Abstract
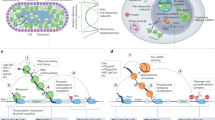
Bacteria make extensive use of riboswitches1,2 to sense metabolites and control gene expression, and typically do so by modulating premature transcription termination or translation initiation. The most widespread riboswitch class known in bacteria responds to the coenzyme thiamine pyrophosphate (TPP)3,4, which is a derivative of vitamin B1. Representatives of this class have also been identified5,6 in fungi and plants, where they are predicted5,7 to control messenger RNA splicing or processing. We examined three TPP riboswitches in the filamentous fungus Neurospora crassa, and found that one activates and two repress gene expression by controlling mRNA splicing. A detailed mechanism involving riboswitch-mediated base-pairing changes and alternative splicing control was elucidated for precursor NMT1 mRNAs, which code for a protein involved in TPP metabolism. These results demonstrate that eukaryotic cells employ metabolite-binding RNAs to regulate RNA splicing events that are important for the control of key biochemical processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mandal, M. & Breaker, R. R. Gene regulation by riboswitches. Nature Rev. Mol. Cell Biol. 5, 451–463 (2004)
Winkler, W. C. & Breaker, R. R. Regulation of bacterial gene expression by riboswitches. Annu. Rev. Microbiol. 59, 487–517 (2005)
Winkler, W. C., Nahvi, A. & Breaker, R. R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 419, 952–956 (2002)
Mironov, A. S. et al. Sensing small molecules by nascent RNA: a mechanism to control transcription in bacteria. Cell 111, 747–756 (2002)
Sudarsan, N., Barrick, J. E. & Breaker, R. R. Metabolite-binding RNA domains are present in the genes of eukaryotes. RNA 9, 644–647 (2003)
Galagan, J. E. et al. Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature 438, 1105–1115 (2005)
Kubodera, T. et al. Thiamine-regulated gene expression of Aspergillus oryzae thiA requires splicing of the intron containing a riboswitch-like domain in the 5′-UTR. FEBS Lett. 555, 516–520 (2003)
McColl, D., Valencia, C. A. & Vierula, P. J. Characterization and expression of the Neurospora crassa nmt-1 gene. Curr. Genet. 44, 216–223 (2003)
Faou, P. & Tropschug, M. A novel binding protein for a member of CyP40-type Cyclophilins: N. crassa CyPBP37, a growth and thiamine regulated protein homolog to yeast Thi4p. J. Mol. Biol. 333, 831–844 (2003)
Faou, P. & Tropschug, M. Neurospora crassa CyPBP37: a cytosolic stress protein that is able to replace yeast Thi4p function in the synthesis of vitamin B1. J. Mol. Biol. 344, 1147–1157 (2004)
Maundrell, K. nmt1 of fission yeast: a highly expressed gene completely repressed by thiamine. J. Biol. Chem. 265, 10857–10864 (1989)
Soukup, G. A. & Breaker, R. R. Relationship between internucleotide linkage geometry and the stability of RNA. RNA 5, 1308–1325 (1999)
Thore, S., Leibundgut, M. & Ban, N. Structure of the eukaryotic thiamine pyrophosphate riboswitch with its regulatory ligand. Science 312, 1208–1211 (2006)
Serganov, A., Polonskaia, A., Phan, A. T., Breaker, R. R. & Patel, D. J. Structural basis for gene regulation by a thiamine pyrophosphate-sensing riboswitch. Nature 441, 1167–1171 (2006)
Edwards, T. E. & Ferré-D’Amaré, A. R. Crystal structures of the Thi-box riboswitch bound to thiamine pyrophosphate analogs reveal adaptive RNA–small molecule recognition. Structure 14, 1459–1468 (2006)
Welz, R. & Breaker, R. R. Ligand binding and gene control characteristics of tandem riboswitches in Bacillus anthracis. RNA advance online publication (16 February, doi: 10.1261/rna.407707 2007)
Vilela, C. & McCarthy, J. E. Regulation of fungal gene expression via short open reading frames in the mRNA 5′ untranslated region. Mol. Microbiol. 49, 859–867 (2003)
Nahvi, A. et al. Genetic control by a metabolite binding mRNA. Chem. Biol. 9, 1043–1049 (2002)
Romfo, C. M., Alvarez, C. J., van Heeckeren, W. J., Webb, C. J. & Wise, J. A. Evidence for splice site pairing via intron definition in Schizosaccharomyces pombe. Mol. Cell. Biol. 20, 7955–7970 (2000)
Matlin, A. J., Clark, F. & Smith, C. W. Understanding alternative splicing: towards a cellular code. Nature Rev. Mol. Cell Biol. 6, 386–398 (2005)
Blencowe, B. J. Alternative splicing: new insights from global analyses. Cell 126, 37–47 (2006)
Buratti, E. & Baralle, F. E. Influence of RNA secondary structure on the pre-mRNA splicing process. Mol. Cell. Biol. 24, 10505–10514 (2004)
Colot, H. V., Loros, J. J. & Dunlap, J. C. Temperature-modulated alternative splicing and promoter use in the circadian clock gene frequency. Mol. Biol. Cell 16, 5563–5571 (2005)
Borsuk, P. et al. L-Arginine influences the structure and function of arginase mRNA in Aspergillus nidulans. Biol. Chem. 388, 135–144 (2007)
Kim, D.-S., Gusti, V., Pillai, S. G. & Gaur, R. K. An artificial riboswitch for controlling pre-mRNA splicing. RNA 11, 1667–1677 (2005)
Eddy, S. R. & Durbin, R. RNA sequence analysis using covariance models. Nucleic Acids Res. 22, 2079–2088 (1994)
Eddy, S. R. INFERNAL. Version 0.55. Distributed by the author. Department of Genetics, Washington Univ. School of Medicine, St. Louis, Missouri.
Seetharaman, S., Zivarts, M., Sudarsan, N. & Breaker, R. R. Immobilized RNA switches for the analysis of complex chemical and biological mixtures. Nature Biotechnol. 19, 336–341 (2001)
Froehlich, A. C., Loros, J. J. & Dunlap, J. C. Rhythmic binding of a WHITE COLLAR-containing complex to the frequency promoter is inhibited by FREQUENCY. Proc. Natl Acad. Sci. USA 100, 5914–5919 (2003)
Mehra, A., Morgan, L., Bell-Pedersen, D., Loros, J. & Dunlap, J. C. Watching the Neurospora clock tick. Soc. Res. Biol. Rhythms 27 (Amelia Island, Florida, Society for Research on Biological Rhythms, 22–25 May, 2002)
Orbach, M. J., Porro, E. B. & Yanofsky, C. Cloning and characterization of the gene for β-tubulin from a benomyl-resistant mutant of Neurospora crassa and its use as a dominant selectable marker. Mol. Cell. Biol. 6, 2452–2461 (1986)
Westergaard, M. & Mitchell, H. K. Neurospora V. A synthetic medium favoring sexual reproduction. Am. J. Bot. 34, 573–577 (1947)
Davis, R. H. Neurospora: Contributions of a Model Organism. (Oxford Univ. Press, New York, New York, 2000)
Loros, J. J. & Dunlap, J. C. Neurospora crassa clock-controlled genes are regulated at the level of transcription. Mol. Cell. Biol. 11, 558–563 (1991)
Vann, D. C. Electroporation-based transformation of freshly harvested conidia of Neurospora crassa. Fungal Genet. Newsl. 42A, 53 (1995)
Ebbole, D. & Sachs, M. S. A rapid and simple method for isolation of Neurospora crassa homokaryons using microconidia. Fungal Genet. Newsl. 37, 17–18 (1990)
Acknowledgements
. We thank J. C. Dunlap for hosting and training M.T.C. in fungal genetics techniques and for supplying us with N. crassa strains, genomic DNA of N. crassa and the luciferase reporter vector. Also, we thank members of the Dunlap laboratory (M. Shi and L. Larrondo) for technical assistance and helpful discussions. A.W. was supported by a postdoctoral fellowship from the German Research Foundation (DFG) and M.T.C. was supported by a Yale College Dean’s Fellowship and an HHMI Future Scientist Fellowship. This work also was supported by an NIH grant (R.R.B).
Author Contributions M.T.C. and A.W. conducted all genetic and molecular biology analyses, and M.T.C. and N.S. conducted biochemical analyses. M.T.C., A.W. and R.R.B. co-wrote the manuscript. All authors contributed to discussions regarding the data and their interpretation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
[Competing Interests Statement: ‘R.R.B. is a cofounder of BioRelix, a company interested in using riboswitches as antimicrobial drug targets.’]
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Figures S1-S10 with Legends and additional references. (PDF 1154 kb)
Rights and permissions
About this article
Cite this article
Cheah, M., Wachter, A., Sudarsan, N. et al. Control of alternative RNA splicing and gene expression by eukaryotic riboswitches. Nature 447, 497–500 (2007). https://doi.org/10.1038/nature05769
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05769
This article is cited by
-
Structure-based insights into recognition and regulation of SAM-sensing riboswitches
Science China Life Sciences (2023)
-
Getting to the bottom of lncRNA mechanism: structure–function relationships
Mammalian Genome (2022)
-
Translational control of enzyme scavenger expression with toxin-induced micro RNA switches
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.