645–648 (1988)
Imoto, K. et al. A ring of uncharged polar amino acids as a component of channel constriction in the nicotinic acetylcholine receptor. FEBS Lett. 289, 193–200 (1991)
Keramidas, A., Moorhouse, A. J., Schofield, P. R. & Barry, P. H. Ligand-gated ion channels: Mechanisms underlying ion selectivity. Prog. Biophys. Mol. Biol. 86, 161–204 (2004)
Corringer, P. J. et al. Mutational analysis of the charge selectivity filter of the α7 nicotinic acetylcholine receptor. Neuron 22, 831–843 (1999)
Hilf, R. J. & Dutzler, R. X-ray structure of a prokaryotic pentameric ligand-gated ion channel. Nature 452, 375–379 (2008)
Miyazawa, A., Fujiyoshi, Y. & Unwin, N. Structure and gating mechanism of the acetylcholine receptor pore. Nature 423, 949–955 (2003)
Unwin, N. Refined structure of the nicotinic acetylcholine receptor at 4 Å resolution. J. Mol. Biol. 346, 967–989 (2005)
Brejc, K. et al. Crystal structure of an ACh-binding protein reveals the ligand-binding domain of nicotinic receptors. Nature 411, 269–276 (2001)
Celie, P. H. et al. Nicotine and carbamylcholine binding to nicotinic acetylcholine receptors as studied in AChBP crystal structures. Neuron 41, 907–914 (2004)
Hansen, S. B. et al. Structures of Aplysia AChBP complexes with nicotinic agonists and antagonists reveal distinctive binding interfaces and conformations. EMBO J. 24, 3635–3646 (2005)
Dellisanti, C. D., Yao, Y., Stroud, J. C., Wang, Z. Z. & Chen, L. Crystal structure of the extracellular domain of nAChR α1 bound to α-bungarotoxin at 1.94 Å resolution. Nature Neurosci. 10, 953–962 (2007)
Giraudat, J., Dennis, M., Heidmann, T., Chang, J. Y. & Changeux, J. P. Structure of the high-affinity binding site for noncompetitive blockers of the acetylcholine receptor: Serine-262 of the delta subunit is labeled by [3H]chlorpromazine. Proc. Natl Acad. Sci. USA 83, 2719–2723 (1986)
Blanton, M. P. & Cohen, J. B. Identifying the lipid-protein interface of the Torpedo nicotinic acetylcholine receptor: Secondary structure implications. Biochemistry 33, 2859–2872 (1994)
Wilson, G. G. & Karlin, A. The location of the gate in the acetylcholine receptor channel. Neuron 20, 1269–1281 (1998)
Cymes, G. D., Ni, Y. & Grosman, C. Probing ion-channel pores one proton at a time. Nature 438, 975–980 (2005)
Wang, H. L., Cheng, X., Taylor, P., McCammon, J. A. & Sine, S. M. Control of cation permeation through the nicotinic receptor channel. PLOS Comput. Biol. 4, e41 (2008)
Beckstein, O. & Sansom, M. S. The influence of geometry, surface character, and flexibility on the permeation of ions and water through biological pores. Phys. Biol. 1, 42–52 (2004)
Harding, M. M. Metal-ligand geometry relevant to proteins and in proteins: Sodium and potassium. Acta Crystallogr. D 58, 872–874 (2002)
Jha, A., Cadugan, D. J., Purohit, P. & Auerbach, A. Acetylcholine receptor gating at extracellular transmembrane domain interface: The cys-loop and M2–M3 linker. J. Gen. Physiol. 130, 547–558 (2007)
Grutter, T. et al. Molecular tuning of fast gating in pentameric ligand-gated ion channels. Proc. Natl Acad. Sci. USA 102, 18207–18212 (2005)
Lee, W. Y. & Sine, S. M. Principal pathway coupling agonist binding to channel gating in nicotinic receptors. Nature 438, 243–247 (2005)
Lummis, S. C. et al. Cis-trans isomerization at a proline opens the pore of a neurotransmitter-gated ion channel. Nature 438, 248–252 (2005)
Taly, A. et al. Normal mode analysis suggests a quaternary twist model for the nicotinic receptor gating mechanism. Biophys. J. 88, 3954–3965 (2005)
Shimizu, H. et al. Global twisting motion of single molecular KcsA potassium channel upon gating. Cell 132, 67–78 (2008)
Li, G. D. et al. Identification of a GABAA receptor anesthetic binding site at subunit interfaces by photolabeling with an etomidate analog. J. Neurosci. 26, 11599–11605 (2006)
Acknowledgements
We thank P. Koehl for his program Aquasol and help with electrostatic calculations; P. Delepelaire and S. Edelstein for discussions; the staff of ESRF (Grenoble) ID14 and ID23 beamlines for data collection; facilities of the Pasteur Institute (A. Haouz for crystallogenesis, P. England for ultracentrifugation experiments, J. d’Alayer for mass spectroscopy controls and J. Bellalou for help in protein expression); and B. De Foresta (CEA, Orsay) for a gift of the two brominated DDM analogues. The latter diffraction data sets were collected at SLS and PSI (Villingen, Switzerland). We thank M. Fuchs for assistance during data collection; and the IDRIS supercomputer centre and its support staff for allocating CPU time at very short notice (project 082292). This work was supported by the Région Ile-de-France (N.B.), the Association Française contre les Myopathies, the Collège de France (C.L.P.), the Commission of the European Communities (Neurocypres project; H.N.) and the Network of European Neuroscience Institutes (ENI-NET).
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Tables S1- S2, Supplementary Figures S1-S8 and Supplementary References (PDF 2411 kb)
Supplementary Movie
The Supplementary Movie file illustrates the conformational dynamics of the system. The GLIC structure remains in an open conformation during the whole production run (20 ns), exhibiting thermal fluctuations around an average structure in its transmembrane part. The extracellular part is more mobile. (ZIP 8194 kb)
Rights and permissions
About this article
Cite this article
Bocquet, N., Nury, H., Baaden, M. et al. X-ray structure of a pentameric ligand-gated ion channel in an apparently open conformation. Nature 457, 111–114 (2009). https://doi.org/10.1038/nature07462
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07462
This article is cited by
-
A release of local subunit conformational heterogeneity underlies gating in a muscle nicotinic acetylcholine receptor
Nature Communications (2024)
-
Review: Nicotinic acetylcholine receptors to regulate important brain activity—what occurs at the molecular level?
Cognitive Neurodynamics (2023)
-
A lipid site shapes the agonist response of a pentameric ligand-gated ion channel
Nature Chemical Biology (2019)
-
The roles of aromatic residues in the glycine receptor transmembrane domain
BMC Neuroscience (2018)
-
PoreDesigner for tuning solute selectivity in a robust and highly permeable outer membrane pore
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.