Abstract
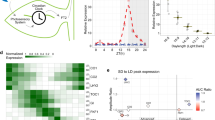
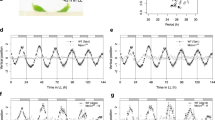
Most organisms use circadian oscillators to coordinate physiological and developmental processes such as growth with predictable daily environmental changes like sunrise and sunset. The importance of such coordination is highlighted by studies showing that circadian dysfunction causes reduced fitness in bacteria1 and plants2, as well as sleep and psychological disorders in humans3. Plant cell growth requires energy and water—factors that oscillate owing to diurnal environmental changes. Indeed, two important factors controlling stem growth are the internal circadian oscillator4,5,6 and external light levels7. However, most circadian studies have been performed in constant conditions, precluding mechanistic study of interactions between the clock and diurnal variation in the environment. Studies of stem elongation in diurnal conditions have revealed complex growth patterns, but no mechanism has been described8,9,10. Here we show that the growth phase of Arabidopsis seedlings in diurnal light conditions is shifted 8–12 h relative to plants in continuous light, and we describe a mechanism underlying this environmental response. We find that the clock regulates transcript levels of two basic helix–loop–helix genes, phytochrome-interacting factor 4 (PIF4) and PIF5, whereas light regulates their protein abundance. These genes function as positive growth regulators; the coincidence of high transcript levels (by the clock) and protein accumulation (in the dark) allows them to promote plant growth at the end of the night. Thus, these two genes integrate clock and light signalling, and their coordinated regulation explains the observed diurnal growth rhythms. This interaction may serve as a paradigm for understanding how endogenous and environmental signals cooperate to control other processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Woelfle, M. A., Ouyang, Y., Phanvijhitsiri, K. & Johnson, C. H. The adaptive value of circadian clocks: an experimental assessment in cyanobacteria. Curr. Biol. 14, 1481–1486 (2004)
Dodd, A. N. et al. Plant circadian clocks increase photosynthesis, growth, survival, and competitive advantage. Science 309, 630–633 (2005)
Dunlap, J. C., Loros, J. J. DeCoursey, P. J. (eds) Chronobiology (Sinauer Associates, Sunderland, Massachusetts, 2005)
Lecharny, A. & Wagner, E. Stem extension rate in light-grown plants. Evidence for an endogenous circadian rhythm in Chenopodium. Physiol. Plant. 60, 437–443 (1984)
Ibrahim, C. A., Lecharny, A. & Millet, B. Circadian endogenous growth rhythm in tomato. Plant Physiol. 67, 113 (1981)
Dowson-Day, M. J. & Millar, A. J. Circadian dysfunction causes aberrant hypocotyl elongation patterns in Arabidopsis. Plant J. 17, 63–71 (1999)
Chen, M., Chory, J. & Fankhauser, C. Light signal transduction in higher plants. Annu. Rev. Genet. 38, 87–117 (2004)
Bertram, L. & Karlsen, P. Patterns in stem elongation rate in chrysanthemum and tomato plants in relation to irradiance and day/night temperature. Sci. Hortic. 58, 139–150 (1994)
Tutty, J. R., Hicklenton, P. R., Kristie, D. N. & McRae, K. B. The influence of photoperiod and temperature on the kinetics of stem elongation in Dendranthema grandiflorum. J. Am. Soc. Hortic. Sci. 119, 138–143 (1994)
Bertram, L. & Lercari, B. Kinetics of stem elongation in light-grown tomato plants. Responses to different photosynthetically active radiation levels by wild-type and aurea mutant plants. Photochem. Photobiol. 66, 396–403 (1997)
Gardner, M. J., Hubbard, K. E., Hotta, C. T., Dodd, A. N. & Webb, A. A. How plants tell the time. Biochem. J. 397, 15–24 (2006)
Schaffer, R. et al. The late elongated hypocotyl mutation of Arabidopsis disrupts circadian rhythms and the photoperiodic control of flowering. Cell 93, 1219–1229 (1998)
Wang, Z. Y. & Tobin, E. M. Constitutive expression of the CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) gene disrupts circadian rhythms and suppresses its own expression. Cell 93, 1207–1217 (1998)
McWatters, H. G., Bastow, R. M., Hall, A. & Millar, A. J. The ELF3 zeitnehmer regulates light signalling to the circadian clock. Nature 408, 716–720 (2000)
Covington, M. F. et al. ELF3 modulates resetting of the circadian clock in Arabidopsis. Plant Cell 13, 1305–1315 (2001)
Thain, S. C. et al. Circadian rhythms of ethylene emission in Arabidopsis. Plant Physiol. 136, 3751–3761 (2004)
Duek, P. D. & Fankhauser, C. bHLH class transcription factors take centre stage in phytochrome signalling. Trends Plant Sci. 10, 51–54 (2005)
Osterlund, M. T., Hardtke, C. S., Wei, N. & Deng, X. W. Targeted destabilization of HY5 during light-regulated development of Arabidopsis. Nature 405, 462–466 (2000)
Kohchi, T. et al. The Arabidopsis HY2 gene encodes phytochromobilin synthase, a ferredoxin-dependent biliverdin reductase. Plant Cell 13, 425–436 (2001)
Huq, E. & Quail, P. H. PIF4, a phytochrome-interacting bHLH factor, functions as a negative regulator of phytochrome B signaling in Arabidopsis. EMBO J. 21, 2441–2450 (2002)
Yamashino, T. et al. A link between circadian-controlled bHLH factors and the APRR1/TOC1 quintet in Arabidopsis thaliana. Plant Cell Physiol. 44, 619–629 (2003)
Khanna, R. et al. A novel molecular recognition motif necessary for targeting photoactivated phytochrome signaling to specific basic helix–loop–helix transcription factors. Plant Cell 16, 3033–3044 (2004)
Fujimori, T., Yamashino, T., Kato, T. & Mizuno, T. Circadian-controlled basic/helix–loop–helix factor, PIL6, implicated in light-signal transduction in Arabidopsis thaliana. Plant Cell Physiol. 45, 1078–1086 (2004)
Park, E. et al. Degradation of phytochrome interacting factor 3 in phytochrome-mediated light signaling. Plant Cell Physiol. 45, 968–975 (2004)
Shen, H., Moon, J. & Huq, E. PIF1 is regulated by light-mediated degradation through the ubiquitin–26S proteasome pathway to optimize photomorphogenesis of seedlings in Arabidopsis. Plant J. 44, 1023–1035 (2005)
Al-Sady, B., Ni, W., Kircher, S., Schafer, E. & Quail, P. H. Photoactivated phytochrome induces rapid PIF3 phosphorylation prior to proteasome-mediated degradation. Mol. Cell 23, 439–446 (2006)
Bünning, E. Die endogene Tagesrhythmik als Grundlage der photoperiodischen reaktion. Ber. Dtsch. Bot. Ges. 54, 590–607 (1936)
Tsuda, M. & Tyree, M. T. Plant hydraulic conductance measured by the high pressure flow meter in crop plants. J. Exp. Bot. 51, 823–828 (2000)
Hsiao, T. C. Plant responses to water stress. Annu. Rev. Plant Physiol. 24, 519–570 (1973)
Strayer, C. et al. Cloning of the Arabidopsis clock gene TOC1, an autoregulatory response regulator homolog. Science 289, 768–771 (2000)
Doyle, M. R. et al. The ELF4 gene controls circadian rhythms and flowering time in Arabidopsis thaliana. Nature 419, 74–77 (2002)
R Development Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, Austria, 2005)
Schmid, M. et al. A gene expression map of Arabidopsis thaliana development. Nature Genet. 37, 501–506 (2005)
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003)
Breitling, R., Armengaud, P., Amtmann, A. & Herzyk, P. Rank products: a simple, yet powerful, new method to detect differentially regulated genes in replicated microarray experiments. FEBS Lett. 573, 83–92 (2004)
Gentleman, R. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004)
Czechowski, T., Bari, R. P., Stitt, M., Scheible, W.-R. & Udvardi, M. K. Real-time RT–PCR profiling of over 1400 Arabidopsis transcription factors: unprecedented sensitivity reveals novel root- and shoot-specific genes. Plant J. 38, 366–379 (2004)
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M. K. & Scheible, W.-R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 139, 5–17 (2005)
Mockler, T. C. et al. Regulation of flowering time in Arabidopsis by K homology domain proteins. Proc. Natl Acad. Sci. USA 101, 12759–12764 (2004)
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2–ΔΔCT method. Methods 25, 402–408 (2001)
Duek, P. D., Elmer, M. V., van Oosten, V. R. & Fankhauser, C. The degradation of HFR1, a putative bHLH class transcription factor involved in light signaling, is regulated by phosphorylation and requires COP1. Curr. Biol. 14, 2296–2301 (2004)
Acknowledgements
We thank A. Wallace for technical assistance; J. C. Lagarias, N. Sinha and C. Wessinger for critical reading and comments on the manuscript; E. Tobin, S. Kay, A. Millar, J. C. Lagarias, T. Mizuno, P. Quail and the Arabidopsis Biological Resources Centre for seeds; and J. C. Lagarias for the loan of computer equipment. This work was supported by grants from the NSF (to J.N.M. and C. Weinig), the Swiss National Science foundation (to C.F.), the HFSP (to C.F., J.N.M. and U. Genick), the NRI of the USDA CSREES (to M.F.C.) and the NIH (to S.L.H.). The microarray data have been deposited in the GEOdatabase (http://www.ncbi.nlm.nih.gov/projects/geo/) under accession number GSE6906.
Author Contributions K.N. performed all experiments. Statistical analysis of growth and microarray data was done by J.N.M. K.N. and J.N.M. wrote the paper. P.D.D., S.L. and C.F. contributed HA-tagged protein overexpressing plants, western blot protocols, and pif4 pif5 double-mutant seed. M.F.C. contributed microarray experimental design. K.N., J.N.M. and S.L.H. contributed to project design. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The microarray data have been deposited in the GEO database (http://www.ncbi.nlm.nih.gov/projects/geo/) under accession number GSE6906. Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Discussion of growth pattern shifts between LL and SD, of additional circadian mutants, and of the microarray data; Supplementary Figures 1-8 with Legends; Supplementary Tables 1 -2 and additional references. (PDF 1290 kb)
Rights and permissions
About this article
Cite this article
Nozue, K., Covington, M., Duek, P. et al. Rhythmic growth explained by coincidence between internal and external cues. Nature 448, 358–361 (2007). https://doi.org/10.1038/nature05946
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05946
This article is cited by
-
The expression of the NPR1-dependent defense response pathway genes in Persea americana (Mill.) following infection with Phytophthora cinnamomi
BMC Plant Biology (2023)
-
Transcription factors RhPIF4/8 and RhHY5 regulate autophagy-mediated petal senescence in rose (Rosa hybrida)
Horticulture Advances (2023)
-
SUPPRESSOR OF PHYTOCHROME B-4 #3 reduces the expression of PIF-activated genes and increases expression of growth repressors to regulate hypocotyl elongation in short days
BMC Plant Biology (2022)
-
Environment-mediated mutagenetic interference on genetic stabilization and circadian rhythm in plants
Cellular and Molecular Life Sciences (2022)
-
Stem respiration and growth in a central Amazon rainforest
Trees (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.