Abstract
Stem cell self-renewal implies proliferation under continued maintenance of multipotency. Small changes in numbers of stem cells may lead to large differences in differentiated cell numbers, resulting in significant physiological consequences. Proliferation is typically regulated in the G1 phase, which is associated with differentiation and cell cycle arrest1. However, embryonic stem (ES) cells may lack a G1 checkpoint2,3. Regulation of proliferation in the ‘DNA damage’ S/G2 cell cycle checkpoint pathway is known for its role in the maintenance of chromatin structural integrity4. Here we show that autocrine/paracrine γ-aminobutyric acid (GABA) signalling by means of GABAA receptors negatively controls ES cell and peripheral neural crest stem (NCS) cell proliferation, preimplantation embryonic growth and proliferation in the boundary-cap stem cell niche, resulting in an attenuation of neuronal progenies from this stem cell niche. Activation of GABAA receptors leads to hyperpolarization, increased cell volume and accumulation of stem cells in S phase, thereby causing a rapid decrease in cell proliferation. GABAA receptors signal through S-phase checkpoint kinases of the phosphatidylinositol-3-OH kinase-related kinase family and the histone variant H2AX. This signalling pathway critically regulates proliferation independently of differentiation, apoptosis and overt damage to DNA. These results indicate the presence of a fundamentally different mechanism of proliferation control in these stem cells, in comparison with most somatic cells, involving proteins in the DNA damage checkpoint pathway.
Similar content being viewed by others
Main
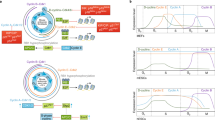
Expression of subunits consistent with functional GABAA receptors (GABAARs) was found in both ES cells positive for stage-specific embryonic antigen 1 (SSEA-1) and NCS cells derived from the boundary cap (BC)5 (Fig. 1a and Supplementary Fig. 1) with a common and possibly critical expression of the β3 subunit. GABA or the GABAAR-specific agonist muscimol evoked an outward current in whole-cell voltage-clamp recordings of ES cells held at -60 mV (n = 15) and the amplitude was decreased significantly by the GABAAR-specific antagonist bicuculline (Fig. 1b and Supplementary Fig. 2a). With the use of the perforated-patch technique the resting membrane potential of ES cells was estimated as -26.05 ± 1.93 mV (n = 20; mean ± s.e.m.). The GABA reversal potential calculated by measuring the amplitude of the induced current at different holding potentials showed a linear relationship that reversed at -78 mV (n = 11) (Fig. 1c and Supplementary Fig. 2b). The patch clamp data were independently confirmed by using a voltage-sensitive dye, bis-(1,3-dibutylbarbituric acid)trimethine oxonol (DiBAC4(3)), in ES and NCS cells (Supplementary Figs 2c, d and 3a). Ca2+ transients were not detected in NCS cells in response to GABAAR activation (Supplementary Fig. 3b). Taken together, these data show that ES and NCS cells possess functional GABAARs that induce a hyperpolarizing current by means of an influx of Cl-.
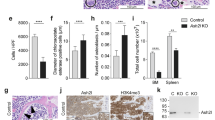
a, Expression analysis of GABAAR subunits and synthesizing enzymes by real-time RT–PCR of SSEA-1+ ES cells and NCS cells isolated by flow cytometry (+, ++, +++, expression levels; -, no expression). b, Electrophysiological analysis of R1 ES cells revealed an outward GABA-induced current that was reversibly blocked by the GABAA receptor antagonist bicuculline (100 μM). c, Reversal potential of the GABA-induced current (-78 mV) estimated by using the perforated-patch voltage-clamp technique and by stimulating with GABA at different holding potentials. d, ES cell numbers after 24 h of exposure to muscimol (n = 3). e, Photomicrograph of ES cell colonies in d. f, Flow cytometric analysis of forward and side scattering (FSC and SSC, respectively) of ES cells revealing the relative numbers of cell doublets reflecting mitotic events after 2 h of agonist treatment. g, Quantification of f (n = 3). h, Block-release experiment showing NCS cell spheres after a three-day treatment with or without muscimol and three days after release. i, j, Cell size measurement by FCS analyses with flow cytometry after muscimol. i, Distribution of cells within FSC levels. The colour code in i is the same as in j. j, Average of FSC levels revealing a rapid increase in cell size by GABAAR activation (n = 3). Incub., incubation. Error bars indicate s.e.m. (two-tailed Student’s t-test). Scale bars, 200 μm (d) and 25 μm (g).
Next we examined the significance of GABAAR activation in stem cell proliferation. The rate of mitotic events was determined by exploiting the tight adhesion that occurs between daughter ES cells after mitosis, resulting in cell doublets that are identified by flow cytometry forward scattering (FSC) 2 h after dissociation (Fig. 1f). Muscimol decreased cell number (Fig. 1d,e) and mitotic events (Fig. 1f, g) in ES cell cultures and potently blocked the proliferation of NCS cells in a reversible manner (Fig. 1h and Supplementary Fig. 3c). The percentage of multipotent SSEA-1+ ES cells and multipotency of NCS cells was unaffected, as were colony morphology and activity and the levels of markers for survival, death or differentiation in response to the agonist (Supplementary Fig. 4). The effect of muscimol seemed to be inversely proportional to cell numbers, because high-density cultures of both cell types showed a lower response. The density dependence of responses to the agonist suggested endogenous signalling. Expression analyses of components necessary for the production and release of GABA in ES and NCS cells showed their presence in both cell types (Fig. 1a and Supplementary Figs 1, 5 and 6); vesicular GABA transporter was localized in subcellular puncta (Fig. 2a and Supplementary Fig. 5). Differentiating ES cells (namely SSEA-1- cells) rapidly lost expression of these markers, indicating that endogenous GABA signalling may be specific to stem cell populations (Supplementary Fig. 6). Inhibition of endogenous signalling with bicuculline in high-density ES cell cultures significantly increased cell numbers (Fig. 2b) and led to a marked increase in incorporation of bromodeoxyuridine (BrdU) into NCS cells (Fig. 2c, d). Suppression of GABAAR β3 expression by RNA interference (RNAi; Supplementary Fig. 1i) led to an increase in ES cells without affecting colony morphology (Fig. 2e, f) and increased mitotic events in the 2 h cell doublet assay (Fig. 2g). These data indicate that endogenously synthesized GABA functions in an autocrine/paracrine manner in stem cells.
a, Immunofluorescence staining for vesicular GABA transporter (VGAT) in ES cells revealing a punctate localization (colour scale shows depth). b, Analysis of autocrine GABA signalling through the GABAAR. ES cell numbers with bicuculline for two passages (n = 3) are shown. c, BrdU pulse (2 h) of NCS cells treated with bicuculline for 12 h. d, Quantification of c (n = 74 clones). e, ES cell numbers four days after transfection with siRNA against the GABAAR β3 subunit (n = 3). f, Photomicrograph of ES cell colonies in e. g, ES cell proliferation 48 h after siRNA transfection by the 2-h doublet assay (n = 3). h, BrdU incorporation over the course of 1.5 h in ES cells exposed to muscimol (n = 3). i, Quantitative analysis showing the percent change of cells in S-phase based on flow cytometric cell cycle profile analyses following muscimol or antagonist (picrotoxin) for 6 h by flow cytometry. j, Cell cycle profiles of i. PI, propidium iodide. Error bars indicate s.e.m. (two-tailed Student’s t-test). Scale bars, 5 μm (a), 75 μm (c) and 400 μm (f).
Consistent with unaffected cell cycle regulation at the G1–S transition (Supplementary Fig. 7a), the rapid effects of muscimol (Fig. 1f, g) suggested a mechanism acting within 2 h of mitosis; that is, in late S, G2 or M phase. BrdU incorporation during 1.5 h of muscimol stimulation showed a rapid threefold decrease in DNA synthesis (Fig. 2h). Cell cycle distribution analysis by flow cytometry revealed an accumulation or decrease in ES cells in S phase after 6 h of activation or inhibition of the receptor, respectively (Fig. 2i, j); this was not observed in differentiated ES cells (Supplementary Fig. 6u). ES cells responded immediately to muscimol by decreasing FSC (Fig. 1i, j) which correlates with osmotic swelling6. Thus, an influx of Cl- ions leads to a rapid increase in cell size followed by a control of proliferation by regulating cell cycle progression in S phase. ES cells within colonies responded synchronously to muscimol and showed dye coupling (Fig. 3c; n = 6). This suggests an electrochemical coupling that may allow hyperpolarizing currents to spread to neighbouring ES cells via gap junctions, coordinating the S-phase checkpoint pathway within a colony.
a, Immunostaining of ES cells for γ-H2AX after 6 h of exposure to muscimol. Open arrow, γ-H2AX+ colony; filled arrow, baseline reactivity. b, Punctate nuclear localization of γ-H2AX staining in a single ES cell. c, Single ES cells filled with a gap-junction-permeable dye to reveal electrochemical coupling between cells within colonies; arrows indicate injected cells (n = 6). d, Western blot (WB) analysis of γ-H2AX in ES cells after exposure to muscimol or bicuculline and ATM/DNA-PK inhibitor (A/D i) for 6 h. e, Comet assay for DNA damage by 2 h of exposure to muscimol in ES cells. f, Quantification of e. g, Quantification of ATR RNAi-treated ES cells in S phase after 2 h of exposure to muscimol. h, i, Number of P-ATMhigh cells by flow cytometric analysis after 2 h of exposure to muscimol in β3 subunit siRNA-treated cells. Dox., doxyrubicin. j, Cell cycle profiles of R1 ES cells with and without ATM/DNA-PK inhibitor (A/D i). k, ES cell numbers at 48 h with control siRNA or H2AX and control or H2AX and GABAAR β3 subunit siRNA (n = 3). l, Effects of muscimol on H2AX siRNA-treated ES cells in a 2 -h cell doublet analysis (n = 3). m, Cell cycle profiles of R1 ES cells transfected with GABAAR β3 and H2AX siRNAs. n, Flow cytometric analysis of ES cells after a 50-min BrdU pulse. BrdU increases with β3/H2AX siRNA. o, Flow cytometric analysis of R1 ES cells treated with the ATM/DNA-PK (A/D) inhibitor. BrdU increases in the presence of inhibitor. Error bars indicate s.e.m. (two-tailed Student’s t-test). Scale bars, 200 μm (a), 5 μm (b), 50 μm (c) and 100 μm (e).
We analysed active phosphorylated histone H2AX (γ-H2AX), which is a critical factor of the S/G2 DNA-damage checkpoint complex7, after exposing ES and NCS cells to muscimol. Muscimol rapidly increased γ-H2AX levels in nuclear foci of ES cells (Fig. 3a, b, d) which could be inhibited by antagonist (Fig. 3d). A significant increase in γ-H2AX was seen also in NCS cells (Supplementary Fig. 3d). DNA tail comet assays revealed no overt DNA damage after 2 h of muscimol treatment (Fig. 3e, f). H2AX is phosphorylated by the phosphatidylinositol-3-OH kinase-related kinase (PIKK) family of kinases, including ataxia telangiectasia mutated (ATM), ataxia telangiectasia, Rad3-related (ATR) and DNA protein kinase (DNA-PK)4. In ES cells, short interfering RNA (siRNA) against ATR abolished the cell cycle distribution response to muscimol (Fig. 3g). ATR activates ATM in response to replication fork stalling8; we therefore analysed activated phosphorylated ATM (P-ATM). P-ATM was increased and decreased by muscimol and β3 siRNA, respectively (Fig. 3h, i). Furthermore, an ATM/DNA-PK inhibitor that blocked the phosphorylation and activity of ATM (Supplementary Fig. 7c, d) decreased γ-H2AX levels (Fig. 3d) and brought about a marked decrease in cells in S phase (Fig. 3j). The inability of the inhibitor to block all γ-H2AX (Fig. 3d) suggests a residual ATM/DNA-PK-independent component of H2AX activation, presumably by ATR. These results show that the PIKK family of kinases mediates GABAAR activation of H2AX in the absence of overt DNA damage.
The effect of GABA on cell proliferation was critically dependent on H2AX, because transfection of ES and NCS cells with siRNA against H2AX resulted in an increase in cell numbers that was not additive when both H2AX and GABAAR expression were decreased (Fig. 3k), and H2AX RNAi eliminated the effect of muscimol in NCS cells (Supplementary Fig. 3e) and ES cells in the 2-h cell doublet assay (Fig. 3l). Muscimol-induced γ-H2AX was consistently decreased in ES cells treated with β3 siRNA (Supplementary Fig. 7f). ES cell cycle distribution analysis revealed a marked decrease in cells in S phase with siRNAs (Fig. 3m and Supplementary Fig. 7b), similar to that with the ATM/DNA-PK inhibitor. This could be caused either by an increase in the time spent in other cell cycle phases or by an increased rate of progression of cells through S phase. β3, H2AX RNAi and ATM/DNA-PK inhibitor all markedly increased BrdU incorporation and decreased the proportion of cells at any given point in S phase (Fig. 3n, o), reflecting an increased rate of progression through S phase.
Next we addressed whether GABAAR activation controlled proliferation in the preimplantation embryo from which the ES cells were derived. Blastocysts responded rapidly to muscimol ex vivo by decreasing DiBAC4(3) fluorescence, which is consistent with hyperpolarization (Fig. 4a). In vivo, muscimol caused a marked decrease in BrdU incorporation in blastocysts (Fig. 4b). To investigate the physiological role of this during early development, GABAAR β3 subunit siRNA was injected into the cytoplasm of mouse zygotes, which developed into blastocysts in vivo. Blastocyst-stage embryos exposed to β3 siRNA developed faster than controls and contained smaller cells and a larger blastocoel (Fig. 4c, d). These results show that proliferation and growth of the preimplantation embryo is regulated by GABAAR signalling, similarly to ES cells.
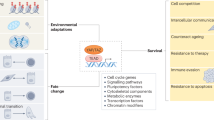
a, Membrane potentiometric analysis of blastocysts with DiBAC4(3) (1 μM) immediately before and 2 min after administration of GABAAR agonist (muscimol). b, BrdU staining of blastocysts after 5 h of exposure in vivo to muscimol and a 1-h BrdU pulse. c, Zygote embryos microinjected with siRNA against GABAAR β3 subunit were implanted into pseudopregnant recipients and developed into morula/blastocysts. Blue denotes blastocoel. d, Quantification of morulae and blastocysts with large or small blastocoels (P < 0.01, χ2 test). e, BrdU and Sox10 staining of Krox20eGFP/+ mice injected with bicuculline at E11 followed by a 2-h BrdU pulse 8 h later. There are fewer BrdU+ (arrowheads) nuclei in the control than in the bicuculline-treated sample. f, Quantification of e (one-way analysis of variance, n = 3). g, Sections of DRG from Krox20Cre/+ R26R-EYFP mice killed at E14 that received muscimol between E11 and E14. h, Quantification of g. Lineage tracing shows a significant decrease in the number of progeny from the BC in the DRG of treated mice (one-way analysis of variance, n = 4). Error bars indicate s.e.m. (two-tailed Student’s t-test). i, Proposed model for GABA-mediated control of proliferation by means of chromatin modification in stem cells. ES and NCS cells synthesize physiological levels of GABA, which activate GABAARs and cause hyperpolarization through Cl- influx. GABA signals by means of PIKKs and activates histone H2AX by phosphorylation of Ser 139 to decrease DNA synthesis and cell cycle progression. j, Proposed consequences of GABA signalling in stem cells. GABA levels correlate with stem cell numbers, which results in an inverse correlation between proliferation rate and stem cell pool size. Scale bars, 25 μm (a, b), 20 μm (c), 10 μm (e) and 50 μm (g).
NCS cells are derived from the BC stem cell niche located at the border between the peripheral nervous system and the central nervous system5,9 and produce peripheral sensory neurons and glia10. To test whether GABAAR activation regulates proliferation in this population in vivo, bicuculline was administered to Krox20eGFP/+ mice expressing the fluorophore enhanced green fluorescent protein (EGFP) specifically in BC cells at embryonic day 11 (E11) (ref. 10). Despite a high rate of proliferation, with more than 20% of the cells incorporating BrdU after a 2-h pulse, blocking endogenous GABA signalling led to a nearly twofold increase in incorporation (Fig. 4e, f). Mice expressing a knock-in allele for the Cre recombinase in the Krox20 locus (Krox20Cre/+)11,12 crossed with a ROSA26 reporter (R26R) mouse strain permanently activating expression of the fluorophore enhanced yellow fluorescent protein (EYFP) on recombination10,13 was used to trace the cell lineage of BC progenies genetically. Consistent with previous findings was the observation of numerous EYFP+ cells within the dorsal root entry zone, at motor exit points and in the dorsal root ganglion (DRG) in E14 sections at thoracic levels (Fig. 4g)10. The administration of muscimol to pregnant mice between E11 and E14 led to a marked decrease in EYFP+ BC-derived cells (Fig. 4g, h). These results show that GABA signalling controls the number of cells produced from the BC stem cell niche during development.
Our results (Fig. 4i, j) show that endogenously produced GABA acting through PIKK-family proteins and H2AX has the following functions: first, it regulates ES and NCS cell proliferation; second, it determines early embryo size; and third, it determines proliferation in the BC stem cell niche in vivo, resulting in direct consequences on progeny cell number. Unlike conventional G1-phase cell cycle control, which is linked to cell differentiation, this S-phase mechanism may provide a way to control proliferation independently of differentiation in the stem cells. Induction of the PIKK/H2AX pathway seems to be independent of DNA damage, in a similar manner to that previously described during sex chromosome inactivation14, hypoxia15 and mitosis16,17. We find it intriguing that ES and NCS cells share a similar mechanism regulating their proliferation, especially in view of the reported effects of GABA on the proliferation of adult brain progenitor cells that are positive for glial fibrillary acidic protein18. Thus, seemingly disparate biological processes may be affected by a common mechanism of control of stem cell proliferation.
Methods Summary
The EB5 and R1 ES strains were cultured in knockout serum replacement and leukaemia inhibitory factor (LIF) or as described19, and NCS cells were cultured as described5. Antibody against SSEA-1 or γ-H2AX together with secondary antibody was used for flow cytometry analysis on a FACScan instrument and cell sorting on a FACSVantage SE/FACS DiVa instrument (Becton Dickinson). Quantification was performed with CellQuest Pro software (Becton Dickinson). Immunostainings, real-time RT–PCR and biochemical analyses were performed in accordance with standard procedures. Two siRNAs (27-mers) were designed for each mRNA and transfected at 5 μM. Control siRNA was used at equimolar levels. For in vivo analyses, zygotes were injected into the cytoplasm with 10 pl of 20 μM siRNA. Picrotoxin, bicuculline and muscimol were used at 50, 20 and 100 μM, respectively, on cultured cells with controls receiving vehicle. For experiments in vivo, bicuculline or muscimol was diluted in PBS and administered intraperitoneally to pregnant females at 4 mg kg-1; for long-term experiments this dose was given twice a day. BrdU (50 mg kg-1, in PBS) was administered intraperitoneally. Mice were held on a pure C57/Bl6 or mixed C57Bl6/DBA2 background10. ATM/DNA-PK inhibitor (Calbiochem) was used at 10 μM.
Online Methods
Cell culture
The ES EB5 strain was used for most experiments, and the R1 ES cell strain cultured on feeder cells was used for confirmation experiments. EB5 cells were cultured in serum-free medium (Knockout DMEM, 15% Knockout Serum Replacement, 1 × non-essential amino acids, 2 mM glutamine, 0.4 mM 2-mercaptoethanol, all from Gibco) or in DMEM/F12, N2 supplement, 0.4 mM 2-mercaptoethanol, 5 mM HEPES, 10 ng ml-1 basic fibroblast growth factor (modified from ref. 19) and 1,000 U ml-1 ESGRO (Chemicon) in suspension or on tissue culture plates coated with 0.1% gelatin. For experiments and cell counts, ES cells were grown on six-well plates. The NCS cells were isolated and cultured as described previously5. ES cells were differentiated in DMEM and 10% FCS without LIF.
Mouse lines
All mouse lines used in this study were maintained in either a pure C57/Bl6 or a mixed C57Bl6/DBA2 background. In the Krox20Cre/+ and Krox20GFP/+ alleles, the Krox20 coding sequence was replaced by the Cre recombinase coding sequence and EGFP, respectively. R26R-EYFP expresses EYFP from the ROSA locus on Cre recombination.
Flow cytometry
For live cell sorting and analysis, cells were dissociated and incubated in primary antibody against SSEA-1 (diluted 1:400 in PBS including 1% BSA; Developmental Studies Hybridoma Bank) for 1 h at 4 °C, followed by incubation in fluorescein isothiocyanate (FITC)-conjugated secondary antibody (diluted 1:400; Jackson Immuno Research) for 1 h at 4 °C followed by incubation in PBS with 1% BSA and 1 μg ml-1 propidium iodide (Sigma) to discern dead cells. Negative controls were treated identically but without primary antibody.
ES cell doublet analysis was performed by first passaging to single cells by incubation in TrypLE Express (Invitrogen) for 15 min at 37 °C and filtering through a 40-μm filter. The cells were then checked in a microscope to ensure that only single cells were present. The cells were then incubated for 2 h at low density under non-adherent conditions in Petri dishes. After the addition of propidium iodide to discern dead cells, live cells were analysed for forward and side scattering and the population of cell doublets was gated and quantified.
For cell cycle analysis, cells were fixed overnight with 70% ethanol and rehydrated in PBS with RNase and propidium iodide. For double staining with propidium iodide and γ-H2AX (γ-H2AX diluted 1:400; Upstate) cells were fixed briefly with 4% paraformaldehyde in PBS before immunostaining. Cell analysis was performed on a FACScan instrument and cell sorting on a FACSVantage SE/FACS DiVa instrument (Becton Dickinson). Cell cycle analysis and quantification were performed with CellQuest Pro software (Becton Dickinson).
Electrophysiology
Whole-cell and perforated-patch voltage-clamp recordings were used to monitor the effect of GABA in ES cells by using the Axopatch 200A amplifier (Axon Instruments). Whole-cell recordings were made with an intracellular solution containing (in mM): potassium gluconate 120, KCl 5, CaCl2 0.0001, Mg2ATP 4, Na4GTP 0.3, HEPES 10, Na2 phosphocreatine 10; pH adjusted to 7.4 with KOH, 275–280 osM. For perforated-patch recordings the tip of the pipette was filled by dipping it briefly (for less than 1 s) into the normal intracellular solution. The rest of the pipette was filled with amphotericin B (240 μg ml-1 intracellular solution). The cells were continuously perfused with culture medium through a gravity-driven microperfusion system with the nozzle positioned close to the recorded cell. The different drugs were added to this solution at appropriate concentrations. Data were acquired and analysed with pClamp software (Axon Instruments). For cell-coupling experiments a gap-junction-permeable dye was injected through the patch pipette during whole-cell patch-clamp recordings.
Immunostaining and western blotting
Cells and sections were fixed in 4% paraformaldehyde in PBS (pH 7.4). Blastocysts were treated with Tyrode’s solution (Sigma) to remove the zona pellucida, fixed in 4% paraformaldehyde and dried on Superfrost Plus glass slides (Microm) before immunostaining.
After being blocked for 2 h in PBS containing 3% BSA and 0.3% Triton X-100 and after two washes in PBS, cells were incubated in primary antibody against VGAT (diluted 1:400; Advanced Targeting Systems), GABAAR β3 subunit (diluted 1:1,000; Chemicon), Gad67 (diluted 1:1,000; Chemicon), GABA (diluted 1:2,000; Sigma), BrdU-FITC (diluted 1:200; Becton Dickinson), γ-H2AX (diluted 1:400; Upstate), Sox10 (guinea pig, diluted 1:1,000, gift from M. Wegner); SSEA-1 (mouse, diluted 1:400; Chemicon); monoamino oxidase B (rabbit, diluted 1:2,000, gift from O. Cases), Sox2 (diluted 1:400; Chemicon), Oct4 (diluted 1:400; Chemicon), BrdU (rat, diluted 1:125; AbCam), phospho-ATM (diluted 1:200; Cell Signaling) and ATM/ATR substrate (diluted 1:200; Cell Signaling) in PBS containing 1% BSA and 0.3% Triton X-100. For immunostaining against GABA, cells were also fixed with 1% glutaraldehyde. For anti-BrdU staining, cells were treated with 0.1 M HCl before being stained.
Imaging was performed with an Axiovert 100M microscope (Zeiss) equipped with a Hamamatsu C4742-95 digital camera and Openlab 5 software (Improvision) or an Axioplan 2 microscope (Zeiss) with an LSM 510 system and software (Zeiss) for confocal imaging.
Western blotting was performed as described20 with anti-γ-H2AX (diluted 1:1,000; Upstate), ATM/ATR substrate (diluted 1:1,000, Cell Signaling) and phospho-ATM (diluted 1:800; Cell Signaling) antibodies. Secondary antibodies were conjugated with horseradish peroxidase (GE Healthcare) and films were developed with the ECL enhanced chemiluminescence system (GE Healthcare).
Real-time RT–PCR
A detailed description is given in Supplementary Information. In brief, total RNA was isolated with an RNeasy extraction kit (Qiagen), reverse transcription was performed with Superscript II enzyme (Invitrogen), primers were designed with Primer Express software (Applied Biosystems) and real-time PCR was conducted with the SYBR green detection method on an ABI PRISM 7000 instrument (Applied Biosystems).
siRNA
Two siRNAs (27-mers, synthesized by MWG Biotech) were designed for each mRNA, pooled or used separated and transfected at 5 μM each. Control siRNA was used at equimolar levels. Transfections were performed with Lipofectamine 2000 (Invitrogen). siRNA sequences were as follows: GABAAR β3-(siRNA1), 5′-GCUACACUACGGAUGACAUUGAAUUUU-3′; GABAAR β3-(siRNA2), 5′-AGAAUGAUCGUUCUAAGAGUGAAAUAA-3′; H2AX-(siRNA1), 5′-CAAACACCUAGAUACCAGCACAAGCCG-3′; H2AX-(siRNA2), 5′-GACAAGAGUUCUAUACCUGCCGCAGGC-3′; ATR-(siRNA1), 5′-AAAAGCAGUUCUCCAGGAAUAUAGAAA-3′ ATR-(siRNA2), 5′-CCAAAGACAGUUGUAUUGAAUUCCGCG-3′; control siRNA21, 5′-AAGCUGACCCUGAAGUUCAUCUGCACC-3′.
Target gene downregulation was confirmed by real-time RT–PCR (see Supplementary Information for primer sequences) and for H2AX also by western blotting with anti-H2AX antibody (diluted 1:2,000; Abcam). For analyses in vivo, zygotes were injected into the cytoplasm with 10 pl of 20 μM siRNA and transferred to pseudopregnant females. At day 3 the mice were killed; embryos were flushed out of the uteri and fixed in 4% paraformaldehyde.
Pharmacological, BrdU and potentiometric analyses
Chemical substances were prepared as stock solutions as follows: picrotoxin (Tocris) and bicuculline (C20H17NO6, catalogue no. 0130; Tocris) to 50 or 100 mM in dimethylsulphoxide (DMSO), respectively; muscimol (Tocris) to 100 mM in water, DiBAC4(3) (Molecular Probes) in DMSO to 40 mM, hydroxyurea (Sigma) to 50 mg ml-1 in water, and BrdU (Sigma) was freshly dissolved to 10 mM in water.
Picrotoxin, bicuculline and muscimol ere used at 50, 20 and 100 μM, respectively, on cultured cells; controls received vehicle. Bicuculline was used in all experiments as antagonist, unless picrotoxin is specifically stated. For in vivo experiments, bicuculline or muscimol was diluted in PBS and administered intraperitoneally to pregnant females to 4 mg kg-1, in lineage tracing experiments the dose was repeated every 8 h. BrdU and muscimol were administered to 3.5-day pregnant females and analysed 5 h later for effects of the agonist on BrdU incorporation in vivo. Cells were loaded with 200 nM, and blastocysts with 1 μM, DiBAC4(3) in PBS or phenol-red-free culture medium for at least 15 min before analysis. Hydroxyurea was used at a concentration of 8 μg ml-1 in cell culture medium, and BrdU was used in cell culture medium at a final concentration of 10 μM. For experiments in vivo, 50 mg kg-1 BrdU was diluted in PBS and administered intraperitoneally. ATM/DNA-PK inhibitor (catalogue no. 118500, Calbiochem) was used at 10 μM.
References
Zhu, L. & Skoultchi, A. I. Coordinating cell proliferation and differentiation. Curr. Opin. Genet. Dev. 11, 91–97 (2001)
Savatier, P., Huang, S., Szekely, L., Wiman, K. G. & Samarut, J. Contrasting patterns of retinoblastoma protein expression in mouse embryonic stem cells and embryonic fibroblasts. Oncogene 9, 809–818 (1994)
Burdon, T., Smith, A. & Savatier, P. Signalling, cell cycle and pluripotency in embryonic stem cells. Trends Cell Biol. 12, 432–438 (2002)
Bartek, J., Lukas, C. & Lukas, J. Checking on DNA damage in S phase. Nature Rev. Mol. Cell Biol. 5, 792–804 (2004)
Hjerling-Leffler, J. et al. The boundary cap: a source of neural crest stem cells that generate multiple sensory neuron subtypes. Development 132, 2623–2632 (2005)
McGann, L. E., Walterson, M. L. & Hogg, L. M. Light scattering and cell volumes in osmotically stressed and frozen-thawed cells. Cytometry 9, 33–38 (1988)
Fernandez-Capetillo, O., Lee, A., Nussenzweig, M. & Nussenzweig, A. H2AX: the histone guardian of the genome. DNA Repair (Amst.) 3, 959–967 (2004)
Stiff, T. et al. ATR-dependent phosphorylation and activation of ATM in response to UV treatment or replication fork stalling. EMBO J. 25, 5775–5782 (2006)
Altman, J. & Bayer, S. A. The development of the rat spinal cord. Adv. Anat. Embryol. Cell Biol. 85, 1–164 (1984)
Maro, G. S. et al. Neural crest boundary cap cells constitute a source of neuronal and glial cells of the PNS. Nature Neurosci. 7, 930–938 (2004)
Voiculescu, O. et al. Hindbrain patterning: Krox20 couples segmentation and specification of regional identity. Development 128, 4967–4978 (2001)
Voiculescu, O., Charnay, P. & Schneider-Maunoury, S. Expression pattern of a Krox-20/Cre knock-in allele in the developing hindbrain, bones, and peripheral nervous system. Genesis 26, 123–126 (2000)
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001)
Fernandez-Capetillo, O. et al. H2AX is required for chromatin remodeling and inactivation of sex chromosomes in male mouse meiosis. Dev. Cell 4, 497–508 (2003)
Hammond, E. M., Dorie, M. J. & Giaccia, A. J. Inhibition of ATR leads to increased sensitivity to hypoxia/reoxygenation. Cancer Res. 64, 6556–6562 (2004)
McManus, K. J. & Hendzel, M. J. ATM-dependent DNA damage-independent mitotic phosphorylation of H2AX in normally growing mammalian cells. Mol. Biol. Cell 16, 5013–5025 (2005)
Ichijima, Y. et al. Phosphorylation of histone H2AX at M phase in human cells without DNA damage response. Biochem. Biophys. Res. Commun. 336, 807–812 (2005)
Liu, X., Wang, Q., Haydar, T. F. & Bordey, A. Nonsynaptic GABA signaling in postnatal subventricular zone controls proliferation of GFAP-expressing progenitors. Nature Neurosci. 8, 1179–1187 (2005)
Ying, Q. L., Nichols, J., Chambers, I. & Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 115, 281–292 (2003)
Lundgren, T. K., Scott, R. P., Smith, M., Pawson, T. & Ernfors, P. Engineering the recruitment of phosphotyrosine binding domain-containing adaptor proteins reveals distinct roles for RET receptor-mediated cell survival. J. Biol. Chem. 281, 29886–29896 (2006)
Kim, D. H. et al. Synthetic dsRNA Dicer substrates enhance RNAi potency and efficacy. Nature Biotechnol. 23, 222–226 (2005)
Acknowledgements
We thank T. Harkany and O. Kiehn for discussions, B. Wester for help with flow cytometry, and J. O. Bratteng for embryo injections. This work was supported by the Swedish Research Council (P.E., C.F.I. and A.E.M), the Swedish foundation for strategic research, CEDB and DBRM grants, Swedish Cancer Foundation, the Swedish Brain Foundation and Bertil Hållsten Research Foundation (P.E.), the Marianne and Marcus Wallenberg Foundation (C.F.I.) and the Karolinska Institutet (A.E.M.). M.A. was supported by grants from the Karolinska Institutet and the Swedish Brain Foundation. G.C.-B. was supported by the Swedish Brain Foundation.
Author Contributions M.A. was the main contributor to the work on ES cells; J.H.-L. was the main contributor to the work on NCS cells.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-9 with Legends, Supplementary Materials and Methods and additional references. (PDF 3542 kb)
Rights and permissions
About this article
Cite this article
Andäng, M., Hjerling-Leffler, J., Moliner, A. et al. Histone H2AX-dependent GABAA receptor regulation of stem cell proliferation. Nature 451, 460–464 (2008). https://doi.org/10.1038/nature06488
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06488
This article is cited by
-
DNA damage signalling histone H2AX is required for tumour growth
Cell Death Discovery (2024)
-
Identification of Nanog as a novel inhibitor of Rad51
Cell Death & Disease (2022)
-
Single cell transcriptomics reveals the heterogeneity of the human cornea to identify novel markers of the limbus and stroma
Scientific Reports (2021)
-
Endoplasmic reticulum retention and degradation of a mutation in SLC6A1 associated with epilepsy and autism
Molecular Brain (2020)
-
GABA inhibits proliferation and self-renewal of mouse retinal progenitor cell
Cell Death Discovery (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.