Abstract
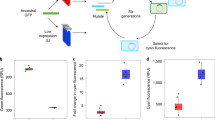
Protein expression is a stochastic process that leads to phenotypic variation among cells1,2,3,4,5,6. The cell–cell distribution of protein levels in microorganisms has been well characterized7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23 but little is known about such variability in human cells. Here, we studied the variability of protein levels in human cells, as well as the temporal dynamics of this variability, and addressed whether cells with higher than average protein levels eventually have lower than average levels, and if so, over what timescale does this mixing occur. We measured fluctuations over time in the levels of 20 endogenous proteins in living human cells, tagged by the gene for yellow fluorescent protein at their chromosomal loci24. We found variability with a standard deviation that ranged, for different proteins, from about 15% to 30% of the mean. Mixing between high and low levels occurred for all proteins, but the mixing time was longer than two cell generations (more than 40 h) for many proteins. We also tagged pairs of proteins with two colours, and found that the levels of proteins in the same biological pathway were far more correlated than those of proteins in different pathways. The persistent memory for protein levels that we found might underlie individuality in cell behaviour and could set a timescale needed for signals to affect fully every member of a cell population.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Spudich, J. L. & Koshland, D. E. Jr. Non-genetic individuality: chance in the single cell. Nature 262, 467–471 (1976)
Novick, A. & Weiner, M. Enzyme induction as an all-or-none phenomenon. Proc. Natl Acad. Sci. USA 43, 553–566 (1957)
Alon, U. An Introduction to Systems Biology: Design Principles of Biological Circuits (Chapman & Hall/CRC Press, Boca Raton, Florida, 2006)
McAdams, H. H. & Arkin, A. Stochastic mechanisms in gene expression. Proc. Natl Acad. Sci. USA 94, 814–819 (1997)
Ferrell, J. E. & Machleder, E. M. The biochemical basis of an all-or-none cell fate switch in Xenopus oocytes. Science 280, 895–898 (1998)
Balaban, N. Q., Merrin, J., Chait, R., Kowalik, L. & Leibler, S. Bacterial persistence as a phenotypic switch. Science 305, 1622–1625 (2004)
Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002)
Bar-Even, A. et al. Noise in protein expression scales with natural protein abundance. Nature Genet. 38, 636–643 (2006)
Newman, J. R. et al. Single-cell proteomic analysis of S. cerevisiae reveals the architecture of biological noise. Nature 441, 840–846 (2006)
Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002)
Pedraza, J. M. & van Oudenaarden, A. Noise propagation in gene networks. Science 307, 1965–1969 (2005)
Rosenfeld, N., Young, J. W., Alon, U., Swain, P. S. & Elowitz, M. B. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005)
Austin, D. W. et al. Gene network shaping of inherent noise spectra. Nature 439, 608–611 (2006)
Raser, J. M. & O'Shea, E. K. Control of stochasticity in eukaryotic gene expression. Science 304, 1811–1814 (2004)
Mihalcescu, I., Hsing, W. & Leibler, S. Resilient circadian oscillator revealed in individual cyanobacteria. Nature 430, 81–85 (2004)
Colman-Lerner, A. et al. Regulated cell-to-cell variation in a cell-fate decision system. Nature 437, 699–706 (2005)
Fraser, H. B., Hirsh, A. E., Giaever, G., Kumm, J. & Eisen, M. B. Noise minimization in eukaryotic gene expression. PLoS Biol. 2, e137 (2004)
Blake, W. J., Kærn, M., Cantor, C. R. & Collins, J. J. Noise in eukaryotic gene expression. Nature 422, 633–637 (2003)
Acar, M., Becskei, A. & van Oudenaarden, A. Enhancement of cellular memory by reducing stochastic transitions. Nature 435, 228–232 (2005)
Volfson, D. et al. Origins of extrinsic variability in eukaryotic gene expression. Nature 439, 861–864 (2005)
Hooshangi, S., Thiberge, S. & Weiss, R. Ultrasensitivity and noise propagation in a synthetic transcriptional cascade. Proc. Natl Acad. Sci. USA 102, 3581–3586 (2005)
Kærn, M., Elston, T. C., Blake, W. J. & Collins, J. J. Stochasticity in gene expression: from theories to phenotypes. Nature Rev. Genet. 6, 451–464 (2005)
Paulsson, J. Summing up the noise in gene networks. Nature 427, 415–418 (2004)
Sigal, A. et al. Dynamic proteomics in individual human cells uncovers widespread cell-cycle dependence of nuclear proteins. Nature Methods 3, 525–531 (2006)
Jarvik, J. W. et al. In vivo functional proteomics: mammalian genome annotation using CD-tagging. Biotechniques 33, 852–4, 856, 858–60 (2002)
Morin, X., Daneman, R., Zavortink, M. & Chia, W. A protein trap strategy to detect GFP-tagged proteins expressed from their endogenous loci in Drosophila. Proc. Natl Acad. Sci. USA 98, 15050–15055 (2001)
Clyne, P. J., Brotman, J. S., Sweeney, S. T. & Davis, G. Green fluorescent protein tagging Drosophila proteins at their native genomic loci with small P elements. Genetics 165, 1433–1441 (2003)
Golding, I., Paulsson, J., Zawilski, S. M. & Cox, E. C. Real-time kinetics of gene activity in individual bacteria. Cell 123, 1025–1036 (2005)
Shaner, N. C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nature Biotechnol. 22, 1567–1572 (2004)
Acknowledgements
We thank the Kahn Family Foundation and the Israel Science Foundation for support. We thank J. Paulsson, J. Pedraza and A. Eldar for discussions of the manuscript, and A. Sharp and E. Ariel for flow cytometry assistance. R.M. thanks the Horowitz Complexity Science Foundation for support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains Supplementary Data, Supplementary Tables 1–4 and Supplementary Discussion. (PDF 208 kb)
Supplementary Figures
This file contains Supplementary Figures 1–5. (PDF 294 kb)
Supplementary Movie 1
Time-lapse movie of transmitted light images of the clone with YFP CD-tagged TOP1, from which frames were obtained for Figure 1b in the article. Movie duration is 100 hours (time-lapse: 1 frame per 10 minutes, each second of movie is ∼ 5 hours of cell growth). (MPG 7931 kb)
Supplementary Movie 2
Time-lapse movie of fluorescence images of the clone with YFP CD-tagged TOP1, shown in Video1, and from which frames were obtained for Figure 1b in the article. Movie duration is 100 hours (time-lapse: 1 frame per 10 minutes, each second of movie is ∼ 5 hours of cell growth). (MPG 2195 kb)
Rights and permissions
About this article
Cite this article
Sigal, A., Milo, R., Cohen, A. et al. Variability and memory of protein levels in human cells. Nature 444, 643–646 (2006). https://doi.org/10.1038/nature05316
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05316
This article is cited by
-
Information-Theoretic Analysis of a Model of CAR-4-1BB-Mediated NFκB Activation
Bulletin of Mathematical Biology (2024)
-
Nonmonotone invasion landscape by noise-aware control of metastasis activator levels
Nature Chemical Biology (2023)
-
Predictive power of non-identifiable models
Scientific Reports (2023)
-
BdLT-Seq as a barcode decay-based method to unravel lineage-linked transcriptome plasticity
Nature Communications (2023)
-
Membrane marker selection for segmenting single cell spatial proteomics data
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.