Abstract
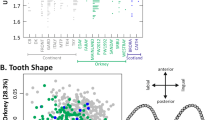
The origin and patterns of dispersal of anatomically modern humans are the focus of considerable debate1,2,3. Global genetic analyses have argued for one single origin, placed somewhere in Africa4,5,6,7. This scenario implies a rapid expansion, with a series of bottlenecks of small amplitude, which would have led to the observed smooth loss of genetic diversity with increasing distance from Africa. Analyses of cranial data, on the other hand, have given mixed results8,9,10,11,12, and have been argued to support multiple origins of modern humans2,9,12. Using a large data set of skull measurements and an analytical framework equivalent to that used for genetic data, we show that the loss in genetic diversity has been mirrored by a loss in phenotypic variability. We find evidence for an African origin, placed somewhere in the central/southern part of the continent, which harbours the highest intra-population diversity in phenotypic measurements. We failed to find evidence for a second origin, and we confirm these results on a large genetic data set. Distance from Africa accounts for an average 19–25% of heritable variation in craniometric measurements—a remarkably strong effect for phenotypic measurements known to be under selection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mellars, P. Going east: New genetic and archaeological perspectives on the modern human colonization of Eurasia. Science 313, 796–800 (2006)
Trinkaus, E. Early modern humans. Annu. Rev. Anthropol. 34, 207–230 (2005)
Mellars, P. Why did modern human populations disperse from Africa ca. 60,000 years ago? A new model. Proc. Natl Acad. Sci. USA 103, 9381–9386 (2006)
Liu, H., Prugnolle, F., Manica, A. & Balloux, F. A geographically explicit genetic model of worldwide human-settlement history. Am. J. Hum. Genet. 79, 230–237 (2006)
Prugnolle, F., Manica, A. & Balloux, F. Geography predicts neutral genetic diversity of human populations. Curr. Biol. 15, R159–R160 (2005)
Ramachandran, S. et al. Support from the relationship of genetic and geographic distance in human populations for a serial founder effect originating in Africa. Proc. Natl Acad. Sci. USA 102, 15942–15947 (2005)
Ray, N., Currat, M., Berthier, P. & Excoffier, L. Recovering the geographic origin of early modern humans by realistic and spatially explicit simulations. Genome Res. 15, 1161–1167 (2005)
Grine, F. E. et al. Late Pleistocene human skull from Hofmeyr, South Africa, and modern human origins. Science 315, 226–229 (2007)
Wolpoff, M. H., Hawks, J. & Caspari, R. Multiregional, not multiple origins. Am. J. Phys. Anthropol. 112, 129–136 (2000)
Brauer, G., Collard, M. & Stringer, C. On the reliability of recent tests of the Out of Africa hypothesis for modern human origins. Anat. Rec. A 279, 701–707 (2004)
Lahr, M. M. The multiregional model of modern human origins — A reassessment of its morphological basis. J. Hum. Evol. 26, 23–56 (1994)
Wolpoff, M. H. in The Human Revolution: Biological Perspectives in the Origins of Modern Humans (eds Mellars, P. & Stringer, C.) 62–108 (Princeton Univ. Press, Princeton, 1989)
Fowler, K. & Whitlock, M. C. The distribution of phenotypic variance with inbreeding. Evolution Int. J. Org. Evolution 53, 1143–1156 (1999)
Frankham, R. Do island populations have less genetic variation than mainland populations? Heredity 78, 311–327 (1997)
Hanihara, T. & Ishida, H. Metric dental variation of major human populations. Am. J. Phys. Anthropol. 128, 287–298 (2005)
Hanihara, T. & Ishida, H. Os incae: variation in frequency in major human population groups. J. Anat. 198, 137–152 (2001)
Roseman, C. C. Detecting interregionally diversifying natural selection on modern human cranial form by using matched molecular and morphometric data. Proc. Natl Acad. Sci. USA 101, 12824–12829 (2004)
Relethford, J. H. Boas and beyond: Migration and craniometric variation. Am. J. Hum. Biol. 16, 379–386 (2004)
Relethford, J. H. & Blangero, J. Detection of differential gene flow from patterns of quantitative variation. Hum. Biol. 62, 5–25 (1990)
Manica, A., Prugnolle, F. & Balloux, F. Geography is a better determinant of human genetic differentiation than ethnicity. Hum. Genet. 118, 366–371 (2005)
Rosenberg, N. A. Standardized subsets of the HGDP-CEPH human genome diversity cell line panel, accounting for atypical and duplicated samples and pairs of close relatives. Ann. Hum. Genet. 70, 841–847 (2006)
Rosenberg, N. A. et al. Genetic structure of human populations. Science 298, 2381–2385 (2002)
Venables, W. N. & Ripley, B. D. Modern Applied Statistics with S (Springer, New York, 2002)
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003)
Carson, E. A. Maximum likelihood estimation of human craniometric heritabilities. Am. J. Phys. Anthropol. 131, 169–180 (2006)
Prugnolle, F. et al. Pathogen-driven selection and worldwide HLA class I diversity. Curr. Biol. 15, 1022–1027 (2005)
Hijmans, R. J., Cameron, S. E., Parra, J. L., Jones, P. G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Climatol. 25, 1965–1978 (2005)
Crawley, M. J. Statistical Computing: An Introduction to Data Analysis Using S-Plus (Wiley, London, 2002)
Burnham, K. P. & Anderson, D. R. Model Selection and Inferences (Springer, New York, 1998)
R Development Core Team (R Foundation for Statistical Computing, Vienna, 2006)
Acknowledgements
We acknowledge J. Goudet for discussions. The Biotechnology and Biological Sciences Research Council provided financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables S1-S4. (PDF 294 kb)
Rights and permissions
About this article
Cite this article
Manica, A., Amos, W., Balloux, F. et al. The effect of ancient population bottlenecks on human phenotypic variation. Nature 448, 346–348 (2007). https://doi.org/10.1038/nature05951
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature05951
This article is cited by
-
Female craniometrics support the ‘two-layer model’ of human dispersal in Eastern Eurasia
Scientific Reports (2021)
-
The origins of the division of labor in pre-industrial times
Journal of Economic Growth (2020)
-
Genomic comparisons reveal biogeographic and anthropogenic impacts in the koala (Phascolarctos cinereus): a dietary-specialist species distributed across heterogeneous environments
Heredity (2019)
-
The great human expansion
Resonance (2019)
-
Manot 1 calvaria and Recent Modern Human Evolution: an Anthropological Perspective
Bulletins et Mémoires de la Société d'Anthropologie de Paris (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.