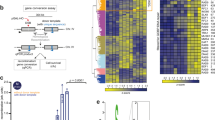
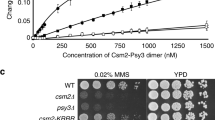
Abstract
In the S and G2 phases of the cell cycle, DNA double-strand breaks (DSBs) are processed into single-stranded DNA, triggering ATR-dependent checkpoint signalling and DSB repair by homologous recombination. Previous work has implicated the MRE11 complex in such DSB-processing events. Here, we show that the human CtIP (RBBP8) protein confers resistance to DSB-inducing agents and is recruited to DSBs exclusively in the S and G2 cell-cycle phases. Moreover, we reveal that CtIP is required for DSB resection, and thereby for recruitment of replication protein A (RPA) and the protein kinase ATR to DSBs, and for the ensuing ATR activation. Furthermore, we establish that CtIP physically and functionally interacts with the MRE11 complex, and that both CtIP and MRE11 are required for efficient homologous recombination. Finally, we reveal that CtIP has sequence homology with Sae2, which is involved in MRE11-dependent DSB processing in yeast. These findings establish evolutionarily conserved roles for CtIP-like proteins in controlling DSB resection, checkpoint signalling and homologous recombination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Wyman, C. & Kanaar, R. DNA double-strand break repair: all's well that ends well. Annu. Rev. Genet. 40, 363–383 (2006)
Lieber, M. R., Ma, Y., Pannicke, U. & Schwarz, K. Mechanism and regulation of human non-homologous DNA end-joining. Nature Rev. Mol. Cell Biol. 4, 712–720 (2003)
West, S. C. Molecular views of recombination proteins and their control. Nature Rev. Mol. Cell Biol. 4, 435–445 (2003)
Sung, P. & Klein, H. Mechanism of homologous recombination: mediators and helicases take on regulatory functions. Nature Rev. Mol. Cell Biol. 7, 739–750 (2006)
Zou, L. & Elledge, S. J. Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science 300, 1542–1548 (2003)
Ira, G. et al. DNA end resection, homologous recombination and DNA damage checkpoint activation require CDK1. Nature 431, 1011–1017 (2004)
Jazayeri, A. et al. ATM- and cell cycle-dependent regulation of ATR in response to DNA double-strand breaks. Nature Cell Biol. 8, 37–45 (2006)
Aylon, Y., Liefshitz, B. & Kupiec, M. The CDK regulates repair of double-strand breaks by homologous recombination during the cell cycle. EMBO J. 23, 4868–4875 (2004)
Lavin, M. F. The Mre11 complex and ATM: a two-way functional interaction in recognising and signaling DNA double strand breaks. DNA Repair (Amst.) 3, 1515–1520 (2004)
D'Amours, D. & Jackson, S. P. The Mre11 complex: at the crossroads of DNA repair and checkpoint signalling. Nature Rev. Mol. Cell Biol. 3, 317–327 (2002)
Schaeper, U., Subramanian, T., Lim, L., Boyd, J. M. & Chinnadurai, G. Interaction between a cellular protein that binds to the C-terminal region of adenovirus E1A (CtBP) and a novel cellular protein is disrupted by E1A through a conserved PLDLS motif. J. Biol. Chem. 273, 8549–8552 (1998)
Fusco, C., Reymond, A. & Zervos, A. S. Molecular cloning and characterization of a novel retinoblastoma-binding protein. Genomics 51, 351–358 (1998)
Wong, A. K. C. et al. Characterization of a carboxy-terminal BRCA1 interacting protein. Oncogene 17, 2279–2285 (1998)
Yu, X., Wu, L. C., Bowcock, A. M., Aronheim, A. & Baer, R. The C-terminal (BRCT) domains of BRCA1 interact in vivo with CtIP, a protein implicated in the CtBP pathway of transcriptional repression. J. Biol. Chem. 273, 25388–25392 (1998)
Greenberg, R. A. et al. Multifactorial contributions to an acute DNA damage response by BRCA1/BARD1-containing complexes. Genes Dev. 20, 34–46 (2006)
Yu, X. & Chen, J. DNA damage-induced cell cycle checkpoint control requires CtIP, a phosphorylation-dependent binding partner of BRCA1 C-terminal domains. Mol. Cell. Biol. 24, 9478–9486 (2004)
Yu, X., Fu, S., Lai, M., Baer, R. & Chen, J. BRCA1 ubiquitinates its phosphorylation-dependent binding partner CtIP. Genes Dev. 20, 1721–1726 (2006)
McKee, A. H. & Kleckner, N. A general method for identifying recessive diploid-specific mutations in Saccharomyces cerevisiae, its application to the isolation of mutants blocked at intermediate stages of meiotic prophase and characterization of a new gene SAE2 . Genetics 146, 797–816 (1997)
Prinz, S., Amon, A. & Klein, F. Isolation of COM1, a new gene required to complete meiotic double-strand break-induced recombination in Saccharomyces cerevisiae . Genetics 146, 781–795 (1997)
Rattray, A. J., McGill, C. B., Shafer, B. K. & Strathern, J. N. Fidelity of mitotic double-strand-break repair in Saccharomyces cerevisiae: a role for SAE2/COM1 . Genetics 158, 109–122 (2001)
Lobachev, K. S., Gordenin, D. A. & Resnick, M. A. The Mre11 complex is required for repair of hairpin-capped double-strand breaks and prevention of chromosome rearrangements. Cell 108, 183–193 (2002)
Clerici, M., Mantiero, D., Lucchini, G. & Longhese, M. P. The S. cerevisiae Sae2 protein promotes resection and bridging of double-strand break ends. J. Biol. Chem. 280, 38631–38638 (2005)
D'Arpa, P., Beardmore, C. & Liu, L. F. Involvement of nucleic acid synthesis in cell killing mechanisms of topoisomerase poisons. Cancer Res. 50, 6919–6924 (1990)
Saleh-Gohari, N. et al. Spontaneous homologous recombination is induced by collapsed replication forks that are caused by endogenous DNA single-strand breaks. Mol. Cell. Biol. 25, 7158–7169 (2005)
Shiloh, Y. ATM and related protein kinases: safeguarding genome integrity. Nature Rev. Cancer 3, 155–168 (2003)
Bakkenist, C. J. & Kastan, M. B. Initiating cellular stress responses. Cell 118, 9–17 (2004)
Cuadrado, M. et al. ATM regulates ATR chromatin loading in response to DNA double-strand breaks. J. Exp. Med. 203, 297–303 (2006)
Shao, R.-G. et al. Replication-mediated DNA damage by camptothecin induces phosphorylation of RPA by DNA-dependent protein kinase and dissociates RPA:DNA-PK complexes. EMBO J. 18, 1397–1406 (1999)
Sakasai, R. et al. Differential involvement of phosphatidylinositol 3-kinase-related protein kinases in hyperphosphorylation of replication protein A2 in response to replication-mediated DNA double-strand breaks. Genes Cells 11, 237–246 (2006)
Lukas, C., Falck, J., Bartkova, J., Bartek, J. & Lukas, J. Distinct spatiotemporal dynamics of mammalian checkpoint regulators induced by DNA damage. Nature Cell Biol. 5, 255–260 (2003)
Bekker-Jensen, S. et al. Spatial organization of the mammalian genome surveillance machinery in response to DNA strand breaks. J. Cell Biol. 173, 195–206 (2006)
Ball, H. L., Myers, J. S. & Cortez, D. ATRIP binding to RPA-ssDNA promotes ATR–ATRIP localization but is dispensable for Chk1 phosphorylation. Mol. Biol. Cell 16, 2372–2381 (2005)
Adams, K. E., Medhurst, A. L., Dart, D. A. & Lakin, N. D. Recruitment of ATR to sites of ionising radiation-induced DNA damage requires ATM and components of the MRN protein complex. Oncogene 25, 3894–3904 (2006)
Myers, J. S. & Cortez, D. Rapid activation of ATR by ionizing radiation requires ATM and Mre11. J. Biol. Chem. 281, 9346–9350 (2006)
Paull, T. T. & Gellert, M. The 3′ to 5′ exonuclease activity of Mre11 facilitates repair of DNA double-strand breaks. Mol. Cell 1, 969–979 (1998)
Assenmacher, N. & Hopfner, K. P. MRE11/RAD50/NBS1: complex activities. Chromosoma 113, 157–166 (2004)
Larson, E. D., Cummings, W. J., Bednarski, D. W. & Maizels, N. MRE11/RAD50 cleaves DNA in the AID/UNG-Dependent Pathway of immunoglobulin gene diversification. Mol. Cell 20, 367–375 (2005)
Trujillo, K. M., Yuan, S.-S. F., Lee, E.-Y. P. & Sung, P. Nuclease activities in a complex of human recombination and DNA repair factors Rad50, Mre11, and p95. J. Biol. Chem. 273, 21447–21450 (1998)
Pierce, A. J., Hu, P., Han, M., Ellis, N. & Jasin, M. Ku DNA end-binding protein modulates homologous repair of double-strand breaks in mammalian cells. Genes Dev. 15, 3237–3242 (2001)
Schaffer, A. A. et al. Improving the accuracy of PSI-BLAST protein database searches with composition-based statistics and other refinements. Nucleic Acids Res. 29, 2994–3005 (2001)
Deng, C., Brown, J. A., You, D. & Brown, J. M. Multiple endonucleases function to repair covalent topoisomerase I complexes in Saccharomyces cerevisiae . Genetics 170, 591–600 (2005)
Lisby, M., Barlow, J. H., Burgess, R. C. & Rothstein, R. Choreography of the DNA damage response: spatiotemporal relationships among checkpoint and repair proteins. Cell 118, 699–713 (2004)
Li, S. et al. Functional link of BRCA1 and ataxia telangiectasia gene product in DNA damage response. Nature 13, 210–215 (2000)
Baroni, E., Viscardi, V., Cartagena-Lirola, H., Lucchini, G. & Longhese, M. P. The functions of budding yeast Sae2 in the DNA damage response require Mec1- and Tel1-dependent phosphorylation. Mol. Cell. Biol. 24, 4151–4165 (2004)
Cartagena-Lirola, H., Guerini, I., Viscardi, V., Lucchini, G. & Longhese, M. P. Budding yeast Sae2 is an in vivo target of the Mec1 and Tel1 checkpoint kinases during meiosis. Cell Cycle 5, 1549–1559 (2006)
Vilkki, S. et al. Screening for microsatellite instability target genes in colorectal cancers. J. Med. Genet. 39, 785–789 (2002)
Wu, G. & Lee, W. H. CtIP, a multivalent adaptor connecting transcriptional regulation, checkpoint control and tumor suppression. Cell Cycle 5, 1592–1596 (2006)
Raderschall, E., Golub, E. I. & Haaf, T. Nuclear foci of mammalian recombination proteins are located at single-stranded DNA regions formed after DNA damage. Proc. Natl Acad. Sci. USA 96, 1921–1926 (1999)
Wu-Baer, F., Lagrazon, K., Yuan, W. & Baer, R. The BRCA1/BARD1 heterodimer assembles polyubiquitin chains through an unconventional linkage involving lysine residue K6 of ubiquitin. J. Biol. Chem. 278, 34743–34746 (2003)
Acknowledgements
We thank T. Paull and J. Falck for providing reagents, and P. Huertas, A. Meier, K. Dry, K. Miller and Y. Pommier for advice and critical reading of the manuscript. This study was supported by Cancer Research UK, the EU and a Swiss National Science Foundation fellowship for advanced researcher (A.A.S). Research in the S.P.J. laboratory is made possible by core infrastructure funding from Cancer Research UK and the Wellcome Trust. C.L., J.L., M.M. and J.B. were supported by grants from the Danish Cancer Society, Danish National Research Foundation, EU (DNA Repair), and the John and Birthe Meyer Foundation and the Czech Ministry of Education. Research in the laboratory of R.B. is supported by the National Institute of Health and S.F. is supported by a fellowship from the New York State Breast Cancer Research Fund.
Author Contributions S.F. and R.B. generated CtIP cDNA, CtIP antibodies and recombinant CtIP protein. C.L., J.L., M.M. and J.B. generated the cell lines with GFP-tagged proteins, conceived, performed and evaluated the real-time imaging experiments, and performed the homologous recombination measurements. All other experiments were conceived by A.A.S. and S.P.J., and were performed by A.A.S. with the help of J.C. A.A.S. and S.P.J. wrote the paper. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Methods with additional references and Supplementary Figures S1-S6 with Legends. (PDF 9614 kb)
Rights and permissions
About this article
Cite this article
Sartori, A., Lukas, C., Coates, J. et al. Human CtIP promotes DNA end resection. Nature 450, 509–514 (2007). https://doi.org/10.1038/nature06337
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06337
This article is cited by
-
CDK9-55 guides the anaphase-promoting complex/cyclosome (APC/C) in choosing the DNA repair pathway choice
Oncogene (2024)
-
Modulation of cell cycle increases CRISPR-mediated homology-directed DNA repair
Cell & Bioscience (2023)
-
Recurrence mutation in RBBP8 gene causing non-syndromic autosomal recessive primary microcephaly; geometric simulation approach for insight into predicted computational models
Journal of Human Genetics (2023)
-
CRISPR screen identifies the role of RBBP8 in mediating unfolded protein response induced liver damage through regulating protein synthesis
Cell Death & Disease (2023)
-
Genome-wide analysis of DNA-PK-bound MRN cleavage products supports a sequential model of DSB repair pathway choice
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.