Abstract
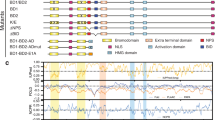
The transcriptional coactivator p300/CBP (CREBBP) is a histone acetyltransferase (HAT) that regulates gene expression by acetylating histones and other transcription factors. Dysregulation of p300/CBP HAT activity contributes to various diseases including cancer1,2,3,4. Sequence alignments, enzymology experiments and inhibitor studies on p300/CBP have led to contradictory results about its catalytic mechanism and its structural relation to the Gcn5/PCAF and MYST HATs5,6,7,8,9. Here we describe a high-resolution X-ray crystal structure of a semi-synthetic heterodimeric p300 HAT domain in complex with a bi-substrate inhibitor, Lys-CoA. This structure shows that p300/CBP is a distant cousin of other structurally characterized HATs, but reveals several novel features that explain the broad substrate specificity and preference for nearby basic residues. Based on this structure and accompanying biochemical data, we propose that p300/CBP uses an unusual ‘hit-and-run’ (Theorell–Chance) catalytic mechanism that is distinct from other characterized HATs. Several disease-associated mutations can also be readily accounted for by the p300 HAT structure. These studies pave the way for new epigenetic therapies involving modulation of p300/CBP HAT activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Goodman, R. H. & Smolik, S. CBP/p300 in cell growth, transformation, and development. Genes Dev. 14, 1553–1577 (2000)
Ogryzko, V. V., Schiltz, R. L., Russanova, V., Howard, B. H. & Nakatani, Y. The transcriptional coactivators p300 and CBP are histone acetyltransferases. Cell 87, 953–959 (1996)
Bannister, A. J. & Kouzarides, T. The CBP co-activator is a histone acetyltransferase. Nature 384, 641–643 (1996)
Iyer, N. G., Ozdag, H. & Caldas, C. p300/CBP and cancer. Oncogene 23, 4225–4231 (2004)
Martinez-Balbas, M. A. et al. The acetyltransferase activity of CBP stimulates transcription. EMBO J. 17, 2886–2893 (1998)
Yuan, L. W. & Giordano, A. Acetyltransferase machinery conserved in p300/CBP-family proteins. Oncogene 21, 2253–2260 (2002)
Poux, A. N., Cebrat, M., Kim, C. M., Cole, P. A. & Marmorstein, R. Structure of the GCN5 histone acetyltransferase bound to a bisubstrate inhibitor. Proc. Natl Acad. Sci. USA 99, 14065–14070 (2002)
Lau, O. D. et al. HATs off: selective synthetic inhibitors of the histone acetyltransferases p300 and PCAF. Mol. Cell 5, 589–595 (2000)
Thompson, P. R., Kurooka, H., Nakatani, Y. & Cole, P. A. Transcriptional coactivator protein p300. Kinetic characterization of its histone acetyltransferase activity. J. Biol. Chem. 276, 33721–33729 (2001)
Thompson, P. R. et al. Regulation of the p300 HAT domain via a novel activation loop. Nature Struct. Mol. Biol. 11, 308–315 (2004)
Karanam, B. et al. Multiple roles for acetylation in the interaction of p300 HAT with ATF-2. Biochemistry 46, 8207–8216 (2007)
Marmorstein, R. Structure of histone acetyltransferases. J. Mol. Biol. 311, 433–444 (2001)
Neuwald, A. F. & Landsman, D. GCN5-related histone N-acetyltransferases belong to a diverse superfamily that includes the yeast SPT10 protein. Trends Biochem. Sci. 22, 154–155 (1997)
Cebrat, M., Kim, C. M., Thompson, P. R., Daugherty, M. & Cole, P. A. Synthesis and analysis of potential prodrugs of coenzyme A analogues for the inhibition of the histone acetyltransferase p300. Bioorg. Med. Chem. 11, 3307–3313 (2003)
Hwang, Y. et al. A selective chemical probe for coenzyme A-requiring enzymes. Angew. Chem. Int. Edn Engl. 46, 7621–7624 (2007)
Segel, I. Enzyme Kinetics (Wiley Interscience, New York, 1975)
Scheibner, K. A., De Angelis, J., Burley, S. K. & Cole, P. A. Investigation of the roles of catalytic residues in serotonin N-acetyltransferase. J. Biol. Chem. 277, 18118–18126 (2002)
Murata, T. et al. Defect of histone acetyltransferase activity of the nuclear transcriptional coactivator CBP in Rubinstein-Taybi syndrome. Hum. Mol. Genet. 10, 1071–1076 (2001)
Muraoka, M. et al. p300 gene alterations in colorectal and gastric carcinomas. Oncogene 12, 1565–1569 (1996)
Gayther, S. A. et al. Mutations truncating the EP300 acetylase in human cancers. Nature Genet. 24, 300–303 (2000)
Kishimoto, M. et al. Mutations and deletions of the CBP gene in human lung cancer. Clin. Cancer Res. 11, 512–519 (2005)
Zheng, Y. et al. Selective HAT inhibitors as mechanistic tools for protein acetylation. Methods Enzymol. 376, 188–199 (2004)
Stimson, L. et al. Isothiazolones as inhibitors of PCAF and p300 histone acetyltransferase activity. Mol. Cancer Ther. 4, 1521–1532 (2005)
Iyer, N. G. et al. p300 regulates p53-dependent apoptosis after DNA damage in colorectal cancer cells by modulation of PUMA/p21 levels. Proc. Natl Acad. Sci. USA 101, 7386–7391 (2004)
Davidson, S. M. et al. The transcriptional coactivator p300 plays a critical role in the hypertrophic and protective pathways induced by phenylephrine in cardiac cells but is specific to the hypertrophic effect of urocortin. ChemBioChem 6, 162–170 (2005)
Zhou, X. Y. et al. Insulin regulation of hepatic gluconeogenesis through phosphorylation of CREB-binding protein. Nature Med. 10, 633–637 (2004)
Varier, R. A. & Kundu, T. K. Chromatin modifications (acetylation/ deacetylation/ methylation) as new targets for HIV therapy. Curr. Pharm. Des. 12, 1975–1993 (2006)
Guidez, F. et al. Histone acetyltransferase activity of p300 is required for transcriptional repression by the promyelocytic leukemia zinc finger protein. Mol. Cell. Biol. 25, 5552–5566 (2005)
Zheng, Y. et al. Synthesis and evaluation of a potent and selective cell-permeable p300 histone acetyltransferase inhibitor. J. Am. Chem. Soc. 127, 17182–17183 (2005)
Qiao, Y., Molina, H., Pandey, A., Zhang, J. & Cole, P. A. Chemical rescue of a mutant enzyme in living cells. Science 311, 1293–1297 (2006)
Acknowledgements
We appreciate advice on the manuscript from W. Cleland, J. Stivers, A. Mildvan and D. Leahy. We thank the National Institutes of Health and the FAMRI, Kaufman and Keck foundations for financial support.
Author Contributions X.L. and L.W. designed and performed reported experiments and prepared manuscript figures and text; K.Z. and P.R.T. developed key reagents and performed preliminary studies that led to the reported experiments; Y.H. developed a key reagent and designed and interpreted some of the experiments. R.M. and P.A.C. designed and supervised experiments and prepared manuscript text. All authors read and approved the submitted manuscript.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
The file contains Supplementary Methods with additional references; Supplementary Figures S1-S10 with Legends and Supplementary Tables S1-S3. (PDF 1922 kb)
Rights and permissions
About this article
Cite this article
Liu, X., Wang, L., Zhao, K. et al. The structural basis of protein acetylation by the p300/CBP transcriptional coactivator. Nature 451, 846–850 (2008). https://doi.org/10.1038/nature06546
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06546
This article is cited by
-
Experimental analysis of bladder cancer-associated mutations in EP300 identifies EP300-R1627W as a driver mutation
Molecular Medicine (2023)
-
TAZ2 truncation confers overactivation of p300 and cellular vulnerability to HDAC inhibition
Nature Communications (2023)
-
Epigenetic mechanisms to propagate histone acetylation by p300/CBP
Nature Communications (2023)
-
Structural mechanism of BRD4-NUT and p300 bipartite interaction in propagating aberrant gene transcription in chromatin in NUT carcinoma
Nature Communications (2023)
-
Modulation of cellular processes by histone and non-histone protein acetylation
Nature Reviews Molecular Cell Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.