Abstract
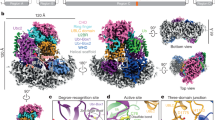
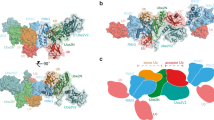
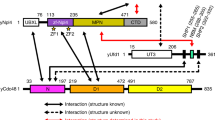
Deubiquitinating enzymes (DUBs) remove ubiquitin from conjugated substrates to regulate various cellular processes. The Zn2+-dependent DUBs AMSH and AMSH-LP regulate receptor trafficking by specifically cleaving Lys 63-linked polyubiquitin chains from internalized receptors. Here we report the crystal structures of the human AMSH-LP DUB domain alone and in complex with a Lys 63-linked di-ubiquitin at 1.2 Å and 1.6 Å resolutions, respectively. The AMSH-LP DUB domain consists of a Zn2+-coordinating catalytic core and two characteristic insertions, Ins-1 and Ins-2. The distal ubiquitin interacts with Ins-1 and the core, whereas the proximal ubiquitin interacts with Ins-2 and the core. The core and Ins-1 form a catalytic groove that accommodates the Lys 63 side chain of the proximal ubiquitin and the isopeptide-linked carboxy-terminal tail of the distal ubiquitin. This is the first reported structure of a DUB in complex with an isopeptide-linked ubiquitin chain, which reveals the mechanism for Lys 63-linkage-specific deubiquitination by AMSH family members.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Glickman, M. H. & Ciechanover, A. The ubiquitin–proteasome proteolytic pathway: destruction for the sake of construction. Physiol. Rev. 82, 373–428 (2002)
Hicke, L. Protein regulation by monoubiquitin. Nature Rev. Mol. Cell Biol. 2, 195–201 (2001)
Miranda, M. & Sorkin, A. Regulation of receptors and transporters by ubiquitination: new insights into surprisingly similar mechanisms. Mol. Interv. 7, 157–167 (2007)
Huang, F., Kirkpatrick, D., Jiang, X., Gygi, S. & Sorkin, A. Differential regulation of EGF receptor internalization and degradation by multiubiquitination within the kinase domain. Mol. Cell 21, 737–748 (2006)
Saksena, S., Sun, J., Chu, T. & Emr, S. D. ESCRTing proteins in the endocytic pathway. Trends Biochem. Sci. 32, 561–573 (2007)
Amerik, A. Y. & Hochstrasser, M. Mechanism and function of deubiquitinating enzymes. Biochim. Biophys. Acta 1695, 189–207 (2004)
Nijman, S. M. et al. A genomic and functional inventory of deubiquitinating enzymes. Cell 123, 773–786 (2005)
Komada, M. Controlling receptor downregulation by ubiquitination and deubiquitination. Curr. Drug Discov. Technol. 5, 78–84 (2008)
Clague, M. J. & Urbe, S. Endocytosis: the DUB version. Trends Cell Biol. 16, 551–559 (2006)
Mizuno, E. et al. Regulation of epidermal growth factor receptor down-regulation by UBPY-mediated deubiquitination at endosomes. Mol. Biol. Cell 16, 5163–5174 (2005)
McCullough, J., Clague, M. J. & Urbe, S. AMSH is an endosome-associated ubiquitin isopeptidase. J. Cell Biol. 166, 487–492 (2004)
Kim, M. S., Kim, J. A., Song, H. K. & Jeon, H. STAM–AMSH interaction facilitates the deubiquitination activity in the C-terminal AMSH. Biochem. Biophys. Res. Commun. 351, 612–618 (2006)
Hu, M. et al. Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell 111, 1041–1054 (2002)
Reyes-Turcu, F. E. et al. The ubiquitin binding domain ZnF UBP recognizes the C-terminal diglycine motif of unanchored ubiquitin. Cell 124, 1197–1208 (2006)
Misaghi, S. et al. Structure of the ubiquitin hydrolase UCH-L3 complexed with a suicide substrate. J. Biol. Chem. 280, 1512–1520 (2005)
Messick, T. E. et al. Structural basis for ubiquitin recognition by the OTU1 ovarian tumor domain protein. J. Biol. Chem. 283, 11038–11049 (2008)
Varadan, R., Assfalg, M., Raasi, S., Pickart, C. & Fushman, D. Structural determinants for selective recognition of a Lys48-linked polyubiquitin chain by a UBA domain. Mol. Cell 18, 687–698 (2005)
Trempe, J. F. et al. Mechanism of Lys48-linked polyubiquitin chain recognition by the Mud1 UBA domain. EMBO J. 24, 3178–3189 (2005)
Eddins, M. J., Carlile, C. M., Gomez, K. M., Pickart, C. M. & Wolberger, C. Mms2–Ubc13 covalently bound to ubiquitin reveals the structural basis of linkage-specific polyubiquitin chain formation. Nature Struct. Mol. Biol. 13, 915–920 (2006)
Pena, V., Liu, S., Bujnicki, J. M., Luhrmann, R. & Wahl, M. C. Structure of a multipartite protein–protein interaction domain in splicing factor Prp8 and its link to Retinitis pigmentosa . Mol. Cell 25, 615–624 (2007)
Zhang, L. et al. Crystal structure of the C-terminal domain of splicing factor Prp8 carrying Retinitis pigmentosa mutants. Protein Sci. 16, 1024–1031 (2007)
Sanches, M., Alves, B. S., Zanchin, N. I. & Guimaraes, B. G. The crystal structure of the human Mov34 MPN domain reveals a metal-free dimer. J. Mol. Biol. 370, 846–855 (2007)
Ambroggio, X. I., Rees, D. C. & Deshaies, R. J. JAMM: a metalloprotease-like zinc site in the proteasome and signalosome. PLoS Biol. 2, E2 (2004)
Tran, H. J., Allen, M. D., Lowe, J. & Bycroft, M. Structure of the Jab1/MPN domain and its implications for proteasome function. Biochemistry 42, 11460–11465 (2003)
Pickart, C. M. & Raasi, S. Controlled synthesis of polyubiquitin chains. Methods Enzymol. 399, 21–36 (2005)
Hofmann, R. M. & Pickart, C. M. In vitro assembly and recognition of Lys-63 polyubiquitin chains. J. Biol. Chem. 276, 27936–27943 (2001)
Tenno, T. et al. Structural basis for distinct roles of Lys63- and Lys48-linked polyubiquitin chains. Genes Cells 9, 865–875 (2004)
Varadan, R. et al. Solution conformation of Lys63-linked di-ubiquitin chain provides clues to functional diversity of polyubiquitin signaling. J. Biol. Chem. 279, 7055–7063 (2004)
Hurley, J. H., Lee, S. & Prag, G. Ubiquitin-binding domains. Biochem. J. 399, 361–372 (2006)
Gupta, S. P. Quantitative structure–activity relationship studies on zinc-containing metalloproteinase inhibitors. Chem. Rev. 107, 3042–3087 (2007)
Lipscomb, W. N. & Strater, N. Recent advances in zinc enzymology. Chem. Rev. 96, 2375–2434 (1996)
Holden, H. M., Tronrud, D. E., Monzingo, A. F., Weaver, L. H. & Matthews, B. W. Slow- and fast-binding inhibitors of thermolysin display different modes of binding: crystallographic analysis of extended phosphonamidate transition-state analogues. Biochemistry 26, 8542–8553 (1987)
Zhu, P. et al. A histone H2A deubiquitinase complex coordinating histone acetylation and H1 dissociation in transcriptional regulation. Mol. Cell 27, 609–621 (2007)
Sobhian, B. et al. RAP80 targets BRCA1 to specific ubiquitin structures at DNA damage sites. Science 316, 1198–1202 (2007)
Maytal-Kivity, V., Reis, N., Hofmann, K. & Glickman, M. H. MPN+, a putative catalytic motif found in a subset of MPN domain proteins from eukaryotes and prokaryotes, is critical for Rpn11 function. BMC Biochem. 3, 28 (2002)
Yao, T. & Cohen, R. E. A cryptic protease couples deubiquitination and degradation by the proteasome. Nature 419, 403–407 (2002)
Verma, R. et al. Role of Rpn11 metalloprotease in deubiquitination and degradation by the 26S proteasome. Science 298, 611–615 (2002)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. D 64, 112–122 (2008)
de. la Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997)
Abrahams, J. P. & Leslie, A. G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D 52, 30–42 (1996)
Cowtan, K. D. & Main, P. Improvement of macromolecular electron-density maps by the simultaneous application of real and reciprocal space constraints. Acta Crystallogr. D 49, 148–157 (1993)
Morris, R. J., Perrakis, A. & Lamzin, V. S. ARP/wARP and automatic interpretation of protein electron density maps. Methods Enzymol. 374, 229–244 (2003)
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007)
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Cryst. 30, 1022–1025 (1997)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Iwai, K. et al. Identification of the von Hippel–Lindau tumor-suppressor protein as part of an active E3 ubiquitin ligase complex. Proc. Natl Acad. Sci. USA 96, 12436–12441 (1999)
Acknowledgements
We thank C. Toyoshima for support of this research. We thank M. Shirakawa and T. Tenno for advice about the K63–Ub2 preparation. We thank S. Kaiser for critical reading and improvement of this manuscript. We thank the beam-line staffs at NW12A of PF-AR (Tsukuba, Japan) for technical help during data collection. This work was supported by grants from MEXT to S.F. and O.N. Y.S. is supported by JSPS research fellowships for young scientists.
Author Contributions Y.S. carried out the crystallization and structure determination of the AMSH-LP DUB domain and its complex with K63–Ub2. A.Y. prepared K63–Ub2 and polyubiquitin chains with support from K.O., M.Y., O.N., K.I. and M.K. Y.S. and A.Y. carried out the in vitro DUB assays. K.I. and M.K. provided resources for K63–Ub2 preparation. A.Y., H.M. and S.F. assisted with the crystallization and structure determination. Y.S. and S.F. wrote the paper, with editing from A.Y., H.M., O.N., K.I. and M.K. S.F. and M.K. designed the research. All authors discussed the results and commented on the manuscript. S.F. supervised the work.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Tables S1-S2, Supplementary Figures S1-S8 with legends. (PDF 8438 kb)
Rights and permissions
About this article
Cite this article
Sato, Y., Yoshikawa, A., Yamagata, A. et al. Structural basis for specific cleavage of Lys 63-linked polyubiquitin chains. Nature 455, 358–362 (2008). https://doi.org/10.1038/nature07254
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07254
This article is cited by
-
Dynamic molecular architecture and substrate recruitment of cullin3–RING E3 ligase CRL3KBTBD2
Nature Structural & Molecular Biology (2024)
-
Neutron-encoded diubiquitins to profile linkage selectivity of deubiquitinating enzymes
Nature Communications (2023)
-
Arabidopsis CSN5A Acts as a Transcriptional Co-activator of DREB2C During Seed Germination
Journal of Plant Biology (2023)
-
Plant deubiquitinases: from structure and activity to biological functions
Plant Cell Reports (2023)
-
STAMBPL1 promotes breast cancer cell resistance to cisplatin partially by stabilizing MKP-1 expression
Oncogene (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.