Abstract
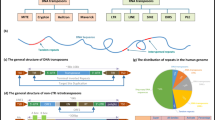
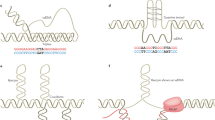
Nearly 30 hereditary disorders in humans result from an increase in the number of copies of simple repeats in genomic DNA. These DNA repeats seem to be predisposed to such expansion because they have unusual structural features, which disrupt the cellular replication, repair and recombination machineries. The presence of expanded DNA repeats alters gene expression in human cells, leading to disease. Surprisingly, many of these debilitating diseases are caused by repeat expansions in the non-coding regions of their resident genes. It is becoming clear that the peculiar structures of repeat-containing transcripts are at the heart of the pathogenesis of these diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Fleischer, B. Uber myotonische Dystrophie mit Katarakt. Albrecht Von Graefes Arch. Klin. Exp. Ophthalmol. 96, 91–133 (1918).
Sherman, S. L. et al. Further segregation analysis of the fragile X syndrome with special reference to transmitting males. Hum. Genet. 69, 289–299 (1985).
Verkerk, A. J. et al. Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 65, 905–914 (1991).
Kremer, E. J. et al. Mapping of DNA instability at the fragile X to a trinucleotide repeat sequence p(CCG)n . Science 252, 1711–1714 (1991).
La Spada, A. R., Wilson, E. M., Lubahn, D. B., Harding, A. E. & Fischbeck, K. H. Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature 352, 77–79 (1991).
Brook, J. D. et al. Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell 68, 799–808 (1992).
Mahadevan, M. et al. Myotonic dystrophy mutation: an unstable CTG repeat in the 3′ untranslated region of the gene. Science 255, 1253–1255 (1992).
Mirkin, S. M. Molecular models for repeat expansions. Chemtracts Biochem. Mol. Biol. 17, 639–662 (2004).
Pearson, C. E., Nichol Edamura, K. & Cleary, J. D. Repeat instability: mechanisms of dynamic mutations. Nature Rev. Genet. 6, 729–742 (2005).
Kunst, C. B. & Warren, S. T. Cryptic and polar variation of the fragile X repeat could result in predisposing normal alleles. Cell 77, 853–861 (1994).
Jodice, C. et al. Effect of trinucleotide repeat length and parental sex on phenotypic variation in spinocerebellar ataxia I. Am. J. Hum. Genet. 54, 959–965 (1994).
Brown, L. Y. & Brown, S. A. Alanine tracts: the expanding story of human illness and trinucleotide repeats. Trends Genet. 20, 51–58 (2004).
Wells, R. D., Dere, R., Hebert, M. L., Napierala, M. & Son, L.S. Advances in mechanisms of genetic instability related to hereditary neurological diseases. Nucleic Acids Res. 33, 3785–3798 (2005).
Mirkin, S. M. DNA structures, repeat expansions and human hereditary disorders. Curr. Opin. Struct. Biol. 16, 351–358 (2006).
Ranum, L. P. & Cooper, T. A. RNA-mediated neuromuscular disorders. Annu. Rev. Neurosci. 29, 259–277 (2006).
Kunkel, T. A. Slippery DNA and diseases. Nature 365, 207–208 (1993).
McMurray, C. T. DNA secondary structure: a common and causative factor for expansion in human disease. Proc. Natl Acad. Sci. USA 96, 1823–1825 (1999).
Gacy, A. M., Goellner, G., Juranic, N., Macura, S. & McMurray, C. T. Trinucleotide repeats that expand in human disease form hairpin structures in vitro. Cell 81, 533–540 (1995).
Dere, R., Napierala, M., Ranum, L. P. & Wells, R. D. Hairpin structure-forming propensity of the (CCTG·CAGG) tetranucleotide repeats contributes to the genetic instability associated with myotonic dystrophy type 2. J. Biol. Chem. 279, 41715–41726 (2004).
Usdin, K. & Woodford, K. J. CGG repeats associated with DNA instability and chromosome fragility from structures that block DNA synthesis in vitro. Nucleic Acids Res. 23, 4202–4209 (1995).
Fry, M. & Loeb, L. A. The fragile X syndrome d(CGG)n nucleotide repeats form a stable tetrahelical structure. Proc. Natl Acad. Sci. USA 91, 4950–4954 (1994).
Pearson, C. E. & Sinden, R. R. Alternative structures in duplex DNA formed within the trinucleotide repeats of the myotonic dystrophy and fragile X loci. Biochemistry 35, 5041–5053 (1996).
Pearson, C. E. et al. Slipped-strand DNAs formed by long (CAG)·(CTG) repeats: slipped-out repeats and slip-out junctions. Nucleic Acids Res. 30, 4534–4547 (2002).
Gacy, A. M. et al. GAA instability in Friedreich's ataxia shares a common, DNA-directed and intraallelic mechanism with other trinucleotide diseases. Mol. Cell 1, 583–593 (1998).
Sakamoto, N. et al. Sticky DNA: self association properties of long (GAA)·(TTC) repeats in R·R·Y triplex structures from Friedreich's ataxia. Mol. Cell 3, 465–475 (1999).
Vetcher, A. A. et al. Sticky DNA, a long (GAA·GAA·TTC) triplex that is formed intramolecularly, in the sequence of intron 1 of the frataxin gene. J. Biol. Chem. 277, 39217–39227 (2002).
Potaman, V. N. et al. Unpaired structures in SCA10 (ATTCT)n·(AGAAT)n repeats. J. Mol. Biol. 326, 1095–1111 (2003).
Moore, H., Greenwell, P. W., Liu, C. P., Arnheim, N. & Petes, T. D. Triplet repeats form secondary structures that escape DNA repair in yeast. Proc. Natl Acad. Sci. USA 96, 1504–1509 (1999).
Sakamoto, N. et al. GGA·TCC-interrupted triplets in long GAA·TTC repeats inhibit the formation of triplex and sticky DNA structures, alleviate transcription inhibition, and reduce genetic instabilities. J. Biol. Chem. 276, 27178–27187 (2001).
Kang, S., Jaworski, A., Ohshima, K. & Wells, R. D. Expansion and deletion of CTG repeats from human disease genes are determined by the direction of replication in E. coli. Nature Genet. 10, 213–218 (1995).
Ohshima, K. & Wells, R. D. Hairpin formation during DNA synthesis primer realignment in vitro in triplet repeat sequences from human hereditary disease genes. J. Biol. Chem. 272, 16798–16806 (1997).
Freudenreich, C. H., Stavenhagen, J. B. & Zakian, V. A. Stability of a CTG·CAG trinucleotide repeat in yeast is dependent on its orientation in the genome. Mol. Cell. Biol. 17, 2090–2098 (1997).
Miret, J. J., Pessoa-Brandao, L. & Lahue, R. S. Orientation-dependent and sequence-specific expansions of CTG·CAG trinucleotide repeats in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 95, 12438–12443 (1998).
Cleary, J. D., Nichol, K., Wang, Y. H. & Pearson, C. E. Evidence of cis-acting factors in replication-mediated trinucleotide repeat instability in primate cells. Nature Genet. 31, 37–46 (2002).
Rindler, M. P., Clark, R. M., Pollard, L. M., De Biase, I. & Bidichandani, S. I. Replication in mammalian cells recapitulates the locus-specific differences in somatic instability of genomic GAA triplet-repeats. Nucleic Acids Res. 34, 6352–6361 (2006).
Bhattacharyya, S. & Lahue, R. S. Saccharomyces cerevisiae Srs2 DNA helicase selectively blocks expansions of trinucleotide repeats. Mol. Cell. Biol. 24, 7324–7330 (2004).
Daee, D. L., Mertz, T., Collins, N. & Lahue, R. S. Post-replication repair inhibits CAG·CTG repeat expansions in Saccharomyces cerevisiae. Mol. Cell. Biol. 27, 102–110 (2007).
Samadashwily, G. M., Raca, G. & Mirkin, S. M. Trinucleotide repeats affect DNA replication in vivo. Nature Genet. 17, 298–304 (1997).
Krasilnikova, M. M. & Mirkin, S. M. Replication stalling at Friedreich's ataxia (GAA)n repeats in vivo. Mol. Cell. Biol. 24, 2286–2295 (2004).
Pelletier, R., Krasilnikova, M. M., Samadashwily, G. M., Lahue, R. S. & Mirkin, S. M. Replication and expansion of trinucleotide repeats in yeast. Mol. Cell. Biol. 23, 1349–1357 (2003).
Fouche, N., Ozgur, S., Roy, D. & Griffith, J. D. Replication fork regression in repetitive DNAs. Nucleic Acids Res. 34, 6044–6050 (2006).
Manley, K., Shirley, T. L., Flaherty, L. & Messer, A. Msh2 deficiency prevents in vivo somatic instability of the CAG repeat in Huntington disease transgenic mice. Nature Genet. 23, 471–473 (1999).
Savouret, C. et al. CTG repeat instability and size variation timing in DNA repair-deficient mice. EMBO J. 22, 2264–2273 (2003).
van den Broek, W. J. et al. Somatic expansion behaviour of the (CTG)n repeat in myotonic dystrophy knock-in mice is differentially affected by Msh3 and Msh6 mismatch-repair proteins. Hum. Mol. Genet. 11, 191–198 (2002).
Kovtun, I. V. & McMurray, C. T. Trinucleotide expansion in haploid germ cells by gap repair. Nature Genet. 27, 407–411 (2001).
Owen, B. A. et al. (CAG)n-hairpin DNA binds to Msh2–Msh3 and changes properties of mismatch recognition. Nature Struct. Mol. Biol. 12, 663–670 (2005).
Savouret, C. et al. MSH2-dependent germinal CTG repeat expansions are produced continuously in spermatogonia from DM1 transgenic mice. Mol. Cell. Biol. 24, 629–637 (2004).
Yoon, S. R., Dubeau, L., de Young, M., Wexler, N. S. & Arnheim, N. Huntington disease expansion mutations in humans can occur before meiosis is completed. Proc. Natl Acad. Sci. USA 100, 8834–8838 (2003).
Anvret, M. et al. Larger expansions of the CTG repeat in muscle compared to lymphocytes from patients with myotonic dystrophy. Hum. Mol. Genet. 2, 1397–1400 (1993).
Kennedy, L. et al. Dramatic tissue-specific mutation length increases are an early molecular event in Huntington disease pathogenesis. Hum. Mol. Genet. 12, 3359–3367 (2003).
Lia, A. S. et al. Somatic instability of the CTG repeat in mice transgenic for the myotonic dystrophy region is age dependent but not correlated to the relative intertissue transcription levels and proliferative capacities. Hum. Mol. Genet. 7, 1285–1291 (1998).
Fortune, M. T., Vassilopoulos, C., Coolbaugh, M. I., Siciliano, M. J. & Monckton, D. G. Dramatic, expansion-biased, age-dependent, tissue-specific somatic mosaicism in a transgenic mouse model of triplet repeat instability. Hum. Mol. Genet. 9, 439–445 (2000).
Kovtun, I. V. et al. OGG1 initiates age-dependent CAG expansion in somatic cells during base excision repair of oxidized bases in vitro and in vivo. Nature 447, 447–452 (2007).
Spiro, C. et al. Inhibition of FEN-1 processing by DNA secondary structure at trinucleotide repeats. Mol. Cell 4, 1079–1085 (1999).
Henricksen, L. A., Tom, S., Liu, Y. & Bambara, R. A. Inhibition of flap endonuclease 1 by flap secondary structure and relevance to repeat sequence expansion. J. Biol. Chem. 275, 16420–16427 (2000).
Panigrahi, G. B., Lau, R., Montgomery, S. E., Leonard, M. R. & Pearson, C. E. Slipped (CTG)·(CAG) repeats can be correctly repaired, escape repair or undergo error-prone repair. Nature Struct. Mol. Biol. 12, 654–662 (2005).
Liu, Y., Zhang, H., Veeraraghavan, J., Bambara, R. A. & Freudenreich, C. H. Saccharomyces cerevisiae flap endonuclease 1 uses flap equilibration to maintain triplet repeat stability. Mol. Cell. Biol. 24, 4049–4064 (2004).
van den Broek, W. J., Nelen, M. R., van der Heijden, G. W., Wansink, D. G. & Wieringa, B. Fen1 does not control somatic hypermutability of the (CTG)n·(CAG)n repeat in a knock-in mouse model for DM1. FEBS Lett. 580, 5208–5214 (2006).
Warren, S. T. Polyalanine expansion in synpolydactyly might result from unequal crossing-over of HOXD13. Science 275, 408–409 (1997).
Richards, R. I. et al. Evidence of founder chromosomes in fragile X syndrome. Nature Genet. 1, 257–260 (1992).
Jakupciak, J. P. & Wells, R. D. Genetic instabilities in (CTG)·(CAG) repeats occur by recombination. J. Biol. Chem. 274, 23468–23479 (1999).
Napierala, M., Dere, R., Vetcher, A. & Wells, R. D. Structure-dependent recombination hot spot activity of GAA·TTC sequences from intron 1 of the Friedreich's ataxia gene. J. Biol. Chem. 279, 6444–6454 (2004).
Dere, R. & Wells, R. D. DM2 CCTG·CAGG repeats are crossover hotspots that are more prone to expansions than the DM1 CTG·CAG repeats in Escherichia coli. J. Mol. Biol. 360, 21–36 (2006).
Freudenreich, C. H., Kantrow, S. M. & Zakian, V. A. Expansion and length-dependent fragility of CTG repeats in yeast. Science 279, 853–856 (1998).
Nag, D. K., Suri, M. & Stenson, E. K. Both CAG repeats and inverted DNA repeats stimulate spontaneous unequal sister-chromatid exchange in Saccharomyces cerevisiae. Nucleic Acids Res. 32, 5677–5684 (2004).
Richard, G.-F., Goellner, G. M., McMurray, C. T. & Haber, J. E. Recombination-induced CAG trinucleotide repeat expansions in yeast involve the MRE11–RAD50–XRS2 complex. EMBO J. 19, 2381–2390 (2000).
Richard, G.-F., Cyncynatus, C. & Dujon, B. Contractions and expansions of CAG·CTG trinucleotide repeats occur during ectopic gene conversion in yeast, by a MUS81-independent mechanism. J. Mol. Biol. 326, 769–782 (2003).
Meservy, J. L. et al. Long CTG tracts from the myotonic dystrophy gene induce deletions and rearrangements during recombination at the APRT locus in CHO cells. Mol. Cell. Biol. 23, 3152–3162 (2003).
Bacolla, A., Wojciechowska, M., Kosmider, B., Larson, J. E. & Wells, R. D. The involvement of non-B DNA structures in gross chromosomal rearrangements. DNA Repair (Amst.) 5, 1161–1170 (2006).
Rolfsmeier, M. L., Dixon, M. J. & Lahue, R. S. Mismatch repair blocks expansions of interrupted trinucleotide repeats in yeast. Mol. Cell 6, 1501–1507 (2000).
Mirkin, S. M. & Smirnova, E. V. Positioned to expand. Nature Genet. 31, 5–6 (2002).
Cleary, J. D. & Pearson, C. E. Replication fork dynamics and dynamic mutations: the fork-shift model of repeat instability. Trends Genet. 21, 272–280 (2005).
Abu-Baker, A. & Rouleau, G. A. in Genetic Instabilities and Neurological Diseases (eds Wells, R. D. & Ashizawa, T.) 487–513 (Elsevier, Amsterdam, 2006).
Davis, B. M., McCurrach, M. E., Taneja, K. L., Singer, R. H. & Housman, D. E. Expansion of a CUG trinucleotide repeat in the 3′ untranslated region of myotonic dystrophy protein kinase transcripts results in nuclear retention of transcripts. Proc. Natl Acad. Sci. USA 94, 7388–7393 (1997).
Amack, J. D. & Mahadevan, M. S. The myotonic dystrophy expanded CUG repeat tract is necessary but not sufficient to disrupt C2C12 myoblast differentiation. Hum. Mol. Genet. 10, 1879–1887 (2001).
Mankodi, A. et al. Myotonic dystrophy in transgenic mice expressing an expanded CUG repeat. Science 289, 1769–1772 (2000).
Tassone, F. et al. Elevated levels of FMR1 mRNA in carrier males: a new mechanism of involvement in the fragile-X syndrome. Am. J. Hum. Genet. 66, 6–15 (2000).
Jin, P. et al. RNA-mediated neurodegeneration caused by the fragile X premutation rCGG repeats in Drosophila. Neuron 39, 739–747 (2003).
Mutsuddi, M., Marshall, C. M., Benzow, K. A., Koob, M. D. & Rebay, I. The spinocerebellar ataxia 8 noncoding RNA causes neurodegeneration and associates with staufen in Drosophila. Curr. Biol. 14, 302–308 (2004).
Lin, X. & Ashizawa, T. Recent progress in spinocerebellar ataxia type-10 (SCA10). Cerebellum 4, 37–42 (2005).
Napierala, M. & Krzyzosiak, W. J. CUG repeats present in myotonin kinase RNA form metastable “slippery” hairpins. J. Biol. Chem. 272, 31079–31085 (1997).
Sobczak, K., de Mezer, M., Michlewski, G., Krol, J. & Krzyzosiak, W. J. RNA structure of trinucleotide repeats associated with human neurological diseases. Nucleic Acids Res. 31, 5469–5482 (2003).
Handa, V., Yeh, H. J., McPhie, P. & Usdin, K. The AUUCU repeats responsible for spinocerebellar ataxia type 10 form unusual RNA hairpins. J. Biol. Chem. 280, 29340–29345 (2005).
Napierala, M., Michalowski, D., de Mezer, M. & Krzyzosiak, W. J. Facile FMR1 mRNA structure regulation by interruptions in CGG repeats. Nucleic Acids Res. 33, 451–463 (2005).
Fardaei, M. et al. Three proteins, MBNL, MBLL and MBXL, co-localize in vivo with nuclear foci of expanded-repeat transcripts in DM1 and DM2 cells. Hum. Mol. Genet. 11, 805–814 (2002).
Miller, J. W. et al. Recruitment of human muscleblind proteins to (CUG)n expansions associated with myotonic dystrophy. EMBO J. 19, 4439–4448 (2000).
Lu, X., Timchenko, N. A. & Timchenko, L. T. Cardiac elav-type RNA-binding protein (ETR-3) binds to RNA CUG repeats expanded in myotonic dystrophy. Hum. Mol. Genet. 8, 53–60 (1999).
Philips, A. V., Timchenko, L. T. & Cooper, T. A. Disruption of splicing regulated by a CUG-binding protein in myotonic dystrophy. Science 280, 737–741 (1998).
Thornton, C. A., Swanson, M. S. & Cooper, T. A. in Genetic Instabilities and Neurological Diseases (eds Wells, R. D. & Ashizawa, T.) 37–54 (Elsevier, Amsterdam, 2006).
Ho, T. H. et al. Muscleblind proteins regulate alternative splicing. EMBO J. 23, 3103–3112 (2004).
Kino, Y. et al. Muscleblind protein, MBNL1/EXP, binds specifically to CHHG repeats. Hum. Mol. Genet. 13, 495–507 (2004).
Kanadia, R. N. et al. A muscleblind knockout model for myotonic dystrophy. Science 302, 1978–1980 (2003).
Kanadia, R. N. et al. Reversal of RNA missplicing and myotonia after muscleblind overexpression in a mouse poly(CUG) model for myotonic dystrophy. Proc. Natl Acad. Sci. USA 103, 11748–11753 (2006).
Iwahashi, C. K. et al. Protein composition of the intranuclear inclusions of FXTAS. Brain 129, 256–271 (2006).
Malinina, L. Possible involvement of the RNAi pathway in trinucleotide repeat expansion diseases. J. Biomol. Struct. Dyn. 23, 233–235 (2005).
Handa, V., Saha, T. & Usdin, K. The fragile X syndrome repeats form RNA hairpins that do not activate the interferon-inducible protein kinase, PKR, but are cut by Dicer. Nucleic Acids Res. 31, 6243–6248 (2003).
Krol, J. et al. Ribonuclease Dicer cleaves triplet repeat hairpins into shorter repeats which silence specific targets. Mol. Cell 25, 575–586 (2007).
Cho, D. H. et al. Antisense transcription and heterochromatin at the DM1 CTG repeats are constrained by CTCF. Mol. Cell 20, 483–489 (2005).
Moseley, M. L. et al. Bidirectional expression of CUG and CAG expansion transcripts and intranuclear polyglutamine inclusions in spinocerebellar ataxia type 8. Nature Genet. 38, 758–769 (2006).
Gray, S. J., Gerhardt, J., Doerfler, W., Small, L. E. & Fanning, E. An origin of DNA replication in the promoter region of the human fragile X mental retardation (FMR1) gene. Mol. Cell. Biol. 27, 426–437 (2007).
Acknowledgements
I thank W. Krzyzosiak, R. Lahue, K. Lobachev, C. McMurray, D. Monckton, C. Pearson, M. Swanson, K. Usdin and R. Wells for sharing their ideas and unpublished results. I also extend my gratitude to all the participants of the 5th International Conference on Unstable Microsatellites and Human Disease (Granada, Spain, 2006) for their intense and productive discussions, which helped to shape this review. I am indebted to my wife, Kate, for her invaluable critical comments. I thank J. White and P. White for their generous support. This work was supported by the National Institutes of Health.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Rights and permissions
About this article
Cite this article
Mirkin, S. Expandable DNA repeats and human disease. Nature 447, 932–940 (2007). https://doi.org/10.1038/nature05977
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05977
This article is cited by
-
Delineating the mechanism of fragility at BCL6 breakpoint region associated with translocations in diffuse large B cell lymphoma
Cellular and Molecular Life Sciences (2024)
-
LUSTR: a new customizable tool for calling genome-wide germline and somatic short tandem repeat variants
BMC Genomics (2024)
-
Comparison of microsatellite distribution in the genomes of Pteropus vampyrus and Miniopterus natalensis (Chiroptera)
BMC Genomic Data (2023)
-
Dynamic alternative DNA structures in biology and disease
Nature Reviews Genetics (2023)
-
Rediscovering tandem repeat variation in schizophrenia: challenges and opportunities
Translational Psychiatry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.