Abstract
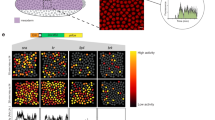
The establishment of complex expression patterns at precise times and locations is key to metazoan development, yet a mechanistic understanding of the underlying transcription control networks is still missing. Here we describe a novel thermodynamic model that computes expression patterns as a function of cis-regulatory sequence and of the binding-site preferences and expression of participating transcription factors. We apply this model to the segmentation gene network of Drosophila melanogaster and find that it predicts expression patterns of cis-regulatory modules with remarkable accuracy, demonstrating that positional information is encoded in the regulatory sequence and input factor distribution. Our analysis reveals that both strong and weaker binding sites contribute, leading to high occupancy of the module DNA, and conferring robustness against mutation; short-range homotypic clustering of weaker sites facilitates cooperative binding, which is necessary to sharpen the patterns. Our computational framework is generally applicable to most protein–DNA interaction systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jackle, H. et al. Transcriptional control by Drosophila gap genes. J. Cell Sci. (suppl.) 16, 39–51 (1992)
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2000)
Berman, B. P. et al. Computational identification of developmental enhancers: conservation and function of transcription factor binding-site clusters in Drosophila melanogaster and Drosophila pseudoobscura. Genome Biol. 5, R61 (2004)
Ochoa-Espinosa, A. et al. The role of binding site cluster strength in Bicoid-dependent patterning in Drosophila. Proc. Natl Acad. Sci. USA 102, 4960–4965 (2005)
Schroeder, M. D. et al. Transcriptional control in the segmentation gene network of Drosophila. PLoS Biol. 2, E271 (2004)
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005)
Tavazoie, S., Hughes, J. D., Campbell, M. J., Cho, R. J. & Church, G. M. Systematic determination of genetic network architecture. Nature Genet. 22, 281–285 (1999)
Albert, R. & Othmer, H. G. The topology of the regulatory interactions predicts the expression pattern of the segment polarity genes in Drosophila melanogaster. J. Theor. Biol. 223, 1–18 (2003)
Segal, E. et al. Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data. Nature Genet. 34, 166–176 (2003)
Granek, J. A. & Clarke, N. D. Explicit equilibrium modeling of transcription-factor binding and gene regulation. Genome Biol. 6, R87 (2005)
Bintu, L. et al. Transcriptional regulation by the numbers: models. Curr. Opin. Genet. Dev. 15, 116–124 (2005)
Zinzen, R. P., Senger, K., Levine, M. & Papatsenko, D. Computational models for neurogenic gene expression in the Drosophila embryo. Curr. Biol. 16, 1358–1365 (2006)
von Dassow, G., Meir, E., Munro, E. M. & Odell, G. M. The segment polarity network is a robust developmental module. Nature 406, 188–192 (2000)
Eldar, A. et al. Robustness of the BMP morphogen gradient in Drosophila embryonic patterning. Nature 419, 304–308 (2002)
Jaeger, J. et al. Dynamic control of positional information in the early Drosophila embryo. Nature 430, 368–371 (2004)
Janssens, H. et al. Quantitative and predictive model of transcriptional control of the Drosophila melanogaster even skipped gene. Nature Genet. 38, 1159–1165 (2006)
Nasiadka, A., Dietrich, B. H. & Krause, H. M. in Advances in Developmental Biology and Biochemistry: Regulation of Gene Expression at the Beginning of Development (ed. DePamphilis, M.). 155–204 (2002)
Rivera-Pomar, R. & Jackle, H. From gradients to stripes in Drosophila embryogenesis: filling in the gaps. Trends Genet. 12, 478–483 (1996)
Furriols, M. & Casanova, J. In and out of Torso RTK signalling. EMBO J. 22, 1947–1952 (2003)
St Johnston, D. & Nusslein-Volhard, C. The origin of pattern and polarity in the Drosophila embryo. Cell 68, 201–219 (1992)
Stormo, G. D. & Hartzell, G. W. Identifying protein-binding sites from unaligned DNA fragments. Proc. Natl Acad. Sci. USA 86, 1183–1187 (1989)
Myasnikova, E., Samsonova, A., Kozlov, K., Samsonova, M. & Reinitz, J. Registration of the expression patterns of Drosophila segmentation genes by two independent methods. Bioinformatics 17, 3–12 (2001)
Rajewsky, N., Vergassola, M., Gaul, U. & Siggia, E. D. Computational detection of genomic cis-regulatory modules applied to body patterning in the early Drosophila embryo. BMC Bioinformatics 3, 30 (2002)
Simpson-Brose, M., Treisman, J. & Desplan, C. Synergy between the hunchback and bicoid morphogens is required for anterior patterning in Drosophila. Cell 78, 855–865 (1994)
Tanay, A. Extensive low-affinity transcriptional interactions in the yeast genome. Genome Res. 16, 962–972 (2006)
Carr, A. & Biggin, M. D. A comparison of in vivo and in vitro DNA-binding specificities suggests a new model for homeoprotein DNA binding in Drosophila embryos. EMBO J. 18, 1598–1608 (1999)
Biggin, M. D. & Tjian, R. Transcriptional regulation in Drosophila: the post-genome challenge. Funct. Integr. Genomics 1, 223–234 (2001)
Raser, J. M. & O'Shea, E. K. Control of stochasticity in eukaryotic gene expression. Science 304, 1811–1814 (2004)
Ptashne, M. & Gann, A. Genes and Signals 26–37 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2002)
Crauk, O. & Dostatni, N. Bicoid determines sharp and precise target gene expression in the Drosophila embryo. Curr. Biol. 15, 1888–1898 (2005)
Lebrecht, D. et al. Bicoid cooperative DNA binding is critical for embryonic patterning in Drosophila. Proc. Natl Acad. Sci. USA 102, 13176–13181 (2005)
Segal, E. et al. A genomic code for nucleosome positioning. Nature 442, 772–778 (2006)
Vashee, S., Melcher, K., Ding, W. V., Johnston, S. A. & Kodadek, T. Evidence for two modes of cooperative DNA binding in vivo that do not involve direct protein–protein interactions. Curr. Biol. 8, 452–458 (1998)
Hoch, M., Seifert, E. & Jackle, H. Gene expression mediated by cis-acting sequences of the Krüppel gene in response to the Drosophila morphogens bicoid and hunchback. EMBO J. 10, 2267–2278 (1991)
Small, S., Kraut, R., Hoey, T., Warrior, R. & Levine, M. Transcriptional regulation of a pair-rule stripe in Drosophila. Genes Dev. 5, 827–839 (1991)
Rivera-Pomar, R., Lu, X., Perrimon, N., Taubert, H. & Jackle, H. Activation of posterior gap gene expression in the Drosophila blastoderm. Nature 376, 253–256 (1995)
Arnosti, D. N., Barolo, S., Levine, M. & Small, S. The eve stripe 2 enhancer employs multiple modes of transcriptional synergy. Development 122, 205–214 (1996)
Sauer, F. & Jackle, H. Heterodimeric Drosophila gap gene protein complexes acting as transcriptional repressors. EMBO J. 14, 4773–4780 (1995)
La Rosee, A., Hader, T., Taubert, H., Rivera-Pomar, R. & Jackle, H. Mechanism and Bicoid-dependent control of hairy stripe 7 expression in the posterior region of the Drosophila embryo. EMBO J. 16, 4403–4411 (1997)
Langeland, J. A., Attai, S. F., Vorwerk, K. & Carroll, S. B. Positioning adjacent pair-rule stripes in the posterior Drosophila embryo. Development 120, 2945–2955 (1994)
Meinhardt, H. Hierarchical inductions of cell states: a model for segmentation in Drosophila. J. Cell Sci. (Suppl.) 4, 357–381 (1986)
Driever, W. & Nusslein-Volhard, C. The bicoid protein determines position in the Drosophila embryo in a concentration-dependent manner. Cell 54, 95–104 (1988)
Ephrussi, A. & St Johnston, D. Seeing is believing: the bicoid morphogen gradient matures. Cell 116, 143–152 (2004)
Hoch, M., Gerwin, N., Taubert, H. & Jackle, H. Competition for overlapping sites in the regulatory region of the Drosophila gene Kruppel. Science 256, 94–97 (1992)
Stanojevic, D., Small, S. & Levine, M. Regulation of a segmentation stripe by overlapping activators and repressors in the Drosophila embryo. Science 254, 1385–1387 (1991)
Sutrias-Grau, M. & Arnosti, D. N. CtBP contributes quantitatively to Knirps repression activity in an NAD binding-dependent manner. Mol. Cell. Biol. 24, 5953–5966 (2004)
Gray, S. & Levine, M. Short-range transcriptional repressors mediate both quenching and direct repression within complex loci in Drosophila. Genes Dev. 10, 700–710 (1996)
Arnosti, D. N., Gray, S., Barolo, S., Zhou, J. & Levine, M. The gap protein knirps mediates both quenching and direct repression in the Drosophila embryo. EMBO J. 15, 3659–3666 (1996)
Acknowledgements
We thank D. Leaman and M. Dandapani for the in vivo analysis of binding sites and are indebted to E. Siggia, S. Sinha and J. Widom for valuable discussions at the outset of the project. This work was supported by a Fellowship from the Center for Studies in Physics and Biology at Rockefeller University (E.S.), by the European Network of Excellence (E.S. and T.R.-S.), by a Rockefeller University Graduate Fellowship (M.S.) and by an NIH grant (U.G.); E.S. is the incumbent of the Soretta and Henry Shapiro career development chair.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-12 with Legends and Supplementary Methods. The Supplementary Figures include additional experimental and computational results. The Supplementary Methods provide a detailed description of the computational model. (PDF 1202 kb)
Rights and permissions
About this article
Cite this article
Segal, E., Raveh-Sadka, T., Schroeder, M. et al. Predicting expression patterns from regulatory sequence in Drosophila segmentation. Nature 451, 535–540 (2008). https://doi.org/10.1038/nature06496
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06496
This article is cited by
-
Systematic analysis of low-affinity transcription factor binding site clusters in vitro and in vivo establishes their functional relevance
Nature Communications (2022)
-
Embryonic development across space and time
Nature Computational Science (2021)
-
Improved recovery of cell-cycle gene expression in Saccharomyces cerevisiae from regulatory interactions in multiple omics data
BMC Genomics (2020)
-
A sensitive mNeonGreen reporter system to measure transcriptional dynamics in Drosophila development
Communications Biology (2020)
-
Eukaryotic transcription factors can track and control their target genes using DNA antennas
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.