Abstract
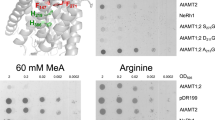
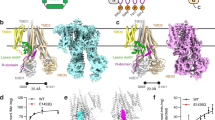
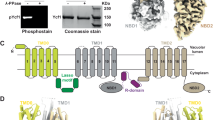
Polytopic membrane proteins are essential for cellular uptake and release of nutrients. To prevent toxic accumulation, rapid shut-off mechanisms are required. Here we show that the soluble cytosolic carboxy terminus of an oligomeric ammonium transporter from Arabidopsis thaliana serves as an allosteric regulator essential for function; mutations in the C-terminal domain, conserved between bacteria, fungi and plants, led to loss of transport activity. When co-expressed with intact transporters, mutants inactivated functional subunits, but left their stability unaffected. Co-expression of two inactive transporters, one with a defective pore, the other with an ablated C terminus, reconstituted activity. The crystal structure of an Archaeoglobus fulgidus ammonium transporter (AMT)1 suggests that the C terminus interacts physically with cytosolic loops of the neighbouring subunit. Phosphorylation of conserved sites in the C terminus2 are proposed as the cognate control mechanism. Conformational coupling between monomers provides a mechanism for tight regulation, for increasing the dynamic range of sensing and memorizing prior events, and may be a general mechanism for transporter regulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Andrade, S. L., Dickmanns, A., Ficner, R. & Einsle, O. Crystal structure of the archaeal ammonium transporter Amt-1 from Archaeoglobus fulgidus. Proc. Natl Acad. Sci. USA 102, 14994–14999 (2005)
Nühse, T. S., Stensballe, A., Jensen, O. N. & Peck, S. C. Phosphoproteomics of the Arabidopsis plasma membrane and a new phosphorylation site database. Plant Cell 16, 2394–2405 (2004)
Blakey, D. et al. Purification of the Escherichia coli ammonium transporter AmtB reveals a trimeric stoichiometry. Biochem. J. 364, 527–535 (2002)
Ludewig, U. et al. Homo- and hetero-oligomerization of ammonium transporter-1 NH4+ uniporters. J. Biol. Chem. 278, 45603–45610 (2003)
Khademi, S. et al. Mechanism of ammonia transport by Amt/MEP/Rh: structure of AmtB at 1.35 A. Science 305, 1587–1594 (2004)
Veenhoff, L. M., Heuberger, E. H. & Poolman, B. Quaternary structure and function of transport proteins. Trends Biochem. Sci. 27, 242–249 (2002)
Jiang, Y. et al. X-ray structure of a voltage-dependent K+ channel. Nature 423, 33–41 (2003)
Borgnia, M., Nielsen, S., Engel, A. & Agre, P. Cellular and molecular biology of the aquaporin water channels. Annu. Rev. Biochem. 68, 425–458 (1999)
Grasberger, B., Minton, A. P., DeLisi, C. & Metzger, H. Interaction between proteins localized in membranes. Proc. Natl Acad. Sci. USA 83, 6258–6262 (1986)
Hamill, S., Cloherty, E. K. & Carruthers, A. The human erythrocyte sugar transporter presents two sugar import sites. Biochemistry 38, 16974–16983 (1999)
Marini, A. M., Springael, J. Y., Frommer, W. B. & André, B. Cross-talk between ammonium transporters in yeast and interference by the soybean SAT1 protein. Mol. Microbiol. 35, 378–385 (2000)
Schulman, H., Hanson, P. I. & Meyer, T. Decoding calcium signals by multifunctional CaM kinase. Cell Calcium 13, 401–411 (1992)
Coutts, G., Thomas, G., Blakey, D. & Merrick, M. Membrane sequestration of the signal transduction protein GlnK by the ammonium transporter AmtB. EMBO J. 21, 536–545 (2002)
Conroy, M. J. et al. Electron and atomic force microscopy of the trimeric ammonium transporter AmtB. EMBO Rep. 5, 1153–1158 (2004)
Zen, K. H., McKenna, E., Bibi, E., Hardy, D. & Kaback, H. R. Expression of lactose permease in contiguous fragments as a probe for membrane-spanning domains. Biochemistry 33, 8198–8206 (1994)
Reinders, A. et al. Intra- and intermolecular interactions in sucrose transporters at the plasma membrane detected by the split-ubiquitin system and functional assays. Structure 10, 763–772 (2002)
Schulze, W. X., Reinders, A., Ward, J., Lalonde, S. & Frommer, W. B. Interactions between co-expressed Arabidopsis sucrose transporters in the split-ubiquitin system. BMC Biochem. 4, 3 (2003)
Kronzucker, H. J., Siddiqi, M. Y. & Glass, A. Kinetics of NH4+ influx in spruce. Plant Physiol. 110, 773–779 (1996)
Rawat, S. R., Silim, S. N., Kronzucker, H. J., Siddiqi, M. Y. & Glass, A. D. AtAMT1 gene expression and NH4+ uptake in roots of Arabidopsis thaliana: evidence for regulation by root glutamine levels. Plant J. 19, 143–152 (1999)
Marini, A. M., Soussi-Boudekou, S., Vissers, S. & Andre, B. A family of ammonium transporters in Saccharomyces cerevisiae. Mol. Cell. Biol. 17, 4282–4293 (1997)
Ludewig, U., von Wiren, N. & Frommer, W. B. Uniport of NH4+ by the root hair plasma membrane ammonium transporter LeAMT1;1. J. Biol. Chem. 277, 13548–13555 (2002)
Ninnemann, O., Jauniaux, J. C. & Frommer, W. B. Identification of a high affinity NH4+ transporter from plants. EMBO J. 13, 3464–3471 (1994)
Javelle, A., Severi, E., Thornton, J. & Merrick, M. Ammonium sensing in Escherichia coli. Role of the ammonium transporter AmtB and AmtB–GlnK complex formation. J. Biol. Chem. 279, 8530–8538 (2004)
Marini, A. M., Boeckstaens, M., Benjelloun, F., Cherif-Zahar, B. & Andre, B. Structural involvement in substrate recognition of an essential aspartate residue conserved in Mep/Amt and Rh-type ammonium transporters. Curr. Genet. 49, 364–374 (2006)
Changeux, J. P. & Edelstein, S. J. Allosteric mechanisms of signal transduction. Science 308, 1424–1428 (2005)
Törnroth-Horsefield, S. et al. Structural mechanism of plant aquaporin gating. Nature 439, 688–694 (2006)
Palmgren, M. G., Sommarin, M., Serrano, R. & Larsson, C. Identification of an autoinhibitory domain in the C-terminal region of the plant plasma membrane H+-ATPase. J. Biol. Chem. 266, 20470–20475 (1991)
Curran, A. C. et al. Autoinhibition of a calmodulin-dependent calcium pump involves a structure in the stalk that connects the transmembrane domain to the ATPase catalytic domain. J. Biol. Chem. 275, 30301–30308 (2000)
Pittman, J. K. & Hirschi, K. D. Regulation of CAX1, an Arabidopsis Ca2+/H+ antiporter. Identification of an N-terminal autoinhibitory domain. Plant Physiol. 127, 1020–1029 (2001)
Levy, J. et al. A putative Ca2+ and calmodulin-dependent protein kinase required for bacterial and fungal symbioses. Science 303, 1361–1364 (2004)
Acknowledgements
We would like to thank L. Yuan (University of Hohenheim) for the Arabidopsis AMT1;1 antiserum. This work was made possible by grants from the Department of Energy and the European Science award from the Körber Foundation to W.B.F.
Author Contributions D.L. created all mutants, developed the co-expression system and did the protein gel blots, S.L. generated GFP fusions and did the imaging, L.L.L. performed structural modelling, N.vW. contributed to production of the serum and was involved in developing the concept. All authors contributed sections of the manuscript. W.B.F. is responsible for the experimental design, developed the hypotheses, and interpreted the results.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Table S1, Supplementary Figures S1-S16 with Legends and additional references. (PDF 1843 kb)
Rights and permissions
About this article
Cite this article
Loqué, D., Lalonde, S., Looger, L. et al. A cytosolic trans-activation domain essential for ammonium uptake. Nature 446, 195–198 (2007). https://doi.org/10.1038/nature05579
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05579
This article is cited by
-
Crosstalk between brassinosteroid signaling and variable nutrient environments
Science China Life Sciences (2023)
-
Tissue specific expression of UMAMIT amino acid transporters in wheat
Scientific Reports (2022)
-
Transcriptomic and genome-wide association study reveal long noncoding RNAs responding to nitrogen deficiency in maize
BMC Plant Biology (2021)
-
Role of protein phosphatases in the regulation of nitrogen nutrition in plants
Physiology and Molecular Biology of Plants (2021)
-
Feedback inhibition of AMT1 NH4+-transporters mediated by CIPK15 kinase
BMC Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.