Abstract
A conserved network of eye field transcription factors (EFTFs) underlies the development of the eye in vertebrates and invertebrates1. To direct eye development, Pax6, a key gene in this network2,3, interacts with genes encoding other EFTFs such as Rx1 and Six3 (refs 4–6). However, the mechanisms that control expression of the EFTFs remain unclear7. Here we show that purine-mediated signalling triggers both EFTF expression and eye development in Xenopus laevis. Overexpression of ectonucleoside triphosphate diphosphohydrolase 2 (E-NTPDase2)8, an ectoenzyme that converts ATP to ADP9, caused ectopic eye-like structures, with occasional complete duplication of the eye, and increased expression of Pax6, Rx1 and Six3. In contrast, downregulation of endogenous E-NTPDase2 decreased Rx1 and Pax6 expression. E-NTPDase2 therefore acts upstream of these EFTFs. To test whether ADP (the product of E-NTPDase2) might act to trigger eye development through P2Y1 receptors, selective in Xenopus for ADP10,11, we simultaneously knocked down expression of the genes encoding E-NTPDase2 and the P2Y1 receptor. This could prevent the expression of Rx1 and Pax6 and eye formation completely. We next measured ATP release12,13,14 in the presumptive eye field, demonstrating a transient release of ATP at a time that could plausibly trigger (once converted to ADP) expression of the EFTFs. This surprising role for transient purine-mediated signalling in eye development may be widely conserved, because alterations to the locus of E-NTPDase2 on human chromosome 9 cause severe head and eye defects, including microphthalmia15,16,17,18. Our results suggest a new mechanism for the initiation of eye development.
Similar content being viewed by others
Main
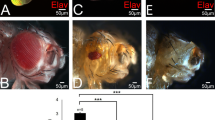
To assess the developmental functions of the E-NTPDases, we simultaneously injected the mRNA for the closely related E-NTPDases1–3 (ref. 8) with lineage tracer into a dorsal animal blastomere at the eight-cell stage to target overexpression of this gene to one side of the nervous system. Overexpression of E-NTPDase2 affected eye development in 27 of 41 embryos, causing in some cases complete duplication of the eye on the injected side (Fig. 1A, a). In contrast, overexpression of E-NTPDase1 decreased eye size (11 of 44 embryos; Fig. 1A, b), whereas overexpression of E-NTPDase3 gave a weaker phenotype somewhat similar to that of E-NTPDase2 (4 of 42 embryos, Fig. 1A, c, Supplementary Tables 1a and 2). E-NTPDases differ in their catalytic activity. Like their mammalian orthologues, E-NTPDase1 can metabolize ATP and ADP with roughly equal efficacy, E-NTPDase2 is highly selective for ATP and hardly metabolizes ADP, and E-NTPDase3 has intermediate selectivity for ATP and ADP (Supplementary Fig. 2). The phenotypes elicited by overexpression of these membrane-bound E-NTPDases correlate with their capacity to metabolize ADP.
A, Overexpression of E-NTPDase genes causes multiple eye phenotypes. D1 (a–i) or V1 blastomeres (j–l) were injected with Xenopus E-NTPDase (NTPD) 1, 2 or 3 mRNAs. E-NTPDase2 caused ectopic eyes (black arrows in a, d and e), occasionally with a lens (g), abnormal eyes (d) or RPE extensions (white dotted arrows, f). Pigmented retina (black arrows) and pineal eye (open arrow) (compare i with h) were immunolabelled with anti-RPE XAR-1. Ventral injection of E-NTPDase2 gave similar ectopic RPE (arrow) (j), sometimes far from the head (k, l). E-NTPDase1 caused a small-eye phenotype (b). E-NTPDase3 caused a mild ectopic-eye phenotype (arrow, c). B, The ectopic eyes exhibit a layered cellular structure. Ectopic eyes (b and c) of stage-48 embryos show typical organization of retinal layers as in normal eyes (a), associated with lenses in some cases (c). Scale bars, 30 μm. CMZ, ciliary margin zone; G, ganglion cell layer; I, inner nuclear layer; L, lens; O, outer nuclear layer; P, retinal pigmented layer.
The eye phenotypes caused by overexpression of E-NTPDase2 (Supplementary Table 1b) included the following: disrupted eye development (Fig. 1A, d); ectopic retinal pigment epithelium (RPE) (Fig. 1A, e); RPE extensions (Fig. 1A, f); and ectopic RPE with an apparent lens (Fig. 1A, g). The antibodies XAR-1, a monoclonal anti-RPE (compare Fig. 1A, i with Fig. 1A, h), rhodopsin and the neural marker 2G9 stained the ectopic structures (Supplementary Fig. 3), showing that overexpression of E-NTPDase2 caused the formation of ectopic retinal tissue. Histological sections through the ectopic eyes (Fig. 1B, b and c) showed a layered structure very similar to normal eyes (Fig. 1B, a). This range of eye phenotypes bears a striking resemblance to that reported after overexpression of the paired transcription factor, Pax6 (ref. 19).
Embryos injected in a ventral animal blastomere, fated to give non-neural ectoderm, exhibited ectopic RPE development (8 of 18 embryos; Supplementary Table 1b), usually close to the otic vesicle, but also in the abdomen and along the tail (Fig. 1A, j and k), also positively identified by staining with XAR-1 (Fig. 1A, l). Overexpression of E-NTPDase2 can therefore recruit a cascade of gene expression sufficient to enable eye formation well away from the head structures and the endogenous eye field.
The genes Rx1 (ref. 5), Pax6 (refs 2, 3) and Six3 (ref. 6) are key members of the EFTF network4. To determine whether the actions of E-NTPDase2 lie upstream or downstream of the transcription factors encoded by these genes, we analysed their expression by in situ hybridization in embryos overexpressing E-NTPDase2 and/or green fluorescent protein (GFP) at stage 13, just after the specification of the eye field20. Abnormal expression of Rx1, Pax6 and Six3 was observed with ectopic patches of staining on the injected side in 21 of 34, 13 of 28, and 14 of 25 embryos, respectively (Fig. 2A, a–c). Rx1, Pax6 and Six3 expression was not perturbed on the uninjected side or in control GFP-injected embryos (Supplementary Table 3).
A, Embryos injected in the D1 blastomere with E-NTPDase2 mRNA were cultured to stage 13. Ectopic expression (arrow) of the EFTF-encoding genes Pax6, Rx1 and Six3 (a–c) occurred on the injected side (Inj.). The eye field was enlarged on the injected side, as also seen with Otx2 and Sox3 expression (open arrow in d and e). B, Embryos injected in the D1 blastomere with mRNAs and MOs were cultured to stage 13. Knockdown of E-NTPDase2 (NTPD2 MO) decreased Pax6 expression on the injected side (inj.) in both anterior (open arrow) and posterior (white arrow) domains (b). Control MO (CMO) had no effect on Pax6 expression (a). Mouse E-NTPDase2 (mNTPD2) caused ectopic expression of Pax6 (c, black arrow). E-NTPDase2 MO simultaneously injected with mNTPD2 mRNA rescued Pax6 expression (compare d with b). E-NTPDase1 (NTPD1) alone had no significant effect (e), whereas E-NTPDase3 (NTPD3) alone caused loss of the posterior Pax6 expression (white arrow) and ectopic Pax6 (black arrow) (g). E-NTPDase3, but not E-NTPDase1, was able to rescue the E-NTPDase2 MO phenotype (compare h and f with b). C, Cumulative probability distributions for the effects of control, E-NTPDase2 MOs, alone or simultaneously injected with E-NTPDase mRNAs on the anterior domain of Pax6 expression. Left: injection of E-NTPDase2 MO substantially decreased the area of Pax6 expression (leftward shift). Simultaneous injection of mE-NTPDase2 mRNA partly rescued this phenotype (shift back to right). Right: simultaneous injection of E-NTPDase3 mRNA slightly lessened the phenotype caused by E-NTPDase2 MO, whereas simultaneous injection of E-NTPDase1 mRNA had little effect.
Otx2 is a homeodomain transcription factor involved during eye development21. The expression of Otx2 is downregulated at stage 12.5, to reveal an expression pattern that delineates the eye field4. E-NTPDase2 overexpression altered Otx2 expression on the injected side and gave an expansion of the eye field (11 of 23 embryos; Fig. 2A, d). A similar effect was observed with the neural marker Sox3 (ref. 22) (13 of 27 embryos; Fig. 2A, e). Analysis at stage 23 confirms that ectopic expression for Otx2 and Sox3 was observed on the injected side, as was also seen for Pax6 (Supplementary Fig. 4), indicating an effect on formation of the nervous system. This effect was further confirmed by analysis of embryos at stage 45, which showed duplication of the pineal eye after overexpression of E-NTPDase2 in a small number of cases (Supplementary Fig. 4).
To test whether endogenous E-NTPDase2 is necessary for Pax6 expression, we knocked down its expression with a specific antisense morpholino oligonucleotide (MO) (Supplementary Fig. 5). Injection of a random control MO (CMO) had no effect on Pax6 expression at stage 13 (Fig. 2B, a). However, injection of E-NTPDase2 MO greatly reduced Pax6 expression in 45 of 48 embryos (P < 0.0001 versus CMO; Fig. 2B, b, C). This effect of the E-NTPDase2 MO was partly rescued by the simultaneous injection of mouse E-NTPDase2 mRNA (P = 0.016; Fig. 2B, d, C), the translation of which was unaffected by the Xenopus-specific E-NTPDase2 MO (Supplementary Fig. 5a, b) and which when injected by itself enhanced Pax6 expression (Fig. 2B, c) and caused ectopic eye phenotype (Supplementary Fig. 6 and Supplementary Table 1c).
Overexpression of E-NTPDase1 had no consistent effect on Pax6 expression (Fig. 2B, e, and Supplementary Table 4). However, if E-NTPDase1 mRNA was simultaneously injected with E-NTPDase2 MO, Pax6 expression was in some cases almost abolished (Fig. 2B, f, C). Overexpression of E-NTPDase1, which can efficiently metabolize ATP and ADP to AMP, will probably exacerbate the effect of knockdown of E-NTPDase2 activity and consequently the decreased production of extracellular ADP. Overexpression of E-NTPDase3 enhanced Pax6 expression (Fig. 2B, g, and Supplementary Table 4) and partly rescued the effects of the E-NTPDase2 MO (P = 0.039; Fig. 2B, h, C).
E-NTPDase2 seems to be involved in the earliest phase of eye development and acts upstream of the EFTFs encoded by Pax6, Rx1 and Six3. As overexpression of E-NTPDases1–3 yields a range of eye phenotypes that correlates with their enzymatic activity towards ADP, we proposed that signalling by ADP could be critical in activating expression of the EFTFs. Xenopus possesses a P2Y1 receptor that is selective for ADP10,11. We therefore directly manipulated the expression levels of the P2Y1 receptor with a specific MO (Supplementary Fig. 7), either on its own or in combination with E-NTPDase2.
Knockdown of E-NTPDase2 and the P2Y1 receptor had synergistic effects on EFTF expression at stage 13 (Fig. 3A, B). The P2Y1 MO alone had no significant effect on Pax6 and Rx1 expression (Fig. 3A, c and g). The E-NTPDase2 MO very significantly decreased Pax6 and Rx1 expression (P = 0.012 and P = 0.04 versus CMO, respectively; Fig. 3A, b and f). Simultaneous injection of both MOs almost eliminated Pax6 and Rx1 expression (P < 0.0001and P = 0.001, respectively; Fig. 3A, a and e, B). Injection of the E-NTPDase2 and P2Y1 MOs, either singly or in combination, had no significant effect on the Sox3 expression that delimits the eye field; however, they did reduce the anterior neural stripe (Fig. 3A, i and j). The CMO had no effect on any of the EFTFs (Fig. 3A, d, h and l).
A, MOs were injected into the D1 blastomere. Knockdown of E-NTPDase2 (NTPD2 MO) decreased Pax6 (b) and Rx1 (f) staining on the injected side (Inj., open arrow). Knockdown of both E-NTPDase2 and P2Y1 induced a stronger phenotype (a, e). Microinjection of both MOs or E-NTPDase2 MO alone resulted in the loss of a stripe of Sox3 expression (i, j) (filled arrow) but there was little effect on the eye field (open arrow). Microinjection of P2Y1 MO alone (c, g, k) or CMO had no effect on the expression of either marker (d, h, l). B, Cumulative probability distributions for the effects of control, P2Y1, E-NTPDase2 and combined MOs on Pax6, Rx1 and Sox3 expression. Injection of both E-NTPDase2 MO and P2Y1 MO resulted in a decrease in the area of Pax6 (anterior domain) and Rx1 expression on the injected side (leftward shift). This was greater than the phenotype seen with E-NTPDase2 MO alone. P2Y1 MO alone had no significant effect on Pax6 and Rx1 expression. There was no significant effect of any of the MOs, either alone or in combination, on Sox3 expression.
We next examined the effects of altering P2Y1 and E-NTPDase2 expression on eye formation. Overexpression of the P2Y1 receptor gave a weak phenotype (Fig. 4A, a) in comparison with overexpression of E-NTPDase2 (Fig. 4A, b). However, when both the P2Y1 receptor and E-NTPDase2 were overexpressed, the resulting ectopic eye phenotype was greatly exaggerated (Fig. 4A, c). Morpholino knockdown of P2Y1 expression decreased eye size in comparison with control in about 60% of embryos (P = 0.057 versus CMO; Fig. 4B, a, D). Knockdown of E-NTPDase2 gave a similar phenotype in a similar proportion of embryos P = 0.048 versus CMO; (Fig. 4B, b, D). However, simultaneous injection of both P2Y1 and E-NTPDase2 MO resulted in a markedly enhanced phenotype at stage 45, characterized by either a very small eye or a complete absence of the eye on the injected side (P = 0.001 versus CMO; Fig. 4B, c, C, D).
A, D1 blastomere-injected E-NTPDase2 (NTPD2) caused the formation of ectopic eyes (b, dotted arrow). On rare occasions, P2Y1 exhibited RPE extensions (a, arrow). Simultaneous injection of E-NTPDase2 with P2Y1 caused multiple larger ectopic eye-like structures (dotted arrow) coupled with RPE extension (arrow) from the endogenous eye (c). B, Combined E-NTPDase2 MO and P2Y1 MO knockdown ablates eye structures. D1 blastomere knockdown of E-NTPDase2 (NTPD2 MO) caused a smaller but fully normal eye (b) on the injected side. A small effect on phenotype was also seen, by measurement, with P2Y1 MO alone (a). Simultaneous knockdown of E-NTPDase2 and P2Y1 could prevent eye development (c, dotted arrow). C, Transverse section of the embryo shown in Fig. 3B, c showing a complete loss of eye (dotted arrow) on the injected side. Scale bars, 30 μm. ey, eye; hb, hindbrain; ph, pharynx. D, Cumulative probability distributions for the effects of control MO (CMO), P2Y1 MO, E-NTPDase2 MO and combined MOs on eye development measured at stage 45. Comparison with the CMO shows that about 30% of the injected embryos showed no eye phenotype. In the remainder, injection with P2Y1 MO and E-NTPDase2 MO together resulted in much smaller eyes than injection with CMO.
Purinergic signalling may thus lie upstream of the EFTFs and be responsible for triggering the expression of these factors. It should therefore be possible to rescue the effects of knockdown of E-NTPDase2 and P2Y1 receptor expression by overexpressing Pax6. Injection of the E-NTPDase2 and P2Y1 MOs resulted in a substantial decrease in eye size (P < 0.0001 versus CMO; Fig. 5a). Injection of Pax6 mRNA by itself also gave a typical eye phenotype, as reported by others19 (P < 0.0001 versus CMO; Fig. 5c). However, when the two MOs and Pax6 mRNA were injected simultaneously, the morphology of the eye on the injected side could be remarkably normal, a phenotype significantly different from those of the double MO phenotype and the control, indicating partial rescue (P = 0.041 and P = 0.028, respectively; Fig. 5b and Supplementary Fig. 8).
The small-eye phenotype at stage 45 induced by simultaneous knockdown of E-NTPDase2 and P2Y1 receptor (NTPDase2 MO + P2Y1 MO injection through blastomere D1) (a) can be rescued by the simultaneous injection of Pax6 mRNA (b). Pax6 overexpression alone (through D1 blastomere) caused ectopic RPE structures (solid arrow) and RPE extensions (c)19 also occasionally seen in b (dotted arrows).
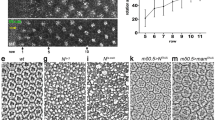
Purinergic signalling by ADP, generated through the actions of E-NTPDase2 and acting through P2Y1 receptors, seems to activate, either directly or indirectly, Pax6 and thus eye formation. Accordingly, the endogenous expression of E-NTPDase2 and P2Y1 genes should precede or coincide with the expression of Pax6 in the eye field. We therefore investigated, by reverse-transcriptase-mediated polymerase chain reaction (RT–PCR), the early expression of these genes in the anterior and posterior neural plate and in the ectodermal or mesodermal/endodermal tissue layers. Expression of E-NTPDase2 becomes detectable in the anterior neural plate at stage 10.5 and thus precedes specification of the eye field20, characterized by the upregulation of expression of Pax6 (ref. 4) (Fig. 6a). Furthermore, expression of E-NTPDase2 is predominantly in the mesodermal/endodermal layer (Fig. 6b). The P2Y1 gene is expressed preferentially in the anterior neural plate from stage 12.5 and occurs only in the mesoderm/endoderm (Fig. 6a, b). These data indicate that the mesoderm/endoderm tissue is the source of purinergic signalling.
a, E-NTPDase2 and P2Y1 are expressed in the eye field. E-NTPDase2 transcripts were detected from stage (St.) 10.5 in the presumptive anterior neural plate (ANP) and posterior neural plate (PNP), whereas Pax6 was detected later, from stage 11.5. E-NTPDase3 was first expressed at stage 13 in the ANP but E-NTPDase1 was not detected at any stage (data not shown). P2Y1 (ANP > PNP) and P2Y8 (ANP = PNP) mRNAs were detected from stage 12.5. Otx2 demonstrated the accuracy of the dissections, and ODC was the loading control. AC, animal cap; W, whole embryo. b, E-NTPDase2 and P2Y1 are expressed in the mesodermal layer. E-NTPDase2 was expressed at higher levels in dissected mesodermal/endodermal (Mes./End.) layers than in the ectoderm (Ect.). P2Y1 was only detected in mesoderm/endoderm dissection. Otx2, Rx1 and Pax6 were expressed in both fractions. Xbra3 and cytokeratin demonstrated the accuracy of the dissections. EF1α was the loading control. c, ATP is released in the neural plate during development. The placement of biosensors is indicated in the diagram. A single large transient event was seen on the ATP biosensor inserted into the anterior neural plate. This event had a rapid rise time and a slower decaying phase (inset) and was unaccompanied by any equivalent signal on the null or posterior ATP biosensors.
To investigate whether purinergic signalling could provide both spatial and temporal cues for the activation of Pax6 expression, we directly measured ATP release in the presumptive eye field by using selective biosensors12. A single event was recorded from ATP biosensors inserted into the anterior but not the posterior neural plate during eye-field development in 10 of 18 embryos (Fig. 6c). No events were recorded on the null biosensor, giving confidence that this signal represented ATP release (Methods). The timing of the event was correlated with the stage of the embryo at the start of the recording. The event had a mean rise time of 25 ± 4.4 s, a duration of 302 ± 53 s and an amplitude of 71 ± 33 pA, equivalent to 1.0 ± 0.4 μM ATP. Towards the end of the recording period, ATP release was also observed in the posterior neural plate, indicating that ATP signalling may regulate the development of the posterior neural plate.
We next tested whether pharmacological manipulation of purinergic signalling can either cause or rescue an eye phenotype. Pyridoxal-phosphate-6-azophenyl-2′,4′-disulphonate (PPADS), a P2 antagonist, injected into the blastocoel at 100 μM, decreased the ectopic expression of Pax6 caused by the overexpression of E-NTPDase2 (Supplementary Fig. 9a), and MRS2179, a more specific P2Y1 receptor antagonist, when applied similarly decreased the eye phenotype measured at stage 45 (Supplementary Fig. 9b). In the converse experiment, the agonist 2-methylthioadenosine-diphosphate (2MeSADP) was able to rescue the effects of the E-NTPDase2 MO on Pax6 expression (Supplementary Fig. 9a). We also applied the P2Y1 agonist 2-MeSADP by means of agarose beads soaked in a 1 mM solution. Beads inserted near the presumptive eye field at stage 12.5 occasionally produced RPE extensions (Supplementary Fig. 9c) typical of overexpression of E-NTPDase2 or Pax6.
Our data provide compelling evidence that a discrete purinergic signalling event localized to the anterior neural plate is critical for triggering the expression of EFTFs and hence initiating eye development (Supplementary Fig. 1). Because mutations to Pax6 can give rise to severe defects of the nervous system23,24,25, we expected similar alterations from manipulation of the purinergic signalling pathway upstream of Pax6. Accordingly, alteration of E-NTPDase2 expression affected both brain development (Supplementary Fig. 4) and Pax6 expression in the posterior neural plate (Fig. 2B, b)—a region destined to form the nervous system. Purinergic signalling thus orchestrates the development of both the anterior nervous system and the eye. E-NTPDase2 has been implicated in the differentiation of stem cells in the vertebrate nervous system26,27.
Human eye and brain development may also depend on E-NTPDase2 activity: spontaneous mutations at 9q34, the region bearing the locus of the E-NTPDase2 gene, consistently give head and brain abnormalities that include a range of eye defects including microphthalmia, analogous to the phenotypes we have reported in the frog15,16,17,18. Our studies raise several fascinating questions such as the identity of the ATP-releasing cells and the chain of events that connect activation of the P2Y1 receptor in the mesoderm/endoderm to initiation of EFTF expression in the ectoderm.
Methods Summary
Embryo culture dissection and microinjections
Xenopus laevis embryos were staged, cultured and dissected by standard methodology. All microinjections of mRNAs, morpholinos or combinations of both were performed into defined blastomeres at the eight-cell stage together with lineage tracer.
RT–PCR
RNAs were isolated and complementary DNAs were reverse-transcribed by following standard procedures from whole or dissected embryos. Non-radioactive PCRs were performed with gene-specific primers.
Whole-mount in situ hybridization, immunohistochemistry and sectioning
Experimentally manipulated embryos were hybridized to digoxigenin-labelled RNA probes produced in vitro from cDNA clones encoding eye and brain markers. Colour reactions were performed and the embryos were bleached before scoring. Immunohistochemically stained embryos were analysed by standard protocols and colour reactions.
Embryo measurement and statistical analysis
Areas of positive in situ hybridization were measured with image analysis software (LuciaG) and recorded. The dimensions of the eyes were measured along orthogonal axes and the product of these values was used as a proxy for volume. Differences between injected and uninjected sides were ranked and cumulative probability distributions were plotted. Statistical analysis was performed by Kruskall–Wallis analysis of variance, with pairwise comparisons by the Kolmogorov–Smirnov test (probability values for these comparisons are given in text where appropriate).
Biosensor recordings
ATP biosensors were used to record ATP release from embryos at the mid-gastrula stage. Null biosensors acted as negative controls and ATP sensors were calibrated at the end of each experiment.
Online Methods
DNA constructs
Full-length Xenopus E-NTPDase1, E-NTPDase2 and E-NTPDase3 cDNAs were isolated from IMAGE clones8, cloned into pBF. The IMAGE clones coding for mouse E-NTPDase2 cDNA (IMAGE clone 3582550, NCBI; accession number BC011241) and Xenopus P2Y1 cDNA (IMAGE clone 5507066, NCBI; accession number CF284315) were ordered from the UK MRC HGMP Resource Centre, and the full coding region was inserted into pBF.
Embryo culture, dissection and microinjections
Embryos, staged as described28, were obtained and cultured as described previously. Embryos at different stages were dissected in Barth X to isolate the animal cap (stage 9) and the anterior neural plate from the posterior neural plate (from stage 10.5 to stage 15). Fine dissection of the anterior neural plate of stage 12.5 embryos was performed to isolate the outer ectoderm from the inner mesodermal and endodermal layers. Capped synthetic mRNAs were generated by transcription in vitro from the expression clones by using the mMessage mMachine kit (Sp6 RNA polymerase; Ambion) and 5 ng was injected into each embryo. All microinjections were performed into a D1 or V1 blastomere (as indicated in the figure legends) of eight-cell-stage embryos. E-NTPDase2 (5′-gccatgttgggctcttgagtgcctc-3′) and P2Y1 (5′-gagagaaagacttctgtcatgatct-3′) MOs were designed and supplied by GeneTools. The MO (5 ng nl-1) was injected alone (20 ng) or in combination with mRNA or with another MO. The random CMO (GeneTools) was used as a control. Either GFP (4 ng) or β-galactosidase (LacZ) (2 ng) mRNA was simultaneously injected as a lineage tracer. X-Gal staining was performed as described29. Each experiment was performed on at least two independent batches of embryos; data presented and analysed in the text are from a representative batch.
RT–PCR
Total RNA from whole or dissected embryos was isolated and used for reverse transcription as described30. PCR was performed with non-radioactive nucleotides (Supplementary Table 5). For each experiment the quantity of input cDNA was determined by equalization of the samples with a housekeeping gene.
Whole-mount in situ hybridization and immunohistochemistry
Microinjected embryos were fixed in MEMFA (0.5 M MOPS pH 7.4, 100 mM EGTA, 1 mM MgSO4, 4% formaldehyde) and hybridized (see ref. 31) with antisense RNA probes produced in vitro from cDNA clones. A 800-base-pair fragment from Sox3 was cloned into pGEMT-easy, transcribed with Sp6 RNA polymerase and linearized with SacII. Clones containing the templates for Pax6, Rx1, Six3 and Otx2 were generated in accordance with published protocols4,32,33,34. The probe was labelled by using a digoxigenin labelling kit (Roche) and the hybridization was revealed with sheep anti-digoxigenin-alkaline phosphatase antibody (Roche) and 4-nitroblue tetrazolium chloride/5-bromo-4-chloro-3-indolylphosphate substrate (NBT/BCIP; Roche). Embryos were bleached by using standard protocols. Whole-mount immunohistochemistry was performed on injected embryos fixed in MEMFA and dehydrated in methanol. Embryos were rehydrated, bleached and immunostained by using standard protocols with the monoclonal anti-RPE antibody XAR-1. The colour reaction was performed with NBT/BCIP.
Embedding and sectioning
Wax sections (12 μm thick) were cut from stage-48 embryos35 and stained with 20% Harris haematoxylin solution (Sigma) and 1% eosin (BDH Laboratory Supplies). The sections were mounted with DePex (BDH Laboratory Supplies).
Measurement of embryos and statistical analysis
A Nikon SMZ1500 microscope and digital DXM1200F camera were used to photograph the embryos. The midline of each embryo was identified and the anterior domain of Pax6 staining, Rx1 domain or the eye field delimited by Sox3 expression on both injected and uninjected sides was measured with the software LuciaG. The difference in area was ranked and plotted as a cumulative probability distribution. The size of eyes of stage-45 embryos was measured along the anterior–posterior and medio-lateral axes by using LuciaG. The product of these two measurements was calculated (in mm2) and used as a proxy statistic for eye volume. The decrease in eye size between the injected and uninjected sides was ranked and plotted as a cumulative probability distribution. The G-test was used to compare the frequency of the phenotypes evoked by each experimental manipulation (Supplementary Table 2). A combination of the Kruskal–Wallis one-way analysis of variance and pairwise comparisons performed with the Kolmogorov–Smirnov two-sample test was used to analyse the cumulative probability distributions presented in Figs 2–4 and Supplementary Fig. 8 (Supplementary Information). For the analysis in Figs 2 and 3, two-tailed comparisons were used; for Fig. 4 and Supplementary Fig. 8, in which our hypothesis predicts a particular result, one-tailed comparisons were used.
Biosensor recordings
The ATP biosensors have been described elsewhere12,13. Those used here were modified with an internal screening layer to enhance the selectivity and greatly reduce their sensitivity to non-specific electroactive reagents. The sensing portion was 500 μm long and 50 μm in diameter. Embryos at late stage 11.5 were placed in a small chamber and wedged between fine tungsten pins so that the developing neural plate was uppermost. The biosensors were inserted into the embryo, to a depth of about 300 μm, through a small hole made in the correct location with a sharpened tungsten pin. Glycerol (5 mM), an essential co-substrate for the biosensor operation, was included in 1/10 Barth X recording medium along with 1 mM MgCl2. The null biosensors, lacking the ATP-detecting enzymes, acted as a control. They were sensitive to any possible interference but did not respond to ATP. A continuous recording was made for at least 2 h (Stage 13). At the end of the experiment, the sensors were withdrawn from the embryo and calibrated with ATP to allow conversion of the sensor signal to units of concentration.
References
Chow, R. & Lang, R. Early eye development in vertebrates. Annu. Rev. Cell Dev. Biol. 17, 255–296 (2001)
Hill, R. E. et al. Mouse Small eye results from mutations in a paired-like homeobox-containing gene. Nature 354, 522–525 (1991)
Quiring, R., Walldorf, U., Kloter, U. & Gehring, W. J. Homology of the eyeless gene of Drosophila to the Small eye gene in mice and Aniridia in humans. Science 265, 785–789 (1996)
Zuber, M., Gestri, G., Viczian, A., Barsacchi, G. & Harris, W. A. Specification of the vertebrate eye by a network of eye field transcription factors. Development 130, 5155–5167 (2003)
Mathers, P., Grinberg, A., Mahon, K. & Jamrich, M. The Rx homeobox gene is essential for vertebrate eye development. Nature 387, 603–607 (1997)
Loosli, F., Winkler, S. & Wittbrodt, J. Six3 overexpression initiates the formation of ectopic retina. Genes Dev. 13, 649–654 (1999)
Esteve, P. & Bovolenta, P. Secreted inducers in vertebrate eye development: more functions for old morphogens. Curr. Opin. Neurobiol. 16, 13–19 (2006)
Massé, K., Eason, R., Bhamra, S., Dale, N. & Jones, E. A. Comparative genomic and expression analysis of the conserved NTPDase gene family in Xenopus . Genomics 87, 366–381 (2006)
Zimmermann, H. Extracellular metabolism of ATP and other nucleotides. Naunyn-Schmiedebergs Arch. Pharmacol. 362, 299–309 (2000)
Brown, P. & Dale, N. Modulation of K+ currents in Xenopus spinal neurons by p2y receptors: a role for ATP and ADP in motor pattern generation. J. Physiol. (Lond.) 540, 843–850 (2002)
Cheng, A. W. et al. cDNA encodes Xenopus P2Y(1) nucleotide receptor: expression at the neuromuscular junctions. Neuroreport 14, 351–357 (2003)
Llaudet, E., Hatz, S., Droniou, M. & Dale, N. Microelectrode biosensor for real-time measurement of ATP in biological tissue. Anal. Chem. 77, 3267–3273 (2005)
Pearson, R. A., Dale, N., Llaudet, E. & Mobbs, P. ATP released via gap junction hemichannels from the pigment epithelium regulates neural retinal progenitor proliferation. Neuron 46, 731–744 (2005)
Gourine, A. V., Llaudet, E., Dale, N. & Spyer, K. M. ATP is a mediator of chemosensory transduction in the central nervous system. Nature 436, 108–111 (2005)
Neas, K. R. et al. Three patients with terminal deletions within the subtelomeric region of chromosome 9q. Am. J. Med. Genet. A 132, 425–430 (2005)
Allderdice, P. W. et al. Duplication 9q34 syndrome. Am. J. Hum. Genet. 35, 1005–1019 (1983)
Hampshire, D. J. et al. MORM syndrome (mental retardation, truncal obesity, retinal dystrophy and micropenis), a new autosomal recessive disorder, links to 9q34. Eur. J. Hum. Genet. 14, 543–548 (2006)
Yatsenko, S. A. et al. Deletion 9q34.3 syndrome: genotype–phenotype correlations and an extended deletion in a patient with features of Opitz C trigonocephaly. J. Med. Genet. 42, 328–335 (2005)
Chow, R., Altmann, C., Lang, R. & Hemmati-Brivanlou, A. Pax6 induces ectopic eyes in vertebrates. Development 126, 4213–4222 (1999)
Li, H., Tierney, C., Wen, L., Wu, J. Y. & Rao, Y. A single morphogenetic field gives rise to two retina primordia under the influence of the prechordal plate. Development 124, 603–615 (1997)
Martinez-Morales, J. R., Signore, M., Acampora, D., Simeone, A. & Bovolenta, P. Otx genes are required for tissue specification in the developing eye. Development 128, 2019–2030 (2001)
Collignon, J. et al. A comparison of the properties of Sox-3 with Sry and two related genes, Sox-1 and Sox-2 . Development 122, 509–520 (1996)
Glaser, T. et al. PAX6 gene dosage effect in a family with congenital cataracts, aniridia, anophthalmia and central nervous system defects. Nature Genet. 7, 463–471 (2004)
Stoykova, A., Fritsch, R., Walther, C. & Gruss, P. Forebrain patterning defects in Small eye mutant mice. Development 122, 3453–3465 (1996)
Mitchell, T. N. et al. Polymicrogyria and absence of pineal gland due to PAX6 mutation. Ann. Neurol. 53, 658–663 (2003)
Braun, N. et al. Expression of the ecto-ATPase NTPDase2 in the germinal zones of the developing and adult rat brain. Eur. J. Neurosci. 17, 1355–1364 (2003)
Mishra, S. K. et al. Extracellular nucleotide signaling in adult neural stem cells: synergism with growth factor-mediated cellular proliferation. Development 133, 675–684 (2006)
Nieuwkoop, P. D & Faber, J. Normal Table of Xenopus laevis (Daudin). (Garland, New York, 1994.
Bourguignon, C., Li, J. & Papalopulu, N. XBF-1, a winged helix transcription factor with dual activity, has a role in positioning neurogenesis in Xenopus competent ectoderm. Development 125, 4889–4900 (1998)
Barnett, M. W., Old, R. W. & Jones, E. A. Neural induction and patterning by fibroblast growth factor, notochord and somite tissue in Xenopus . Dev. Growth Differ. 40, 47–57 (1998)
Harland, R. In situ hybridisation: an improved wholemount method for Xenopus embryos. Methods Cell Biol. 36, 685–695 (1991)
Hirsch, N. & Harris, W. A. Xenopus Pax-6 and retinal development. J. Neurobiol. 32, 45–61 (1997)
Casarosa, S., Andreazzoli, M., Simeone, A. & Barsacchi, G. Xrx1, a novel Xenopus homeobox gene expressed during eye and pineal gland development. Mech. Dev. 61, 187–198 (1997)
Pannese, M. et al. The Xenopus homologue of Otx2 is a maternal homeobox gene that demarcates and specifies anterior body regions. Development 12, 707–720 (1995)
Haldin, C. E., Nijjar, S., Massé, K., Barnett, M. W. & Jones, E. A. Isolation and growth factor inducibility of the Xenopus laevis Lmx1b gene. Int. J. Dev. Biol. 47, 253–262 (2003)
Acknowledgements
We thank P. Jarrett for the maintenance of the frogs and E. Llaudet for the production of biosensors. We thank M. Andreazzoli, G. Guidetti, W. Harris, M. Hodgkin, H. Isaacs, A. Philpott, D. Sakaguchi, C. Smith and P. Stanfield for constructs and antibodies used in this work. This work was supported by the Wellcome Trust.
Author Contributions K.M. performed all molecular biology. K.M. and E.A.J. performed the mis-expression studies and phenotype analysis. E.A.J. performed all microinjections and dissection. S.B. performed in situ hybridization and sectioning, R.E. performed enzymatic activity assays, N.D. performed the ATP biosensor studies and the statistical analysis with K.M. E.A.J. and N.D. supervised the research project. K.M., N.D. and E.A.J. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
N.D. is a director and shareholder in Sarissa Biomedical Ltd., a company that makes biosensors similar to those used in this study.
Supplementary information
Supplementary Information
The file contains Supplementary Tables 1-5 and Supplementary Figures 1-9 with Legends. (PDF 2602 kb)
Rights and permissions
About this article
Cite this article
Massé, K., Bhamra, S., Eason, R. et al. Purine-mediated signalling triggers eye development. Nature 449, 1058–1062 (2007). https://doi.org/10.1038/nature06189
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06189
This article is cited by
-
Generation and characterization of polyclonal and monoclonal antibodies to human NTPDase2 including a blocking antibody
Purinergic Signalling (2017)
-
Adenine Nucleotides Control Proliferation In Vivo of Rat Retinal Progenitors by P2Y1 Receptor
Molecular Neurobiology (2017)
-
Hyperglycemia alters E-NTPDases, ecto-5′-nucleotidase, and ectosolic and cytosolic adenosine deaminase activities and expression from encephala of adult zebrafish (Danio rerio)
Purinergic Signalling (2016)
-
Spatiotemporal quantification of subcellular ATP levels in a single HeLa cell during changes in morphology
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.