Abstract
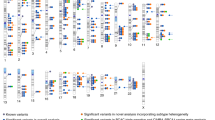
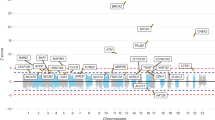
Breast cancer exhibits familial aggregation, consistent with variation in genetic susceptibility to the disease. Known susceptibility genes account for less than 25% of the familial risk of breast cancer, and the residual genetic variance is likely to be due to variants conferring more moderate risks. To identify further susceptibility alleles, we conducted a two-stage genome-wide association study in 4,398 breast cancer cases and 4,316 controls, followed by a third stage in which 30 single nucleotide polymorphisms (SNPs) were tested for confirmation in 21,860 cases and 22,578 controls from 22 studies. We used 227,876 SNPs that were estimated to correlate with 77% of known common SNPs in Europeans at r2 > 0.5. SNPs in five novel independent loci exhibited strong and consistent evidence of association with breast cancer (P < 10-7). Four of these contain plausible causative genes (FGFR2, TNRC9, MAP3K1 and LSP1). At the second stage, 1,792 SNPs were significant at the P < 0.05 level compared with an estimated 1,343 that would be expected by chance, indicating that many additional common susceptibility alleles may be identifiable by this approach.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
28 June 2007
In the AOP version of this Article, Kristy Driver was omitted from The SEARCH collaborators. This and the Supplementary Information acknowledgements have now been corrected for both print and online publication on 28 June 2007.
References
Collaborative Group on Hormonal Factors in Breast Cancer Familial breast cancer: Collaborative reanalysis of individual data from 52 epidemiological studies including 58 209 women with breast cancer and 101 986 women without the disease. Lancet 358, 1389–1399 (2001)
Miki, Y. et al. A strong candidate for the breast and ovarian-cancer susceptibility gene BRCA1. Science 266, 66–71 (1994)
Wooster, R. et al. Identification of the breast cancer susceptibility gene BRCA2. Nature 378, 789–792 (1995)
Antoniou, A. et al. Average risks of breast and ovarian cancer associated with mutations in BRCA1 or BRCA2 detected in case series unselected for family history: A combined analysis of 22 studies. Am. J. Hum. Genet. 72, 1117–1130 (2003)
Smith, P. et al. A genome wide linkage search for breast cancer susceptibility genes. Genes Chromosom. Cancer 45, 646–655 (2006)
Pharoah, P. D. P. et al. Polygenic susceptibility to breast cancer and implications for prevention. Nature Genet. 31, 33–36 (2002)
Antoniou, A. C., Pharoah, P. D. P., Smith, P. & Easton, D. F. The BOADICEA model of genetic susceptibility to breast and ovarian cancer. Br. J. Cancer 91, 1580–1590 (2004)
Rahman, N. et al. PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nature Genet. 39, 165–167 (2007)
Thompson, D. et al. Cancer risks and mortality in heterozygous ATM mutation carriers. J. Natl Cancer Inst. 97, 813–822 (2005)
Meijers-Heijboer, H. et al. Low-penetrance susceptibility to breast cancer due to CHEK2*1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nature Genet. 31, 55–59 (2002)
Erkko, H. et al. A recurrent mutation in PALB2 in Finnish cancer families. Nature 446, 316–319 (2007)
Renwick, A. et al. ATM mutations that cause ataxia-telangiectasia are breast cancer susceptibility alleles. Nature Genet. 38, 873–875 (2006)
Seal, S. et al. Truncating mutations in the Fanconi anemia J gene BRIP1 are low-penetrance breast cancer susceptibility alleles. Nature Genet. 38, 1239–1241 (2006)
The CHEK2 Breast Cancer Case-Control Consortium CHEK2*1100delC and susceptibility to breast cancer: A collaborative analysis involving 10,860 breast cancer cases and 9,065 controls from ten studies. Am. J. Hum. Genet. 74, 1175–1182 (2004)
Cox, A. et al. A common coding variant in CASP8 is associated with breast cancer risk. Nature Genetics 39, 352–358 (2007); corrigendum. 39, 688 (2007)
Easton, D. F. How many more breast cancer predisposition genes are there? Breast Cancer Res. 1, 1–4 (1999)
Kruglyak, L. & Nickerson, D. A. Variation is the spice of life. Nature Genet. 27, 234–236 (2001)
Hinds, D. A. et al. Whole-genome patterns of common DNA variation in three human populations. Science 307, 1072–1079 (2005)
Satagopan, J. M., Verbel, D. A., Venkatraman, E. S., Offit, K. E. & Begg, C. B. Two-stage designs for gene-disease association studies. Biometrics 58, 163–170 (2002)
Antoniou, A. C. & Easton, D. F. Polygenic inheritance of breast cancer: Implications for design of association studies. Genet. Epidemiol. 25, 190–202 (2003)
Altshuler, D. et al. A haplotype map of the human genome. Nature 437, 1299–1320 (2005)
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999)
Thomas, D. C., Haile, R. W. & Duggan, D. Recent developments in genomewide association scans: A workshop summary and review. Am. J. Hum. Genet. 77, 337–345 (2005)
Amundadottir, L. T. et al. A common variant associated with prostate cancer in European and African populations. Nature Genet. 38, 652–658 (2006)
Yeager, M. et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nature Genet. 39, 645–649 (2007)
Gudmundsson, J. et al. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nature Genet. 39, 631–637 (2007)
Ferretti, V. et al. PReMod: a database of genome-wide mammalian cis-regulatory module predictions. Nucleic Acids Res. 35, D122–D126 (2007)
Moffa, A. B., Tannheimer, S. L. & Ethier, S. P. Transforming potential of alternatively spliced variants of fibroblast growth factor receptor 2 in human mammary epithelial cells. Mol. Cancer Res. 2, 643–652 (2004)
Adnane, J. et al. Bek and Flg, 2 receptors to members of the Fgf family, are amplified in subsets of human breast cancers. Oncogene 6, 659–663 (1991)
Jang, J. H., Shin, K. H. & Park, J. G. Mutations in fibroblast growth factor receptor 2 and fibroblast growth factor receptor 3 genes associated with human gastric and colorectal cancers. Cancer Res. 61, 3541–3543 (2001)
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007)
Lesueur, F. et al. Allelic association of the human homologue of the mouse modifier Ptprj with breast cancer. Hum. Mol. Genet. 14, 2349–2356 (2005)
Day, N. et al. EPIC-Norfolk: Study design and characteristics of the cohort. Br. J. Cancer 80, 95–103 (1999)
Breast Cancer Association Consortium. Commonly studied SNPs and breast cancer: Negative results from 12,000 – 32,000 cases and controls from the Breast Cancer Association Consortium. J. Natl Cancer Inst. 98, 1382–1396 (2006)
Hubbard, T. et al. The Ensembl genome database project. Nucleic Acids Res. 30, 38–41 (2002)
Dean, F. B. et al. Comprehensive human genome amplification using multiple displacement amplification. Proc. Natl Acad. Sci. USA 99, 5261–5266 (2002)
de Bakker, P. I. W. et al. Efficiency and power in genetic association studies. Nature Genet. 37, 1217–1223 (2005)
Tyrer, J., Pharoah, P. D. P. & Easton, D. F. The admixture maximum likelihood test: A novel experiment-wise test of association between disease and multiple SNPs. Genet. Epidemiol. 30, 636–643 (2006)
Schaid, D. J., Rowland, C. M., Tines, D. E., Jacobson, R. M. & Poland, G. A. Score tests for association between traits and haplotypes when linkage phase is ambiguous. Am. J. Hum. Genet. 70, 425–434 (2002)
Acknowledgements
The authors thank the women who took part in this research, and all the funders and support staff who made this study possible. The principal funding for this study was provided by Cancer Research UK. Detailed acknowledgements are provided in Supplementary Information.
Author Contributions D.F.E., A.M.D., P.D.P.P., D.R.C. and B.A.J.P. designed the study and obtained financial support. D.G.B. and D.R.C. directed the genotyping of stages 1 and 2. D.F.E. and D.T. conducted the statistical analysis. K.A.P. and A.M.D. coordinated the genotyping for stage 3 and the fine-scale mapping of the FGFR2 and TNRC9 loci. J.P.S. and J.Z. performed resequencing and analysis of the FGFR2 locus. K.A.P., S.A., C.S.H., R.B., C.A.H., L.K.K., B.E.H., L.L.M., P.B., S.S., V.G., F.O., C-Y. S., P-E.W. and H-C.W. conducted genotyping for the fine-scale mapping. R.L., J.M., H.F. and K.B.M. provided bioinformatics support. D.E., D.G.E., J.P., O.F., N.J., S.S., M.R.S. and N.R. coordinated the studies used in stage 1. N.W. and N.E.D. coordinated the EPIC study used in stages 1 and 2. The remaining authors coordinated the studies in stage 3 and undertook genotyping in those studies. D.F.E. drafted the manuscript, with substantial contributions from K.A.P., A.M.D., P.D.P.P. and B.A.J.P. All authors contributed to the final paper.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information 1
This file contains Supplementary Figures S1-S3, Supplementary Tables 1-11 and additional references. (PDF 1623 kb)
Supplementary Information 2
This file contains Supplementary Data (XLS 83 kb)
Supplementary Information 3
This file contains additional acknowledgements. (PDF 16 kb)
Rights and permissions
About this article
Cite this article
Easton, D., Pooley, K., Dunning, A. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007). https://doi.org/10.1038/nature05887
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05887
This article is cited by
-
An apparent quandary: adoption of polygenics and gene panels for personalised breast cancer risk stratification
BJC Reports (2023)
-
DNA methylation at quantitative trait loci (mQTLs) varies with cell type and nonheritable factors and may improve breast cancer risk assessment
npj Precision Oncology (2023)
-
Screening properties of trend tests in genetic association studies
Scientific Reports (2023)
-
MAP3K1 expression is associated with progression and poor prognosis of hormone receptor-positive, HER2-negative early-stage breast cancer
Cellular Oncology (2023)
-
MTHSA-DHEI: multitasking harmony search algorithm for detecting high-order SNP epistatic interactions
Complex & Intelligent Systems (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.