Abstract
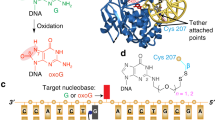
The enzyme uracil DNA glycosylase (UNG) excises unwanted uracil bases in the genome using an extrahelical base recognition mechanism. Efficient removal of uracil is essential for prevention of C-to-T transition mutations arising from cytosine deamination, cytotoxic U•A pairs arising from incorporation of dUTP in DNA, and for increasing immunoglobulin gene diversity during the acquired immune response. A central event in all of these UNG-mediated processes is the singling out of rare U•A or U•G base pairs in a background of approximately 109 T•A or C•G base pairs in the human genome. Here we establish for the human and Escherichia coli enzymes that discrimination of thymine and uracil is initiated by thermally induced opening of T•A and U•A base pairs and not by active participation of the enzyme. Thus, base-pair dynamics has a critical role in the genome-wide search for uracil, and may be involved in initial damage recognition by other DNA repair glycosylases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lindahl, T. & Wood, R. D. Quality control by DNA repair. Science 286, 1897–1905 (1999)
Stivers, J. T. & Jiang, Y. L. A mechanistic perspective on the chemistry of DNA repair glycosylases. Chem. Rev. 103, 2729–2759 (2003)
Seiple, L., Jaruga, P., Dizdaroglu, M. & Stivers, J. T. Linking uracil base excision repair and 5-fluorouracil toxicity in yeast. Nucleic Acids Res. 34, 140–151 (2006)
Kavli, B., Otterlei, M., Slupphaug, G. & Krokan, H. E. Uracil in DNA—General mutagen, but normal intermediate in acquired immunity. DNA Repair 6, 505–516 (2006)
Slupphaug, G. et al. A nucleotide-flipping mechanism from the structure of human uracil–DNA glycosylase bound to DNA. Nature 384, 87–92 (1996)
Parikh, S. S. et al. Uracil-DNA glycosylase–DNA substrate and product structures: conformational strain promotes catalytic efficiency by coupled stereoelectronic effects. Proc. Natl Acad. Sci. USA 97, 5083–5088 (2000)
Kavli, B. et al. Excision of cytosine and thymine from DNA by mutants of human uracil-DNA glycosylase. EMBO J. 15, 3442–3447 (1996)
Cao, C., Jiang, Y. L., Stivers, J. T. & Song, F. Dynamic opening of DNA during the enzymatic search for a damaged base. Nature Struct. Mol. Biol. 11, 1230–1236 (2004)
Cao, C., Jiang, Y. L., Krosky, D. J. & Stivers, J. T. The catalytic power of uracil DNA glycosylase in the opening of thymine base pairs. J. Am. Chem. Soc. 128, 13034–13035 (2006)
Krosky, D. J., Song, F. & Stivers, J. T. The origins of high-affinity enzyme binding to an extrahelical DNA base. Biochemistry 44, 5949–5959 (2005)
Krosky, D. J., Schwarz, F. P. & Stivers, J. T. Linear free energy correlations for enzymatic base flipping: How do damaged base pairs facilitate specific recognition? Biochemistry 43, 4188–4195 (2004)
Moran, S., Ren, R. X. F., Sheils, C. J., Rumney, S. & Kool, E. T. Non-hydrogen bonding 'terminator' nucleosides increase the 3′-end homogeneity of enzymatic RNA and DNA synthesis. Nucleic Acids Res. 24, 2044–2052 (1996)
Jiang, Y. L. & Stivers, J. T. Mutational analysis of the base flipping mechanism of uracil DNA glycosylase. Biochemistry 41, 11236–11247 (2002)
Banerjee, A., Yang, W., Karplus, M. & Verdine, G. L. Structure of a repair enzyme interrogating undamaged DNA elucidates recognition of damaged DNA. Nature 434, 612–618 (2005)
Chen, L., Haushalter, K. A., Lieber, C. M. & Verdine, G. L. Direct visualization of a DNA glycosylase searching for damage. Chem. Biol. 9, 345–350 (2002)
Gueron, M. & Leroy, J. Studies of base pair kinetics by NMR measurement of proton exchange. Methods Enzymol. 261, 383–413 (1995)
Snoussi, K. & Leroy, J. L. Alteration of A•T base-pair opening kinetics by the ammonium cation in DNA A-tracts. Biochemistry 41, 12467–12474 (2002)
Banerjee, A., Santos, W. L. & Verdine, G. L. Structure of a DNA glycosylase searching for lesions. Science 311, 1153–1157 (2006)
Gueron, M. et al. Applications of imino proton exchange to nucleic acid kinetics and structures. In Structure and Methods (ed. Sarma, R. H.) 113–137 (Adenine press, New York, 1990)
Moe, J. G. & Russu, I. M. Kinetics and energetics of base-pair opening in 5′-d(CGCGAATTCGCG)-3′ and a substituted dodecamer containing ĠT mismatches. Biochemistry 31, 8421–8428 (1992)
Verdine, G. L. & Norman, D. P. Covalent trapping of protein–DNA complexes. Annu. Rev. Biochem. 72, 337–366 (2003)
Ren, R. X.-F., Chaudhuri, N. C., Paris, P. L., Rumney, S. & Kool, E. T. Naphthalene, phenanthrene, and pyrene as DNA base analogues: Synthesis, structure, and fluorescence in DNA. J. Am. Chem. Soc. 118, 7671–7678 (1996)
Drohat, A. C., Jagadeesh, J., Ferguson, E. & Stivers, J. T. The role of electrophilic and base catalysis in the mechanism of Escherichia coli uracil DNA glycosylase. Biochemistry 38, 11866–11875 (1999)
Drohat, A. C. et al. Hetronuclear NMR and crystallographic studies of wild-type and H187Q Escherichia coli uracil DNA glycosylase: Electrophilic catalysis of uracil expulsion by a neutral histidine 187. Biochemistry 38, 11876–11886 (1999)
Xiao, G. et al. Crystal structure of Escherichia coli uracil DNA glycosylase and its complexes with uracil and glycerol: structure and glycosylase mechanism revisited. Proteins 35, 13–24 (1999)
Slupphaug, G. et al. Properties of a recombinant human uracil-DNA glycosylase from the UNG gene and evidence that UNG encodes the major uracil-DNA glycosylase. Biochemistry 34, 128–138 (1995)
Otwinoski, Z. & Minor, W. Processing of X-ray diffraction data in oscillation mode. Methods Enzymol. 276, 307–325 (1997)
McCoy, A. J., Grosse-Kunstleve, R. W., Storoni, L. C. & Read, R. J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D 61, 458–464 (2005)
Collaborative Computational Project, Number 4. The CCP4 suite: Programs for protein crystallography. Acta Crystallogr. D 50, 760–776 (1994)
Esnouf, R. M. Further additions to Molscript version 1.4, including reading and contouring of electron density maps. Acta Crystallogr. D 55, 938–940 (1999)
Acknowledgements
We thank A. Majumdar for assistance with NMR experiments. This work was supported by NIH grants (J.T.S. and L.M.A.) and a major research instrumentation grant from the NSF.
Author Contributions J.B.P. prepared all mutant enzymes and performed the NMR experiments; M.A.B. collected, processed and refined X-ray data and performed structural analyses; D.J.K. conceived the structural approach and purified and crystallized the complexes; J.I.F. performed NMR studies on the L272G mutant; L.M.A. performed structural analyses, discussed and commented on the manuscript; J.T.S. analysed data and wrote the paper.
Coordinates and structure factor files for the EI complex and the abasic DNA complex have been deposited in the Protein Data Bank with accession numbers 2OXM and 2OYT, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-4 with Legends and Supplementary Tables 1-3. The Supplementary Figures contain: (1) a scheme describing the general findings, (2) electron density maps of the flipped thymine and the abasic sugar excision product, (3) imino proton spectra of the free T/A 10 mer DNA duplex and its complexes with wild-type and mutant UNG enzymes, and (4) magnetization transfer time courses for the imino protons of residues G3, G4 and T2 of free and UNG-bound DNA. The Supplementary Tables detail (1) crystallographic data collection and refinement statistics, (2) imino proton exchange rates of free and bound DNA, and (3) chemical shifts of imino protons in each enzyme complex. (PDF 1951 kb)
Rights and permissions
About this article
Cite this article
Parker, J., Bianchet, M., Krosky, D. et al. Enzymatic capture of an extrahelical thymine in the search for uracil in DNA. Nature 449, 433–437 (2007). https://doi.org/10.1038/nature06131
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06131
This article is cited by
-
Uracil-DNA glycosylase efficiency is modulated by substrate rigidity
Scientific Reports (2023)
-
“Flexible hinge” dynamics in mismatched DNA revealed by fluorescence correlation spectroscopy
Journal of Biological Physics (2022)
-
DNA repair glycosylase hNEIL1 triages damaged bases via competing interaction modes
Nature Communications (2021)
-
Both DNA global deformation and repair enzyme contacts mediate flipping of thymine dimer damage
Scientific Reports (2017)
-
Sulfamic acid-catalyzed, environmentally benign synthesis of bis-tetronic acids at ambient temperature
Research on Chemical Intermediates (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.