Abstract
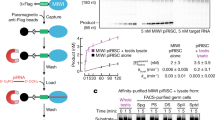
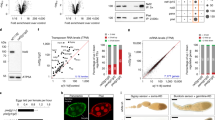
RNA silencing is a conserved mechanism in which small RNAs trigger various forms of sequence-specific gene silencing by guiding Argonaute complexes to target RNAs by means of base pairing1,2. RNA silencing is thought to have evolved as a form of nucleic-acid-based immunity to inactivate viruses and transposable elements. Although the activity of transposable elements in animals has been thought largely to be restricted to the germ line, recent studies have shown that they may also actively transpose in somatic cells, creating somatic mosaicism in animals3. In the Drosophila germ line, Piwi-interacting RNAs arise from repetitive intergenic elements including retrotransposons by a Dicer-independent pathway and function through the Piwi subfamily of Argonautes to ensure silencing of retrotransposons4,5,6,7,8,9. Here we show that, in cultured Drosophila S2 cells, Argonaute 2 (AGO2), an AGO subfamily member of Argonautes, associates with endogenous small RNAs of 20–22 nucleotides in length, which we have collectively named endogenous short interfering RNAs (esiRNAs). esiRNAs can be divided into two groups: one that mainly corresponds to a subset of retrotransposons, and the other that arises from stem–loop structures. esiRNAs are produced in a Dicer-2-dependent manner from distinctive genomic loci, are modified at their 3′ ends and can direct AGO2 to cleave target RNAs. Mutations in Dicer-2 caused an increase in retrotransposon transcripts. Together, our findings indicate that different types of small RNAs and Argonautes are used to repress retrotransposons in germline and somatic cells in Drosophila.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
Small RNA sequences were deposited in the Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/geo/) under the accession number GPL6452.
References
Chapman, E. J. & Carrington, J. C. Specialization and evolution of endogenous small RNA pathways. Nature Rev. Genet. 8, 884–896 (2007)
Peters, L. & Meister, G. Argonaute proteins: mediators of RNA silencing. Mol. Cell 26, 611–623 (2007)
Muotri, A. R. et al. Somatic mosaicism in neuronal precursor cells mediated by L1 reterotranspositioin. Nature 435, 903–910 (2005)
Vagin, V. V. et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006)
Saito, K. et al. Specific association of Piwi with rasiRNAs derived from retrotransposon and heterochromatic regions in the Drosophila genome. Genes Dev. 20, 2214–2222 (2006)
Gunawardane, L. S. et al. A Slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila . Science 315, 1587–1590 (2007)
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila . Cell 128, 1089–1103 (2007)
Yin, H. & Lin, H. An epigenetic activation role of Piwi and a Piwi-associated piRNAs in Drosophila melanogaster . Nature 450, 304–308 (2007)
Nishida, K. M. et al. Gene silencing mechanisms mediated by Aubergine–piRNA complexes in Drosophila male gonad. RNA 13, 1911–1922 (2007)
Hammond, S. M., Boettcher, S., Caudy, A. A., Kobayashi, R. & Hannon, G. J. Argonaute2, a link between genetic and biochemical analyses of RNAi. Science 293, 1146–1150 (2001)
Bernstein, E., Caudy, A. A., Hammond, S. M. & Hannon, G. J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001)
Okamura, K., Ishizuka, A., Siomi, H. & Siomi, M. C. Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev. 18, 1655–1666 (2004)
Lee, Y. S. et al. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 117, 69–81 (2004)
Williams, R. W. & Rubin, G. M. ARGONAUTE1 is required for efficient RNA interference in Drosophila embryos. Proc. Natl Acad. Sci. USA 99, 6889–6894 (2002)
Wang, X. H. et al. RNA interference directs innate immunity against viruses in adult Drosophila . Science 312, 452–454 (2006)
Ruby, J. G. et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans . Cell 127, 1193–1207 (2006)
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005)
Kim, V. N. Small RNAs just got bigger: Piwi-interacting RNAs (piRNAs) in mammalian testes. Genes Dev. 20, 1993–1997 (2006)
Nishikura, K. Editor meets silencer: crosstalk between RNA editing and RNA interference. Nature Rev. Mol. Cell Biol. 7, 919–931 (2006)
Girard, A., Sachidanandam, R., Hannon, G. J. & Carmell, M. A. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442, 199–202 (2006)
Saito, K. et al. Pimet, the Drosophila homolog of HEN1, mediates 2′-O-methylation of Piwi-interacting RNAs at their 3′ ends. Genes Dev. 21, 1603–1608 (2007)
Horwich, M. D. et al. The Drosophila RNA methyltransferase, DmHen1, modifies germline piRNAs and single-stranded siRNAs in RISC. Curr. Biol. 17, 1265–1272 (2007)
Muotri, A. R., Marchetto, M. C. N., Coufal, N. G. & Gage, F. H. The necessary junk: new functions for transposable elements. Hum. Mol. Genet. 16, R159–R167 (2007)
Grewal, S. I. & Elgin, S. C. Transcription and RNA interference in the formation of heterochromatin. Nature 447, 399–406 (2007)
Peng, J. C. & Karpen, G. H. H3K9 methylation and RNA interference regulate nucleolar organization and repeated DNA stability. Nature Cell Biol. 9, 25–35 (2007)
Deshpande, G., Calhoun, G. & Schedl, P. Drosophila argonute-2 is required early in embryogenesis for the assembly of centric/centromeric heterochromatin, nuclear division, nuclear migration, and germ-cell formation. Genes Dev. 19, 1680–1685 (2005)
Miyoshi, K., Tsukumo, H., Nagami, T., Siomi, H. & Siomi, M. C. Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev. 19, 2837–2848 (2005)
Celniker, S. E. et al. Finishing a whole genome shotgun: Release 3 of the Drosophila melanogaster euchromatic genome sequence. Gen. Biol. 3, research0079.1–0079.14 (2002)
Crosby, M. A., Coodman, J. L., Strelets, V. B., Zhang, P. & Gelbart, W. M. the FlyBase Consortium. FlyBase: genomes by the dozen. Nucleic Acids Res. 35 (Database issue). D486–D491 (2007)
Acknowledgements
We are grateful to T. Suzuki and members of the Siomi laboratory for discussion and comments on this manuscript. We thank K. Yamada and E. Hattori for expert assistance in AGO2-associated small RNA mapping and annotation, N. Iwanami and T. Hirose for help in qRT–PCR, and M. Itakura for continuous support and encouragement. This work was supported by MEXT grants to M.C.S. and H.S., a MEXT (Ministry of Education, Culture, Sports, Science and Technology, Japan) 21st COE (Centers of Excellence) postdoctoral fellowship to K.S. and T.S., and NEDO (New Energy and Industrial Technology Development Organization) grants to M.C.S., T.K. and K.A. M.C.S. is supported by CREST from JST. H.S. is a member of the Genome Network Project (MEXT).
Author Contributions Y.K., K.S., T.N.O. and M.C.S. performed AGO2 immunoprecipitations, northern blotting, RNAi, the in vitro cleavage assay, β-elimination and qRT–PCR, and prepared the AGO2-associated small RNA library. T.S. characterized and purified the AGO2 antibody. The bioinformatics analyses of AGO2-associated small RNAs were designed and carried out by T.K., K.S., Y.O. and K.A. M.C.S., K.S., Y.K. and H.S. designed the experiments, discussed the interpretation of the results and co-wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-9 with Legends, Supplementary Tables 1-3 and additional references. (PDF 14560 kb)
Rights and permissions
About this article
Cite this article
Kawamura, Y., Saito, K., Kin, T. et al. Drosophila endogenous small RNAs bind to Argonaute 2 in somatic cells. Nature 453, 793–797 (2008). https://doi.org/10.1038/nature06938
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06938
This article is cited by
-
Regulation of insect behavior by non-coding RNAs
Science China Life Sciences (2024)
-
Mitochondrially mediated RNA interference, a retrograde signaling system affecting nuclear gene expression
Heredity (2023)
-
tRNA-derived fragments from wheat are potentially involved in susceptibility to Fusarium head blight
BMC Plant Biology (2022)
-
Structure of the Dicer-2–R2D2 heterodimer bound to a small RNA duplex
Nature (2022)
-
Conserved chromosomal functions of RNA interference
Nature Reviews Genetics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.