Abstract
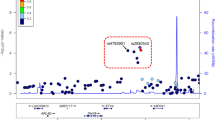
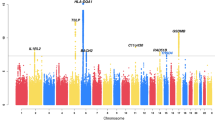
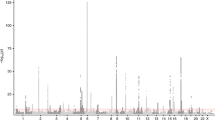
Asthma is caused by a combination of poorly understood genetic and environmental factors1,2. We have systematically mapped the effects of single nucleotide polymorphisms (SNPs) on the presence of childhood onset asthma by genome-wide association. We characterized more than 317,000 SNPs in DNA from 994 patients with childhood onset asthma and 1,243 non-asthmatics, using family and case-referent panels. Here we show multiple markers on chromosome 17q21 to be strongly and reproducibly associated with childhood onset asthma in family and case-referent panels with a combined P value of P < 10-12. In independent replication studies the 17q21 locus showed strong association with diagnosis of childhood asthma in 2,320 subjects from a cohort of German children (P = 0.0003) and in 3,301 subjects from the British 1958 Birth Cohort (P = 0.0005). We systematically evaluated the relationships between markers of the 17q21 locus and transcript levels of genes in Epstein–Barr virus (EBV)-transformed lymphoblastoid cell lines from children in the asthma family panel used in our association study. The SNPs associated with childhood asthma were consistently and strongly associated (P < 10-22) in cis with transcript levels of ORMDL3, a member of a gene family that encodes transmembrane proteins anchored in the endoplasmic reticulum3. The results indicate that genetic variants regulating ORMDL3 expression are determinants of susceptibility to childhood asthma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Cookson, W. The immunogenetics of asthma and eczema: a new focus on the epithelium. Nature Rev. Immunol. 4, 978–988 (2004)
Ober, C. & Hoffjan, S. Asthma genetics 2006: the long and winding road to gene discovery. Genes Immun. 7, 95–100 (2006)
Hjelmqvist, L. et al. ORMDL proteins are a conserved new family of endoplasmic reticulum membrane proteins. Genome Biol. 3, RESEARCH0027 (2002)
Weiland, S. K. et al. Phase II of the International Study of Asthma and Allergies in Childhood (ISAAC II): rationale and methods. Eur. Respir. J. 24, 406–412 (2004)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Statist. Soc. B. 57, 289–300 (1995)
Setakis, E., Stirnadel, H. & Balding, D. J. Logistic regression protects against population structure in genetic association studies. Genome Res. 16, 290–296 (2006)
Schadt, E. E. et al. Genetics of gene expression surveyed in maize, mouse and man. Nature 422, 297–302 (2003)
Morley, M. et al. Genetic analysis of genome-wide variation in human gene expression. Nature 430, 743–747 (2004)
Standards . for the diagnosis and care of patients with chronic obstructive pulmonary disease (COPD) and asthma. This official statement of the American Thoracic Society was adopted by the ATS Board of Directors, November 1986. Am. Rev. Respir. Dis. 136, 225–244 (1987)
British . guideline on the management of asthma. Thorax 58 (Suppl 1). i1–i94 (2003)
Abecasis, G., Cardon, L & Cookson, W. Selection strategies for disequilibrium mapping of quantitative traits in nuclear families. Am. J. Hum. Genet. 65, A245 (1999)
Gunderson, K. L., Steemers, F. J., Lee, G., Mendoza, L. G. & Chee, M. S. A genome-wide scalable SNP genotyping assay using microarray technology. Nature Genet. 37, 549–554 (2005)
Steemers, F. J. et al. Whole-genome genotyping with the single-base extension assay. Nature Methods 3, 31–33 (2006)
Buetow, K. H. et al. High-throughput development and characterization of a genomewide collection of gene-based single nucleotide polymorphism markers by chip-based matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Proc. Natl Acad. Sci. USA 98, 581–584 (2001)
Williams, R. L. A note on robust variance estimation for cluster-correlated data. Biometrics 56, 645–646 (2000)
Clayton, D. A generalization of the transmission/disequilibrium test for uncertain-haplotype transmission. Am. J. Hum. Genet. 65, 1170–1177 (1999)
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003)
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003)
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003)
Abecasis, G. R., Cherny, S. S., Cookson, W. O. & Cardon, L. R. Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nature Genet. 30, 97–101 (2002)
Burdick, J. T., Chen, W. M., Abecasis, G. R. & Cheung, V. G. In silico method for inferring genotypes in pedigrees. Nature Genet. 38, 1002–1004 (2006)
Abecasis, G. R. & Cookson, W. O. GOLD–graphical overview of linkage disequilibrium. Bioinformatics 16, 182–183 (2000)
Acknowledgements
The study was funded by the Wellcome Trust, the Medical Research Council, the French Ministry of Higher Education and Research, the German Ministry of education and research (BMBF), the national genome research network (NGFN), the National Institutes of Health (NHGRI and NHLBI; G.R.A.), and the European Commission as part of GABRIEL (a multidisciplinary study to identify the genetic and environmental causes of asthma in the European Community). We acknowledge use of genotype data from the British 1958 Birth Cohort DNA collection, funded by the Medical Research Council and the Wellcome Trust. We thank J. Todd for genotyping rs3894194 in the 1958 British Birth cohort.
Microarray and chromosome 17 genotyping data have been deposited in the GEO database, with accession number GSE8052.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-4 and Supplementary Figures 1-2 with Legends (PDF 141 kb)
Rights and permissions
About this article
Cite this article
Moffatt, M., Kabesch, M., Liang, L. et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature 448, 470–473 (2007). https://doi.org/10.1038/nature06014
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06014
This article is cited by
-
Shared genetic architecture between autoimmune disorders and B-cell acute lymphoblastic leukemia: insights from large-scale genome-wide cross-trait analysis
BMC Medicine (2024)
-
Prevalence, risk factors, impact and management of pneumonia among preschool children in Chinese seven cities: a cross-sectional study with interrupted time series analysis
BMC Medicine (2023)
-
Simultaneous reduction of all ORMDL proteins decreases the threshold of mast cell activation
Scientific Reports (2023)
-
Ceramide sensing by human SPT-ORMDL complex for establishing sphingolipid homeostasis
Nature Communications (2023)
-
Dietary long-chain omega 3 fatty acids modify sphingolipid metabolism to facilitate airway hyperreactivity
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.