Abstract
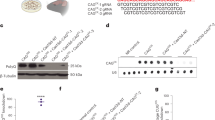
Non-human primates are valuable for modelling human disorders and for developing therapeutic strategies; however, little work has been reported in establishing transgenic non-human primate models of human diseases. Huntington’s disease (HD) is an autosomal dominant neurodegenerative disorder characterized by motor impairment, cognitive deterioration and psychiatric disturbances followed by death within 10–15 years of the onset of the symptoms1,2,3,4. HD is caused by the expansion of cytosine-adenine-guanine (CAG, translated into glutamine) trinucleotide repeats in the first exon of the human huntingtin (HTT) gene5. Mutant HTT with expanded polyglutamine (polyQ) is widely expressed in the brain and peripheral tissues2,6, but causes selective neurodegeneration that is most prominent in the striatum and cortex of the brain. Although rodent models of HD have been developed, these models do not satisfactorily parallel the brain changes and behavioural features observed in HD patients. Because of the close physiological7, neurological and genetic similarities8,9 between humans and higher primates, monkeys can serve as very useful models for understanding human physiology and diseases10,11. Here we report our progress in developing a transgenic model of HD in a rhesus macaque that expresses polyglutamine-expanded HTT. Hallmark features of HD, including nuclear inclusions and neuropil aggregates, were observed in the brains of the HD transgenic monkeys. Additionally, the transgenic monkeys showed important clinical features of HD, including dystonia and chorea. A transgenic HD monkey model may open the way to understanding the underlying biology of HD better, and to the development of potential therapies. Moreover, our data suggest that it will be feasible to generate valuable non-human primate models of HD and possibly other human genetic diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Cummings, C. J. & Zoghbi, H. Y. Trinucleotide repeats: mechanisms and pathophysiology. Annu. Rev. Genomics Hum. Genet. 1, 281–328 (2000)
Davies, S. & Ramsden, D. B. Huntington's disease. Mol. Pathol. 54, 409–413 (2001)
Myers, R. H., Marans, K. S. & MacDonald, M. E. Huntington’s Disease in Genetic Instabilities and Hereditary Neurological Diseases (eds Wells, R. D. & Warren, S. T.) 301–324 (Academic, San Diego, 1998)
Rubinsztein, D. C. Lessons from animal models of Hungtington's disease. Trends Genet. 18, 202–209 (2002)
MacDonald, M. E. et al. HD research collaborative groups. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntintgton’s disease chromosome. Cell 72, 971–983 (1993)
Sharp, A. H. et al. Widespread expression of Huntington’s disease gene (IT15) protein product. Neuron 14, 1065–1074 (1995)
Lane, M. A. Nonhuman primate models in biogerontology. Exp. Gerontol. 35, 533–541 (2000)
King, M. C. & Wilson, A. C. Evolution at two levels in humans and chimpanzees. Science 188, 107–116 (1975)
McConkey, E. H. & Varki, A. A primate genome project deserves high priority. Science 289, 1295–1296 (2000)
Chan, A. W. S., Chong, K. Y., Martinovich, C., Simerly, C. & Schatten, G. Transgenic monkeys produced by retroviral gene transfer into mature oocytes. Science 291, 309–312 (2001)
Chan, A. W. S., Chong, K. Y. & Schatten, G. in Transgenic Animal Technology: A Laboratory Handbook (ed. Pinkert, C. A.) (Academic, San Diego, 2002)
Zhou, H. et al. Huntingtin forms toxic NH2-terminal fragment complexes that are promoted by the age-dependent decrease in proteasome activity. J. Cell Biol. 163, 109–118 (2003)
Gutekunst, C. A. et al. Nuclear and neuropil aggregates in Huntington’s disease: relationship to neuropathology. J. Neurosci. 19, 2522–2534 (1999)
Bates, G. P., Mangiarini, L., Wanker, E. E. & Davies, S. W. Polyglutamine expansion and Huntington’s disease. Biochem. Soc. Trans. 26, 471–475 (1998)
Lin, C. H. et al. Neurological abnormalities in a knock-in mouse model of Huntington’s disease. Hum. Mol. Genet. 10, 137–144 (2001)
Li, H. et al. Ultrastructural localization and progressive formation of neuropil aggregates in Huntington’s disease transgenic mice. Hum. Mol. Genet. 8, 1227–1236 (1999)
Li, H., Li, S. H., Yu, Z. X., Shelbourne, P. & Li, X. J. Huntingtin aggregate-associated axonal degeneration is an early pathological event in Huntington’s disease mice. J. Neurosci. 21, 8473–8481 (2001)
Yu, Z. X. et al. Mutant Huntingtin causes context-dependent neurodegeneration in mice with Huntington’s disease. J. Neurosci. 23, 2193–2202 (2003)
Davies, S. W. et al. Formation of neuronal intranuclear inclusions underlies the neurological dysfunction in mice transgenic for the HD mutation. Cell 90, 537–548 (1997)
DiFiglia, M. et al. Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neuritis in brain. Science 277, 1990–1993 (1997)
Yang, S. H. et al. Enhanced transgenesis by intracytoplasmic injection of envelope-free lentivirus. Genesis 45, 177–183 (2007)
Acknowledgements
We thank J. Fanton (deceased), M. Zelinski-Wooten, M. Sparman and D. Wolf for consultation, C. Lois for lentivirus backbone and C. Testa for critical review of the manuscript. We also thank F. Zhang, T. Caspary, S. Warren, J. Greene, T. Wichmann, M. Wilson, S. Chikazawa, K. Gould, L. Walker, K. Layug, E. Strobert, J. Else, J. Ksiazek, K. Strait, F. Stroud, J. Jenkins, J. Cohen, J. Pare, S. Jenkins, K. Paul, S. Lackey, J. Johnson-Ward, the veterinary staff, the animal resource and the endocrine core laboratory at the Yerkes National Primate Research Center. Acryline was provided by NICHD/NIH. All transgenic HD monkeys were housed under the guideline of the IACUC approved procedures and the support of YNPRC Division of Animal Resources. All newborn monkeys were closely monitored by the veterinary staff and infant care personnel. All procedures were approved by YNPRC/Emory Animal Care and Biosafety Committees. The YNPRC is supported by NIH/NCRR. A.W.S.C., S.H.L. and X.J.-L are supported by grants awarded by the NIH.
Author Contributions S.-H.Y. carried out assisted reproductive technique (ART) in monkeys, viral gene transfer, construct design and molecular analysis; P.-H.C., construct design and evaluation; K.P.-N., ART in monkeys; H.B., animal management; behavioural testing and all animal procedures; K.L., animal care and behavioural testing; E.C.H.C., molecular analysis; J.-J.Y., preparation of high titre lentiviruses; B.S., J.L. and Z.H.F., neuropathological analysis; J.O., surgical procedures and animal care; Y.S., neuropathological analysis; J.B., design of behavioural and cognitive testing; S.M.Z., experimental design and manuscript preparation; S.H.L. and X.J.-L., construct design, analysis and manuscript preparation; A.W.S.C., ART in monkey, viral gene transfer, experimental design, construct design, molecular analysis and manuscript preparation.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figure
The file contains Supplementary Figure S1 with Legend. (PDF 340 kb)
Rights and permissions
About this article
Cite this article
Yang, SH., Cheng, PH., Banta, H. et al. Towards a transgenic model of Huntington’s disease in a non-human primate. Nature 453, 921–924 (2008). https://doi.org/10.1038/nature06975
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06975
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.