Abstract
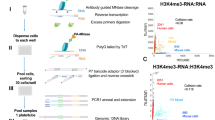
We report the application of single-molecule-based sequencing technology for high-throughput profiling of histone modifications in mammalian cells. By obtaining over four billion bases of sequence from chromatin immunoprecipitated DNA, we generated genome-wide chromatin-state maps of mouse embryonic stem cells, neural progenitor cells and embryonic fibroblasts. We find that lysine 4 and lysine 27 trimethylation effectively discriminates genes that are expressed, poised for expression, or stably repressed, and therefore reflect cell state and lineage potential. Lysine 36 trimethylation marks primary coding and non-coding transcripts, facilitating gene annotation. Trimethylation of lysine 9 and lysine 20 is detected at satellite, telomeric and active long-terminal repeats, and can spread into proximal unique sequences. Lysine 4 and lysine 9 trimethylation marks imprinting control regions. Finally, we show that chromatin state can be read in an allele-specific manner by using single nucleotide polymorphisms. This study provides a framework for the application of comprehensive chromatin profiling towards characterization of diverse mammalian cell populations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Surani, M. A., Hayashi, K. & Hajkova, P. Genetic and epigenetic regulators of pluripotency. Cell 128, 747–762 (2007)
Bernstein, B. E., Meissner, A. & Lander, E. S. The mammalian epigenome. Cell 128, 669–681 (2007)
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007)
Buck, M. J. & Lieb, J. D. ChIP-chip: considerations for the design, analysis, and application of genome-wide chromatin immunoprecipitation experiments. Genomics 83, 349–360 (2004)
Mockler, T. C. et al. Applications of DNA tiling arrays for whole-genome analysis. Genomics 85, 1–15 (2005)
Roh, T. Y., Cuddapah, S. & Zhao, K. Active chromatin domains are defined by acetylation islands revealed by genome-wide mapping. Genes Dev. 19, 542–552 (2005)
Service, R. F. Gene sequencing. The race for the $1000 genome. Science 311, 1544–1546 (2006)
Conti, L. et al. Niche-independent symmetrical self-renewal of a mammalian tissue stem cell. PLoS Biol. 3, e283 (2005)
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006)
Ringrose, L. & Paro, R. Epigenetic regulation of cellular memory by the Polycomb and Trithorax group proteins. Annu. Rev. Genet. 38, 413–443 (2004)
Schuettengruber, B., Chourrout, D., Vervoort, M., Leblanc, B. & Cavalli, G. Genome regulation by polycomb and trithorax proteins. Cell 128, 735–745 (2007)
Azuara, V. et al. Chromatin signatures of pluripotent cell lines. Nature Cell Biol. 8, 532–538 (2006)
Saxonov, S., Berg, P. & Brutlag, D. L. A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc. Natl Acad. Sci. USA 103, 1412–1417 (2006)
Weber, M. et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nature Genet. 39, 457–466 (2007)
Bernstein, B. E. et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120, 169–181 (2005)
Kim, T. H. et al. A high-resolution map of active promoters in the human genome. Nature 436, 876–880 (2005)
Boyer, L. A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, 349–353 (2006)
Lee, T. I. et al. Control of developmental regulators by polycomb in human embryonic stem cells. Cell 125, 301–313 (2006)
Squazzo, S. L. et al. Suz12 binds to silenced regions of the genome in a cell-type-specific manner. Genome Res. 16, 890–900 (2006)
Pasini, D., Bracken, A. P., Hansen, J. B., Capillo, M. & Helin, K. The Polycomb Group protein Suz12 is required for Embryonic Stem Cell differentiation. Mol. Cell. Biol. 27, 3769–3779 (2007)
Klose, R. J. & Bird, A. P. Genomic DNA methylation: the mark and its mediators. Trends Biochem. Sci. 31, 89–97 (2006)
Wang, X., Su, H. & Bradley, A. Molecular mechanisms governing Pcdh-γ gene expression: evidence for a multiple promoter and cis-alternative splicing model. Genes Dev. 16, 1890–1905 (2002)
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005)
Alexander, D. L., Ganem, L. G., Fernandez-Salguero, P., Gonzalez, F. & Jefcoate, C. R. Aryl-hydrocarbon receptor is an inhibitory regulator of lipid synthesis and of commitment to adipogenesis. J. Cell Sci. 111, 3311–3322 (1998)
Lengner, C. J. et al. Primary mouse embryonic fibroblasts: a model of mesenchymal cartilage formation. J. Cell. Physiol. 200, 327–333 (2004)
Garreta, E., Genove, E., Borros, S. & Semino, C. E. Osteogenic differentiation of mouse embryonic stem cells and mouse embryonic fibroblasts in a three-dimensional self-assembling peptide scaffold. Tissue Eng. 12, 2215–2227 (2006)
Doetsch, F. The glial identity of neural stem cells. Nature Neurosci. 6, 1127–1134 (2003)
Krichevsky, A. M., Sonntag, K. C., Isacson, O. & Kosik, K. S. Specific microRNAs modulate embryonic stem cell-derived neurogenesis. Stem Cells 24, 857–864 (2006)
Rao, B., Shibata, Y., Strahl, B. D. & Lieb, J. D. Dimethylation of histone H3 at lysine 36 demarcates regulatory and nonregulatory chromatin genome-wide. Mol. Cell. Biol. 25, 9447–9459 (2005)
Bannister, A. J. et al. Spatial distribution of di- and tri-methyl lysine 36 of histone H3 at active genes. J. Biol. Chem. 280, 17732–17736 (2005)
Kim, A., Kiefer, C. M. & Dean, A. Distinctive signatures of histone methylation in transcribed coding and noncoding human β-globin sequences. Mol. Cell. Biol. 27, 1271–1279 (2007)
Vakoc, C. R., Sachdeva, M. M., Wang, H. & Blobel, G. A. Profile of histone lysine methylation across transcribed mammalian chromatin. Mol. Cell. Biol. 26, 9185–9195 (2006)
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007)
Fantes, J. et al. Mutations in SOX2 cause anophthalmia. Nature Genet. 33, 461–463 (2003)
Hutchinson, J. N. et al. A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics 8, 39 (2007)
Seitz, H. et al. A large imprinted microRNA gene cluster at the mouse Dlk1-Gtl2 domain. Genome Res. 14, 1741–1748 (2004)
Cullen, B. R. Transcription and processing of human microRNA precursors. Mol. Cell 16, 861–865 (2004)
Zaratiegui, M., Irvine, D. V. & Martienssen, R. A. Noncoding RNAs and gene silencing. Cell 128, 763–776 (2007)
Verdel, A. & Moazed, D. RNAi-directed assembly of heterochromatin in fission yeast. FEBS Lett. 579, 5872–5878 (2005)
Martens, J. H. et al. The profile of repeat-associated histone lysine methylation states in the mouse epigenome. EMBO J. 24, 800–812 (2005)
Baust, C. et al. Structure and expression of mobile ETnII retroelements and their coding-competent MusD relatives in the mouse. J. Virol. 77, 11448–11458 (2003)
Svoboda, P. et al. RNAi and expression of retrotransposons MuERV-L and IAP in preimplantation mouse embryos. Dev. Biol. 269, 276–285 (2004)
Cho, D. H. et al. Antisense transcription and heterochromatin at the DM1 CTG repeats are constrained by CTCF. Mol. Cell 20, 483–489 (2005)
Feng, Y. Q. et al. The human β-globin locus control region can silence as well as activate gene expression. Mol. Cell. Biol. 25, 3864–3874 (2005)
Edwards, C. A. & Ferguson-Smith, A. C. Mechanisms regulating imprinted genes in clusters. Curr. Opin. Cell Biol. 19, 281–289 (2007)
Delaval, K. et al. Differential histone modifications mark mouse imprinting control regions during spermatogenesis. EMBO J. 26, 720–729 (2007)
Feil, R. & Berger, F. Convergent evolution of genomic imprinting in plants and mammals. Trends Genet. 23, 192–199 (2007)
Strausberg, R. L. et al. Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences. Proc. Natl Acad. Sci. USA 99, 16899–16903 (2002)
Acknowledgements
We thank S. Fisher, M. Kellis, B. Birren and M. Zody for technical assistance and constructive discussions. We acknowledge L. Zagachin in the MGH Nucleic Acid Quantitation core for assistance with real-time PCR. E.M. was supported by an institutional training grant from NIH. M.W. was supported by fellowships from the Human Frontiers Science Organization Program and the Ellison Foundation. This research was supported by funds from the National Human Genome Research Institute, the National Cancer Institute, the Burroughs Wellcome Fund, Massachusetts General Hospital, and the Broad Institute of MIT and Harvard.
All analysed data sets can be obtained from http://www.broad.mit.edu/seq_platform/chip/. Microarray data have been submitted to the GEO repository under accession number GSE8024.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Notes which includes the ChIP-Seq read requirement, genome coverage and accuracy and Supplementary Figures 1-10 with Legends (PDF 9260 kb)
Supplementary Table 1
This file contains Supplementary Table 1 which includes the list of datasets analyzed. (XLS 15 kb)
Supplementary Table 2
This file contains Supplementary Table 2 which includes the primers for RT-PCR validation of ChIP-Seq. (XLS 24 kb)
Supplementary Table 3
This file contains Supplementary Table 3 which includes the list of analyzed promoters and their chromatin state in ES cells, neural progenitors and embryonic fibroblasts. (XLS 3206 kb)
Supplementary Table 4
This file contains Supplementary Table 4 which includes the expression levels for analyzed genes in ES cells, neural progenitors and embryonic fibroblasts. (XLS 2070 kb)
Supplementary Table 5
This file contains Supplementary Table 5 which includes the Gene Ontology categories associated with monovalent and bivalent promoters in ES cells. (XLS 35 kb)
Supplementary Table 6
This file contains Supplementary Table 6 which includes the chromatin state of promoters associated with known regulators or markers of differentiated cell types (XLS 23 kb)
Supplementary Table 7
This file contains Supplementary Table 7 which includes the allelic bias observed at verified or putative imprinting control regions. (XLS 21 kb)
Rights and permissions
About this article
Cite this article
Mikkelsen, T., Ku, M., Jaffe, D. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007). https://doi.org/10.1038/nature06008
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06008
This article is cited by
-
Giant pandas in captivity undergo short-term adaptation in nerve-related pathways
BMC Zoology (2024)
-
Per-cell histone acetylation is associated with terminal differentiation in human T cells
Clinical Epigenetics (2024)
-
Deubiquitinase USP7 stabilizes KDM5B and promotes tumor progression and cisplatin resistance in nasopharyngeal carcinoma through the ZBTB16/TOP2A axis
Cell Death & Differentiation (2024)
-
Set7/9 aggravates ischemic brain injury via enhancing glutamine metabolism in a blocking Sirt5 manner
Cell Death & Differentiation (2024)
-
Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.