Abstract
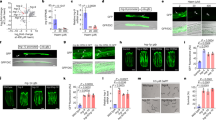
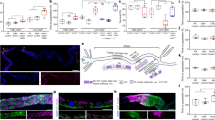
Haems are metalloporphyrins that serve as prosthetic groups for various biological processes including respiration, gas sensing, xenobiotic detoxification, cell differentiation, circadian clock control, metabolic reprogramming and microRNA processing1,2,3,4. With a few exceptions, haem is synthesized by a multistep biosynthetic pathway comprising defined intermediates that are highly conserved throughout evolution5. Despite our extensive knowledge of haem biosynthesis and degradation, the cellular pathways and molecules that mediate intracellular haem trafficking are unknown. The experimental setback in identifying haem trafficking pathways has been the inability to dissociate the highly regulated cellular synthesis and degradation of haem from intracellular trafficking events6. Caenorhabditis elegans and related helminths are natural haem auxotrophs that acquire environmental haem for incorporation into haemoproteins, which have vertebrate orthologues7. Here we show, by exploiting this auxotrophy to identify HRG-1 proteins in C. elegans, that these proteins are essential for haem homeostasis and normal development in worms and vertebrates. Depletion of hrg-1, or its paralogue hrg-4, in worms results in the disruption of organismal haem sensing and an abnormal response to haem analogues. HRG-1 and HRG-4 are previously unknown transmembrane proteins, which reside in distinct intracellular compartments. Transient knockdown of hrg-1 in zebrafish leads to hydrocephalus, yolk tube malformations and, most strikingly, profound defects in erythropoiesis—phenotypes that are fully rescued by worm HRG-1. Human and worm proteins localize together, and bind and transport haem, thus establishing an evolutionarily conserved function for HRG-1. These findings reveal conserved pathways for cellular haem trafficking in animals that define the model for eukaryotic haem transport. Thus, uncovering the mechanisms of haem transport in C. elegans may provide insights into human disorders of haem metabolism and reveal new drug targets for developing anthelminthics to combat worm infestations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ponka, P. Cell biology of heme. Am. J. Med. Sci. 318, 241–256 (1999)
Kaasik, K. & Lee, C. C. Reciprocal regulation of haem biosynthesis and the circadian clock in mammals. Nature 430, 467–471 (2004)
Yin, L. et al. Rev-erbα, a heme sensor that coordinates metabolic and circadian pathways. Science 318, 1786–1789 (2007)
Faller, M. et al. Heme is involved in microRNA processing. Nature Struct. Mol. Biol. 14, 23–29 (2007)
Medlock, A. E. & Dailey, H. A. in Tetrapyrroles (eds Warren, M. & Smith, A. G.) 116–127 (Landes Bioscience and Springer Science + Business Media, Austin, TX, 2007)
Hamza, I. Intracellular trafficking of porphyrins. Am. Chem. Soc. Chem. Biol. 1, 627–629 (2006)
Rao, A. U., Carta, L. K., Lesuisse, E. & Hamza, I. Lack of heme synthesis in a free-living eukaryote. Proc. Natl Acad. Sci. USA 102, 4270–4275 (2005)
Dailey, H. A. Terminal steps of haem biosynthesis. Biochem. Soc. Trans. 30, 590–595 (2002)
Nass, R. & Hamza, I. in Current Protocols in Toxicology (eds Maines, M. D. et al.) 1.9.1–1.9.17 (Wiley, New York, 2007)
Kanehisa, M. et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 34, D354–D357 (2006)
Bonifacino, J. S. & Traub, L. M. Signals for sorting of transmembrane proteins to endosomes and lysosomes. Annu. Rev. Biochem. 72, 395–447 (2003)
Schmidt, P. M. et al. Residues stabilizing the heme moiety of the nitric oxide sensor soluble guanylate cyclase. Eur. J. Pharmacol. 513, 67–74 (2005)
Pellicena, P. et al. Crystal structure of an oxygen-binding heme domain related to soluble guanylate cyclases. Proc. Natl Acad. Sci. USA 101, 12854–12859 (2004)
Goldman, B. S., Beck, D. L., Monika, E. M. & Kranz, R. G. Transmembrane heme delivery systems. Proc. Natl Acad. Sci. USA 95, 5003–5008 (1998)
McGhee, J. D. et al. The ELT-2 GATA-factor and the global regulation of transcription in the C. elegans intestine. Dev. Biol. 302, 627–645 (2007)
Shafizadeh, E. & Paw, B. H. Zebrafish as a model of human hematologic disorders. Curr. Opin. Hematol. 11, 255–261 (2004)
Bennett, C. M. et al. Myelopoiesis in the zebrafish, Danio rerio. Blood 98, 643–651 (2001)
Lin, H. F. et al. Analysis of thrombocyte development in CD41-GFP transgenic zebrafish. Blood 106, 3803–3810 (2005)
Fleming, M. D. et al. Microcytic anaemia mice have a mutation in Nramp2, a candidate iron transporter gene. Nature Genet. 16, 383–386 (1997)
Gunshin, H. et al. Cloning and characterization of a mammalian proton-coupled metal-ion transporter. Nature 388, 482–488 (1997)
Mao, X. et al. A histidine-rich cluster mediates the ubiquitination and degradation of the human zinc transporter, hZIP4, and protects against zinc cytotoxicity. J. Biol. Chem. 282, 6992–7000 (2007)
Rees, E. M. & Thiele, D. J. From aging to virulence: forging connections through the study of copper homeostasis in eukaryotic microorganisms. Curr. Opin. Microbiol. 7, 175–184 (2004)
De Domenico, I., McVey Ward, D. & Kaplan, J. Regulation of iron acquisition and storage: consequences for iron-linked disorders. Nature Rev. Mol. Cell Biol. 9, 72–81 (2008)
Acknowledgements
The Hamza laboratory thanks the DC-Baltimore WormClub for advice and criticism. We also thank P. Aplan, H. Dailey, I. Mather and S. Severance for critical discussions and reading of the manuscript; M. Cam and G. Poy for expertise with microarrays; J. Lippincott-Schwartz and D. Hailey for organelle markers; H.-F. Lin and R. Handin for the Tg(CD41–GFP) transgenic zebrafish line; J. Italiano for use of the Orca IIER charge-coupled device camera/Metamorph software; P. Krieg for the pT7TS Xenopus oocyte expression vector; P. Ponka and R. Eisenstein for the SIH iron chelation; D. Beckett for use of the fluorescent spectrophotometer; and M. Petris for the hZIP4 plasmid. Many of the worm strains were provided by the Caenorhabditis Genetics Center. This work was supported by funding from the National Institutes of Health (NIH) (I.H., M.D.F. and B.H.P.), the March of Dimes Birth Defects Foundation (I.H. and B.H.P.), the NIH/National Institute of Diabetes and Digestive and Kidney Diseases Intramural Research Program (M.K.), Council for Scientific and Industrial Research and Kanwal Rekhi Fellowships (S.K.U.), and a Howard Hughes Medical Institute Undergraduate Science Education Program grant (S.U.).
Author Contributions Experimental design and execution were as follows: worm experiments and microarrays, A.R., A.U.R., M.K. and I.H.; mammalian experiments, A.R., A.U.R., M.T., C.H., S.U., M.D.F. and I.H.; zebrafish experiments, J.A. and B.H.P.; Xenopus injections and measurements, S.K.U. and M.K.M. I.H. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary information
The file contains Supplementary Figures S1-S8 and Supplementary Table S1 with Legends, Supplementary Methods followed by Supplementary Notes. The anti-sense hHRG1 primer was corrected on 06 January 2009 (PDF 868 kb)
Rights and permissions
About this article
Cite this article
Rajagopal, A., Rao, A., Amigo, J. et al. Haem homeostasis is regulated by the conserved and concerted functions of HRG-1 proteins. Nature 453, 1127–1131 (2008). https://doi.org/10.1038/nature06934
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06934
This article is cited by
-
Mechanisms controlling cellular and systemic iron homeostasis
Nature Reviews Molecular Cell Biology (2024)
-
Identification and characterization of a heme exporter from the MRP family in Drosophila melanogaster
BMC Biology (2022)
-
Cryo-EM of CcsBA reveals the basis for cytochrome c biogenesis and heme transport
Nature Chemical Biology (2022)
-
Generation and characterization of human U-2 OS cell lines with the CRISPR/Cas9-edited protoporphyrinogen oxidase IX gene
Scientific Reports (2022)
-
HRG-9 homologues regulate haem trafficking from haem-enriched compartments
Nature (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.