Abstract
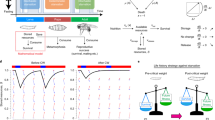
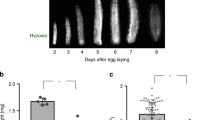
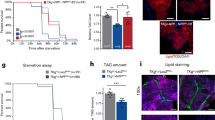
Lipid metabolism is essential for growth and generates much of the energy needed during periods of starvation. In Drosophila, fasting larvae release large quantities of lipid from the fat body but it is unclear how and where this is processed. Here we identify the oenocyte as the principal cell type accumulating lipid droplets during starvation. Tissue-specific manipulations of the Slimfast amino-acid channel, the Lsd2 fat-storage regulator and the Brummer lipase indicate that oenocytes act downstream of the fat body. In turn, oenocytes are required for depleting stored lipid from the fat body during fasting. Hence, lipid-metabolic coupling between the fat body and oenocytes is bidirectional. When food is plentiful, oenocytes have critical roles in regulating growth, development and feeding behaviour. In addition, they specifically express many different lipid-metabolizing proteins, including Cyp4g1, an ω-hydroxylase regulating triacylglycerol composition. These findings provide evidence that some lipid-processing functions of the mammalian liver are performed in insects by oenocytes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Gibbons, G. F., Islam, K. & Pease, R. J. Mobilisation of triacylglycerol stores. Biochim. Biophys. Acta 1483, 37–57 (2000)
Lee, C. H., Olson, P. & Evans, R. M. Minireview: lipid metabolism, metabolic diseases, and peroxisome proliferator-activated receptors. Endocrinology 144, 2201–2207 (2003)
Yu, S., Rao, S. & Reddy, J. K. Peroxisome proliferator-activated receptors, fatty acid oxidation, steatohepatitis and hepatocarcinogenesis. Curr. Mol. Med. 3, 561–572 (2003)
Zechner, R., Strauss, J. G., Haemmerle, G., Lass, A. & Zimmermann, R. Lipolysis: pathway under construction. Curr. Opin. Lipidol. 16, 333–340 (2005)
Reddy, J. K. & Hashimoto, T. Peroxisomal β-oxidation and peroxisome proliferator-activated receptor-α: an adaptive metabolic system. Annu. Rev. Nutr. 21, 193–230 (2001)
Wanders, R. J. Peroxisomes, lipid metabolism, and peroxisomal disorders. Mol. Genet. Metab. 83, 16–27 (2004)
Ntambi, J. M. & Miyazaki, M. Regulation of stearoyl-CoA desaturases and role in metabolism. Prog. Lipid Res. 43, 91–104 (2004)
Wanders, R. J. et al. Peroxisomal fatty acid α- and β-oxidation in health and disease: new insights. Adv. Exp. Med. Biol. 544, 293–302 (2003)
Browning, J. D. & Horton, J. D. Molecular mediators of hepatic steatosis and liver injury. J. Clin. Invest. 114, 147–152 (2004)
McKay, R. M., McKay, J. P., Avery, L. & Graff, J. M. C. elegans: a model for exploring the genetics of fat storage. Dev. Cell 4, 131–142 (2003)
Ashrafi, K. et al. Genome-wide RNAi analysis of Caenorhabditis elegans fat regulatory genes. Nature 421, 268–272 (2003)
Butterworth, F. M., Bodenstein, D. & King, R. C. Adipose tissue of Drosophila melanogaster. I. An experimental study of larval fat body. J. Exp. Zool. 158, 141–153 (1965)
Keeley, L. L. in Comprehensive Insect Physiology, Biochemistry and Pharmacology (eds Kerkut, G. A. & Gilbert, L. I.) 211–248 (Pergamonn, New York, 1985)
Dean, R. L., Locke, M. & Collins, J. V. in Comprehensive Insect Physiology, Biochemistry and Pharmacology (eds Kerkut, G. A. & Gilbert, L. I.) 155–210 (Pergamonn, New York, 1985)
Canavoso, L. E., Jouni, Z. E., Karnas, K. J., Pennington, J. E. & Wells, M. A. Fat metabolism in insects. Annu. Rev. Nutr. 21, 23–46 (2001)
Dantuma, N. P. et al. An insect homolog of the vertebrate very low density lipoprotein receptor mediates endocytosis of lipophorins. J. Lipid Res. 40, 973–978 (1999)
Lee, C. S. et al. Wax moth, Galleria mellonella, high density lipophorin receptor: alternative splicing, tissue-specific expression, and developmental regulation. Insect Biochem. Mol. Biol. 33, 761–771 (2003)
Zhang, H., Stallock, J. P., Ng, J. C., Reinhard, C. & Neufeld, T. P. Regulation of cellular growth by the Drosophila target of rapamycin dTOR. Genes Dev. 14, 2712–2724 (2000)
Colombani, J. et al. A nutrient sensor mechanism controls Drosophila growth. Cell 114, 739–749 (2003)
Gronke, S. et al. Brummer lipase is an evolutionary conserved fat storage regulator in Drosophila.. Cell Metab. 1, 323–330 (2005)
Gronke, S. et al. Control of fat storage by a Drosophila PAT domain protein. Curr. Biol. 13, 603–606 (2003)
Teixeira, L., Rabouille, C., Rorth, P., Ephrussi, A. & Vanzo, N. F. Drosophila Perilipin/ADRP homologue Lsd2 regulates lipid metabolism. Mech. Dev. 120, 1071–1081 (2003)
Britton, J. S. & Edgar, B. A. Environmental control of the cell cycle in Drosophila: nutrition activates mitotic and endoreplicative cells by distinct mechanisms. Development 125, 2149–2158 (1998)
Martin, J. F., Hersperger, E., Simcox, A. & Shearn, A. minidiscs encodes a putative amino acid transporter subunit required non-autonomously for imaginal cell proliferation. Mech. Dev. 92, 155–167 (2000)
Britton, J. S., Lockwood, W. K., Li, L., Cohen, S. M. & Edgar, B. A. Drosophila's insulin/PI3-kinase pathway coordinates cellular metabolism with nutritional conditions. Dev. Cell 2, 239–249 (2002)
Lillie, R. D. Various oil soluble dyes as fat stains in the supersaturated isopropanol technic. Stain Technol. 19, 55–58 (1944)
Lawrence, P. A. & Johnston, P. Observations on cell lineage of internal organs of Drosophila.. J. Embryol. Exp. Morphol. 91, 251–266 (1986)
Elstob, P. R., Brodu, V. & Gould, A. P. spalt-dependent switching between two cell fates that are induced by the Drosophila EGF receptor. Development 128, 723–732 (2001)
Rusten, T. E. et al. Spalt restricts EGFR mediated induction of chordotonal precursors in the embryonic PNS of Drosophila.. Development 128, 711–722 (2001)
Brodu, V., Elstob, P. R. & Gould, A. P. EGF receptor signaling regulates pulses of cell delamination from the Drosophila ectoderm. Dev. Cell 7, 885–895 (2004)
Cherbas, L., Hu, X., Zhimulev, I., Belyaeva, E. & Cherbas, P. EcR isoforms in Drosophila: testing tissue-specific requirements by targeted blockade and rescue. Development 130, 271–284 (2003)
Wu, Q., Zhang, Y., Xu, J. & Shen, P. Regulation of hunger-driven behaviors by neural ribosomal S6 kinase in Drosophila. Proc. Natl Acad. Sci. USA 102, 13289–13294 (2005)
Beadle, G. W., Tatum, E. L. & Clancy, C. W. Food level in relation to rate of development and eye pigmentation in Drosophila. Biol. Bull. 75, 447–462 (1938)
Riddiford, L. M. in The Development of Drosophila melanogaster (eds Bate, M. & Martinez Arias, A.) 899–939 (Cold Spring Harbor Laboratory Press, New York, 1993)
Gilbert, L. I. Halloween genes encode P450 enzymes that mediate steroid hormone biosynthesis in Drosophila melanogaster. Mol. Cell. Endocrinol. 215, 1–10 (2004)
McGuire, S. E., Le, P. T., Osborn, A. J., Matsumoto, K. & Davis, R. L. Spatiotemporal rescue of memory dysfunction in Drosophila. Science 302, 1765–1768 (2003)
Ueyama, M., Chertemps, T., Labeur, C. & Wicker-Thomas, C. Mutations in the desat1 gene reduces the production of courtship stimulatory pheromones through a marked effect on fatty acids in Drosophila melanogaster. Insect Biochem. Mol. Biol. 35, 911–920 (2005)
Sundseth, S. S., Nix, C. E. & Waters, L. C. Isolation of insecticide resistance-related forms of cytochrome P-450 from Drosophila melanogaster. Biochem. J. 265, 213–217 (1990)
Griswold, C. M., Matthews, A. L., Bewley, K. E. & Mahaffey, J. W. Molecular characterization and rescue of acatalasemic mutants of Drosophila melanogaster. Genetics 134, 781–788 (1993)
Simpson, A. E. The cytochrome P450 4 (CYP4) family. Gen. Pharmacol. 28, 351–359 (1997)
Tomancak, P. et al. Systematic determination of patterns of gene expression during Drosophila embryogenesis. Genome Biol. 3, RESEARCH0088 (2002)
Landois, L. Ueber die Funktion des Fettkörpers. Zeitschr. f. Wissensch. Zoologie 15, 371–372 (1865)
Koschevnikov, G. Ueber den Fettkörper und die Oenocyten der Honigbiene (Apis mellifera). Z. Anz. Bd. 23, 657–661 (1900)
Koller, G. Die innere Sekretion bei wirbellosen Tieren. Biol. Rev. 4, 269–306 (1928)
Wigglesworth, V. The physiology of the cuticle and of ecdysis in Rhodnius prolixus (Triatomidae, Hemiptera); with special reference to the function of the oenocytes and of the dermal glands. Quart. J. Micros. Sci. 76, 269–318 (1933)
Zinke, I., Kirchner, C., Chao, L. C., Tetzlaff, M. T. & Pankratz, M. J. Suppression of food intake and growth by amino acids in Drosophila: the role of pumpless, a fat body expressed gene with homology to vertebrate glycine cleavage system. Development 126, 5275–5284 (1999)
Oike, Y., Akao, M., Kubota, Y. & Suda, T. Angiopoietin-like proteins: potential new targets for metabolic syndrome therapy. Trends Mol. Med. 11, 473–479 (2005)
Duerden, J. M. & Gibbons, G. F. Secretion and storage of newly synthesized hepatic triacylglycerol fatty acids in vivo in different nutritional states and in diabetes. Biochem. J. 255, 929–935 (1988)
Burdge, G. C., Wright, P., Jones, A. E. & Wootton, S. A. A method for separation of phosphatidylcholine, triacylglycerol, non-esterified fatty acids and cholesterol esters from plasma by solid-phase extraction. Br. J. Nutr. 84, 781–787 (2000)
Acknowledgements
We thank G. Gibbons and K. Frayn for providing advice and facilities for metabolic profiling, and P. Elstob, V. Brodu, S. Mahadevaiah and P. Sarchet for assistance with enhancer trap screening and sequencing. We also thank S. Celniker and BDGP for providing many of the panels used in Supplementary Fig. 4, and R. Barrio, B. Bello, L. Michaut, A. Brand, R. Carthew, W. Janning, S. Kennel, C. O’Kane, R. Kühnlein, P. Leopold, I. Miguel-Aliaga, I. Salecker, R. Schultz, N. Tapon, C. Thummel, J.-P. Vincent, T. Xu, Flyview, The NP consortium, the DGRC at Kyoto Institute of Technology, the Bloomington, Umeå, and Szeged stock centres and the DSHB at the University of Iowa for DNA constructs, flies and antibodies. We also thank I. Robinson for discussions and J. Briscoe, X. Franch-Marro, G. Gibbons, I. Miguel-Aliaga, E. Ober, E. Piddini, I. Salecker, P. Serpente and D. Wilkinson for critical reading of the manuscript. This work was supported by the Medical Research Council (E.G., D.W and A.P.G.), the Mexican National Council for Science and Technology (E.G.) and the Wellcome Trust (B.F.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Figures and Legends 1-4, Supplementary Table 1 and Supplementary Notes. (PDF 3187 kb)
Rights and permissions
About this article
Cite this article
Gutierrez, E., Wiggins, D., Fielding, B. et al. Specialized hepatocyte-like cells regulate Drosophila lipid metabolism. Nature 445, 275–280 (2007). https://doi.org/10.1038/nature05382
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05382
This article is cited by
-
Histone acetyltransferase NAA40 modulates acetyl-CoA levels and lipid synthesis
BMC Biology (2022)
-
Modeling exercise using optogenetically contractible Drosophila larvae
BMC Genomics (2022)
-
Aging-Related Variation of Cuticular Hydrocarbons in Wild Type and Variant Drosophila melanogaster
Journal of Chemical Ecology (2022)
-
Examination of Niddm20 candidate genes of OLETF rats in Drosophila melanogaster using inducible GeneSwitch GAL4 system
Journal of Genetics (2022)
-
Sulfakinins influence lipid composition and insulin-like peptides level in oenocytes of Zophobas atratus beetles
Journal of Comparative Physiology B (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.