Abstract
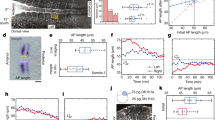
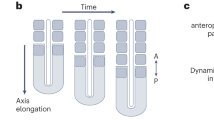
The vertebrate body axis is subdivided into repeated segments, best exemplified by the vertebrae that derive from embryonic somites. The number of somites is precisely defined for any given species but varies widely from one species to another. To determine the mechanism controlling somite number, we have compared somitogenesis in zebrafish, chicken, mouse and corn snake embryos. Here we present evidence that in all of these species a similar ‘clock-and-wavefront’1,2,3 mechanism operates to control somitogenesis; in all of them, somitogenesis is brought to an end through a process in which the presomitic mesoderm, having first increased in size, gradually shrinks until it is exhausted, terminating somite formation. In snake embryos, however, the segmentation clock rate is much faster relative to developmental rate than in other amniotes, leading to a greatly increased number of smaller-sized somites.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cooke, J. & Zeeman, E. C. A clock and wavefront model for control of the number of repeated structures during animal morphogenesis. J. Theor. Biol. 58, 455–476 (1976)
Palmeirim, I., Henrique, D., Ish-Horowicz, D. & Pourquié, O. Avian hairy gene expression identifies a molecular clock linked to vertebrate segmentation and somitogenesis. Cell 91, 639–648 (1997)
Dequeant, M. L. & Pourquié, O. Segmental patterning of the vertebrate embryonic axis. Nature Rev . Genet. 9, 370–382 (2008)
Dequeant, M. L. et al. A complex oscillating network of signaling genes underlies the mouse segmentation clock. Science 314, 1595–1598 (2006)
Dubrulle, J., McGrew, M. J. & Pourquie, O. FGF signaling controls somite boundary position and regulates segmentation clock control of spatiotemporal Hox gene activation. Cell 106, 219–232 (2001)
Sawada, A. et al. Fgf/MAPK signalling is a crucial positional cue in somite boundary formation. Development 128, 4873–4880 (2001)
Aulehla, A. et al. Wnt3a plays a major role in the segmentation clock controlling somitogenesis. Dev. Cell 4, 395–406 (2003)
Aulehla, A. et al. A β-catenin gradient links the clock and wavefront systems in mouse embryo segmentation. Nature Cell Biol. 10, 186–193 (2008)
Richardson, M. K., Allen, S. P., Wright, G. M., Raynaud, A. & Hanken, J. Somite number and vertebrate evolution. Development 125, 151–160 (1998)
Delfini, M. C., Dubrulle, J., Malapert, P., Chal, J. & Pourquie, O. Control of the segmentation process by graded MAPK/ERK activation in the chick embryo. Proc. Natl Acad. Sci. USA 102, 11343–11348 (2005)
Wittler, L. et al. Expression of Msgn1 in the presomitic mesoderm is controlled by synergism of WNT signalling and Tbx6 . EMBO Rep. 8, 784–789 (2007)
Nakajima, Y., Morimoto, M., Takahashi, Y., Koseki, H. & Saga, Y. Identification of Epha4 enhancer required for segmental expression and the regulation by Mesp2. Development 133, 2517–2525 (2006)
Niederreither, K., Subbarayan, V., Dolle, P. & Chambon, P. Embryonic retinoic acid synthesis is essential for early mouse post-implantation development. Nature Genet. 21, 444–448 (1999)
Diez del Corral, R. et al. Opposing FGF and retinoid pathways control ventral neural pattern, neuronal differentiation, and segmentation during body axis extension. Neuron 40, 65–79 (2003)
Burgess, R., Cserjesi, P., Ligon, K. L. & Olson, E. N. Paraxis: a basic helix–loop–helix protein expressed in paraxial mesoderm and developing somites. Dev. Biol. 168, 296–306 (1995)
Mansouri, A. et al. Paired-related murine homeobox gene expressed in the developing sclerotome, kidney, and nervous system. Dev. Dyn. 210, 53–65 (1997)
Yoon, J. K. & Wold, B. The bHLH regulator pMesogenin1 is required for maturation and segmentation of paraxial mesoderm. Genes Dev. 14, 3204–3214 (2000)
Sassoon, D. et al. Expression of two myogenic regulatory factors myogenin and MyoD1 during mouse embryogenesis. Nature 341, 303–307 (1989)
Pownall, M. E. & Emerson, C. P. J. Sequential activation of three myogenic regulatory genes during somite morphogenesis in quail embryos. Dev. Biol. 151, 67–79 (1992)
Yoon, J. K., Moon, R. T. & Wold, B. The bHLH class protein pMesogenin1 can specify paraxial mesoderm phenotypes. Dev. Biol. 222, 376–391 (2000)
McGrew, M. J., Dale, J. K., Fraboulet, S. & Pourquie, O. The lunatic Fringe gene is a target of the molecular clock linked to somite segmentation in avian embryos. Curr. Biol. 8, 979–982 (1998)
Forsberg, H., Crozet, F. & Brown, N. A. Waves of mouse Lunatic fringe expression, in four-hour cycles at two-hour intervals, precede somite boundary formation. Curr. Biol. 8, 1027–1030 (1998)
Zug, G. R., Vitt, L. J. & Caldwell, J. P. Herpetology: an Introductory Biology of Amphibians and Reptiles 2nd edn (Academic, San Diego, 2001)
Tam, P. P. The control of somitogenesis in mouse embryos. J. Embryol. Exp. Morphol. 65 (Suppl). 103–128 (1981)
Primmett, D. R., Norris, W. E., Carlson, G. J., Keynes, R. J. & Stern, C. D. Periodic segmental anomalies induced by heat shock in the chicken embryo are associated with the cell cycle. Development 105, 119–130 (1989)
Giudicelli, F., Ozbudak, E. M., Wright, G. J. & Lewis, J. Setting the tempo in development: an investigation of the zebrafish somite clock mechanism. PLoS Biol. 5, e150 (2007)
Cambray, N. & Wilson, V. Two distinct sources for a population of maturing axial progenitors. Development 134, 2829–2840 (2007)
Sanders, E. J., Khare, M. K., Ooi, V. C. & Bellairs, R. An experimental and morphological analysis of the tail bud mesenchyme of the chicken embryo. Anat. Embryol. (Berl.) 174, 179–185 (1986)
Shum, A. S. et al. Retinoic acid induces down-regulation of Wnt-3a, apoptosis and diversion of tail bud cells to a neural fate in the mouse embryo. Mech. Dev. 84, 17–30 (1999)
Henrique, D. et al. Expression of a Delta homologue in prospective neurons in the chicken. Nature 375, 787–790 (1995)
Acknowledgements
The authors thank M. Gibson, B. Rubinstein, P. Francois and members of the Pourquié laboratory for critical reading and discussions, M. Wahl for the mouse LFNG pictures, members of the Reptile and Aquatics Department, J. Chatfield for editorial assistance, and S. Esteban for artwork. Research was supported by Stowers Institute for Medical Research, and in part by a Defense Advanced Research Projects Agency (DARPA) grant (O.P.). J.L. is supported by Cancer Research UK. Zebrafish were obtained from the Zebrafish International Resource Center (ZIRC) at the University of Oregon, which is supported by a grant from the NIH-NCRR. O.P. is a Howard Hughes Medical Institute Investigator.
Author Contributions C.G. and O.P. designed the experiments, C.G. cloned the snake genes and performed the mouse, chicken and snake in situ hybridizations, E.M.O. performed the fish in situs, C.G. and E.M.O. performed the measurements and analysed the data with O.P. D.B. established the corn snake and zebrafish colony and produced the embryos. C.G. and J.W. performed the cell cycle analysis. J.L. performed the mathematical modelling. C.G., E.M.O., J.L. and O.P. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
The file contains Supplementary Methods with additional references, Supplementary Figures 1-5 with Legends, Supplementary Tables 1-4 and Supplementary Boxes 1-2. (PDF 4543 kb)
Rights and permissions
About this article
Cite this article
Gomez, C., Özbudak, E., Wunderlich, J. et al. Control of segment number in vertebrate embryos. Nature 454, 335–339 (2008). https://doi.org/10.1038/nature07020
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07020
This article is cited by
-
Stochastic gene expression and environmental stressors trigger variable somite segmentation phenotypes
Nature Communications (2023)
-
Metabolic regulation of species-specific developmental rates
Nature (2023)
-
Morphogenesis beyond in vivo
Nature Reviews Physics (2023)
-
The effect of sleep deprivation on postural stability among physically active young adults
Scientific Reports (2023)
-
Fgf8 dynamics and critical slowing down may account for the temperature independence of somitogenesis
Communications Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.