Abstract
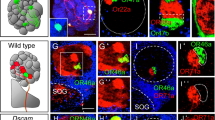
Sensory processing centres in both the vertebrate and the invertebrate brain are often organized into reiterated columns, thus facilitating an internal topographic representation of the external world. Cells within each column are arranged in a stereotyped fashion and form precise patterns of synaptic connections within discrete layers. These connections are largely confined to a single column, thereby preserving the spatial information from the periphery. Other neurons integrate this information by connecting to multiple columns. Restricting axons to columns is conceptually similar to tiling. Axons and dendrites of neighbouring neurons of the same class use tiling to form complete, yet non-overlapping, receptive fields1,2,3. It is thought that, at the molecular level, cell-surface proteins mediate tiling through contact-dependent repulsive interactions1,2,4,5, but proteins serving this function have not yet been identified. Here we show that the immunoglobulin superfamily member Dscam2 restricts the connections formed by L1 lamina neurons to columns in the Drosophila visual system. Our data support a model in which Dscam2 homophilic interactions mediate repulsion between neurites of L1 cells in neighbouring columns. We propose that Dscam2 is a tiling receptor for L1 neurons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Blackshaw, S. E., Nicholls, J. G. & Parnas, I. Expanded receptive fields of cutaneous mechanoreceptor cells after single neurone deletion in leech central nervous system. J. Physiol. (Lond.) 326, 261–268 (1982)
Grueber, W. B., Ye, B., Moore, A. W., Jan, L. Y. & Jan, Y. N. Dendrites of distinct classes of Drosophila sensory neurons show different capacities for homotypic repulsion. Curr. Biol. 13, 618–626 (2003)
Sagasti, A., Guido, M. R., Raible, D. W. & Schier, A. F. Repulsive interactions shape the morphologies and functional arrangement of zebrafish peripheral sensory arbors. Curr. Biol. 15, 804–814 (2005)
Jan, Y. N. & Jan, L. Y. The control of dendrite development. Neuron 40, 229–242 (2003)
Kramer, A. P. & Stent, G. S. Developmental arborization of sensory neurons in the leech Haementeria ghilianii. II. Experimentally induced variations in the branching pattern. J. Neurosci. 5, 768–775 (1985)
Clandinin, T. R. & Zipursky, S. L. Making connections in the fly visual system. Neuron 35, 827–841 (2002)
Meinertzhagen, I. A. & Hanson Thomas, E. The Development of Drosophila melanogaster (Cold Spring Harbor Laboratory Press, New York, 1993)
Barlow, G. M., Lyons, G. E., Richardson, J. A., Sarnat, H. B. & Korenberg, J. R. DSCAM: an endogenous promoter drives expression in the developing CNS and neural crest. Biochem. Biophys. Res. Commun. 299, 1–6 (2002)
Fusaoka, E., Inoue, T., Mineta, K., Agata, K. & Takeuchi, K. Structure and function of primitive immunoglobulin superfamily neural cell adhesion molecules: a lesson from studies on planarian. Genes Cells 11, 541–555 (2006)
Graveley, B. R. et al. The organization and evolution of the dipteran and hymenopteran Down syndrome cell adhesion molecule (Dscam) genes. RNA 10, 1499–1506 (2004)
Yamakawa, K. et al. DSCAM: a novel member of the immunoglobulin superfamily maps in a Down syndrome region and is involved in the development of the nervous system. Hum. Mol. Genet. 7, 227–237 (1998)
Schmucker, D. et al. Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity. Cell 101, 671–684 (2000)
Chen, B. E. et al. The molecular diversity of Dscam is functionally required for neuronal wiring specificity in Drosophila. Cell 125, 607–620 (2006)
Hummel, T. et al. Axonal targeting of olfactory receptor neurons in Drosophila is controlled by Dscam. Neuron 37, 221–231 (2003)
Wang, J., Zugates, C. T., Liang, I. H., Lee, C. H. & Lee, T. Drosophila Dscam is required for divergent segregation of sister branches and suppresses ectopic bifurcation of axons. Neuron 33, 559–571 (2002)
Zhu, H. et al. Dendritic patterning by Dscam and synaptic partner matching in the Drosophila antennal lobe. Nature Neurosci. 9, 349–355 (2006)
Zipursky, S. L., Wojtowicz, W. M. & Hattori, D. Got diversity? Wiring the fly brain with Dscam. Trends Biochem. Sci. 31, 581–588 (2006)
Wojtowicz, W. M., Flanagan, J. J., Millard, S. S., Zipursky, S. L. & Clemens, J. C. Alternative splicing of Drosophila Dscam generates axon guidance receptors that exhibit isoform-specific homophilic binding. Cell 118, 619–633 (2004)
Matthews, B. J. et al. Dendrite self-avoidance is controlled by Dscam. Cell 129, 593–604 (2007)
Hughes, M. E. et al. Homophilic Dscam interactions control complex dendrite morphology. Neuron 54, 417–427 (2007)
Soba, P. et al. Drosophila sensory neurons require Dscam for dendritic self-avoidance and proper dendritic field organization. Neuron 54, 403–416 (2007)
Gong, W. J. & Golic, K. G. Ends-out, or replacement, gene targeting in Drosophila. Proc. Natl Acad. Sci. USA 100, 2556–2561 (2003)
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999)
Pappu, K. S. et al. Dual regulation and redundant function of two eye-specific enhancers of the Drosophila retinal determination gene dachshund. Development 132, 2895–2905 (2005)
Grueber, W. B., Jan, L. Y. & Jan, Y. N. Tiling of the Drosophila epidermis by multidendritic sensory neurons. Development 129, 2867–2878 (2002)
Lee, T., Winter, C., Marticke, S. S., Lee, A. & Luo, L. Essential roles of Drosophila RhoA in the regulation of neuroblast proliferation and dendritic but not axonal morphogenesis. Neuron 25, 307–316 (2000)
Zhan, X. L. et al. Analysis of Dscam diversity in regulating axon guidance in Drosophila mushroom bodies. Neuron 43, 673–686 (2004)
Neves, G., Zucker, J., Daly, M. & Chess, A. Stochastic yet biased expression of multiple Dscam splice variants by individual cells. Nature Genet. 36, 240–246 (2004)
Lee, C. H., Herman, T., Clandinin, T. R., Lee, R. & Zipursky, S. L. N-cadherin regulates target specificity in the Drosophila visual system. Neuron 30, 437–450 (2001)
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker (MARCM) for Drosophila neural development. Trends Neurosci. 24, 251–254 (2001)
Acknowledgements
We thank E. DeRobertis, U. Banerjee, W. Grueber, I. Meinertzhagen, A. Sagasti and members of the Zipursky laboratory for comments on the manuscript; J. Clemens for identifying and characterizing the Dscam paralogues; D. Gunning for cloning Dscam3 and Dscam4; G. Marden for providing unpublished reagents that allowed us to generate the Dac-FLP construct; and Y. Zhu and R. Imondi for identifying the two medulla neuron Gal4 lines used in this study. S.L.Z. is an investigator of the Howard Hughes Medical Institute. This work was also supported by the NIH (S.L.Z). S.S.M. was supported by a Cellular Neurobiology training grant from the NIH.
The Dscam2–4 sequences can be found in the NCBI database under accession numbers AE014296, AE003718 and AE003556, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The Dscam2–4 sequences can be found in the NCBI database under accession numbers AE014296, AE003718 and AE003556, respectively. The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures S1-S2 with Legends. The Supplementary Figure S1 shows confocal images of Dscam2 mutant phenotypes in three different classes of neurons that project into the medulla (T1 and C3 medulla neurons and lamina neurons). The Supplementary Figure S2 shows confocal images of 10 examples of wild type L1 lamina neurons compared to autonomous and non-autonomous Dscam2 phenotypes generated by MARCM and reverse MARCM. (PDF 2085 kb)
Rights and permissions
About this article
Cite this article
Millard, S., Flanagan, J., Pappu, K. et al. Dscam2 mediates axonal tiling in the Drosophila visual system. Nature 447, 720–724 (2007). https://doi.org/10.1038/nature05855
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature05855
This article is cited by
-
A novel method for identifying key genes in macroevolution based on deep learning with attention mechanism
Scientific Reports (2023)
-
Revisiting Dscam diversity: lessons from clustered protocadherins
Cellular and Molecular Life Sciences (2019)
-
DSCAM differentially modulates pre- and postsynaptic structural and functional central connectivity during visual system wiring
Neural Development (2018)
-
Strategies for assembling columns and layers in the Drosophila visual system
Neural Development (2018)
-
Coping with living in the soil: the genome of the parthenogenetic springtail Folsomia candida
BMC Genomics (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.