Abstract
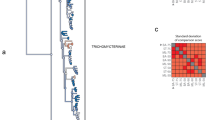
Deuterostomes comprise vertebrates, the related invertebrate chordates (tunicates and cephalochordates) and three other invertebrate taxa: hemichordates, echinoderms and Xenoturbella1. The relationships between invertebrate and vertebrate deuterostomes are clearly important for understanding our own distant origins. Recent phylogenetic studies of chordate classes and a sea urchin have indicated that urochordates might be the closest invertebrate sister group of vertebrates, rather than cephalochordates, as traditionally believed2,3,4,5. More remarkable is the suggestion that cephalochordates are closer to echinoderms than to vertebrates and urochordates, meaning that chordates are paraphyletic2. To study the relationships among all deuterostome groups, we have assembled an alignment of more than 35,000 homologous amino acids, including new data from a hemichordate, starfish and Xenoturbella. We have also sequenced the mitochondrial genome of Xenoturbella. We support the clades Olfactores (urochordates and vertebrates) and Ambulacraria (hemichordates and echinoderms6). Analyses using our new data, however, do not support a cephalochordate and echinoderm grouping and we conclude that chordates are monophyletic. Finally, nuclear and mitochondrial data place Xenoturbella as the sister group of the two ambulacrarian phyla1. As such, Xenoturbella is shown to be an independent phylum, Xenoturbellida, bringing the number of living deuterostome phyla to four.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Bourlat, S. J., Nielsen, C., Lockyer, A. E., Littlewood, D. T. J. & Telford, M. J. Xenoturbella is a deuterostome that eats molluscs. Nature 424, 925–928 (2003)
Delsuc, F., Brinkmann, H., Chourrout, D. & Philippe, H. Tunicates and not cephalochordates are the closest living relatives of vertebrates. Nature 439, 965–968 (2006)
Oda, H. et al. A novel amphioxus cadherin that localises to epithelial adherens junctions has an unusual domain organisation with implications for chordate phylogeny. Evol. Dev. 4, 426–434 (2002)
Wada, H., Okuyama, M., Satoh, N. & Zhang, S. Molecular evolution of fibrillar collagen in chordates, with implications for the evolution of vertebrate skeletons and chordate phylogeny. Evol. Dev. 8, 370–377 (2006)
Jeffery, W. R., Strickler, A. G. & Yamamoto, Y. Migratory neural crest-like cells form body pigmentation in a urochordate embryo. Nature 431, 696–699 (2004)
Turbeville, J. M., Schulz, J. R. & Raff, R. A. Deuterostome phylogeny and the sister group of the chordates: evidence from molecules and morphology. Mol. Biol. Evol. 11, 648–655 (1994)
Telford, M. J. & Holland, P. W. H. The phylogenetic affinities of the chaetognaths: a molecular analysis. Mol. Biol. Evol. 10, 660–676 (1993)
Halanych, K. M. et al. Evidence from 18S ribosomal DNA that the lophophorates are protostome animals. Science 267, 1641–1643 (1995)
Ruppert, E. E. Key characters uniting hemichordates and chordates: homologies or homoplasies?. Can. J. Zool. 83, 8–23 (2005)
Jefferies, R. P. S. in Biological Asymmetry and Handedness (eds Bock, G. R. & Marsh, J.) 94–127 (John Wiley, Chichester, 1991)
Ogasawara, M., Wada, H., Peers, H. & Satoh, N. Developmental expression of Pax 1/9 genes in urochordate and hemichordate gills: insight into function and evolution of the pharyngeal epithelium. Development 126, 2539–2550 (1999)
Rychel, A. L. & Smith, S.E., Shimamoto, H. T. Swalla, B. J. Evolution and development of the chordates: collagen and pharyngeal cartilage. Mol. Biol. Evol. 23, 541–549 (2006)
Telford, M. J., Herniou, E. A., Russell, R. B. & Littlewood, D. T. J. Changes in mitochondrial genetic codes as phylogenetic characters: two examples from the flatworms. Proc. Natl Acad. Sci. USA 97, 11359–11364 (2000)
Stach, T. et al. Nerve cells of Xenoturbella (phylum uncertain) and Harrimania kupfferi (Enteropneusta) are positively immunoreactive to antobodies rasied against echinoderm neuropeptides. J. Mar. Biol. Assoc. UK 85, 1519–1524 (2005)
Reisinger, E. Was ist Xenoturbella?. Z. Wissenschaftlische Zool. 164, 188–198 (1960)
Lowe, C. J. et al. Anteroposterior patterning in hemichordates and the origins of the chordate nervous system. Cell 113, 853–865 (2003)
Philippe, H., Lartillot, N. & Brinkmann, H. Multigene analyses of bilaterian animals corroborate the monophyly of Ecdysozoa, Lophotrochozoa, and Protostomia. Mol. Biol. Evol. 22, 1246–1253 (2005)
Aguinaldo, A. M. A. & Lake, J. A. Evolution of the multicellular animals. Am. Zool. 38, 878–887 (1998)
Jobb, G. TREEFINDER version of May 2006. 〈http://www.treefinder.de〉 (2006)
Phillipe, H. & Telford, M. J. Large-scale sequencing and the new animal phylogeny. Trends Ecol. Evol. (in the press).
Wiens, J. J. Can incomplete taxa rescue phylogenetic analyses from long-branch attraction?. Syst. Biol. 54, 731–742 (2005)
Felsenstein, J. Cases in which parsimony or compatibility methods will be positively misleading. Syst. Zool. 27, 401–410 (1978)
Yokobori, S., Watanabe, Y. & Oshima, T. Mitochondrial genome of Ciona savigny (Urochordata, Ascidiacea, Enterogona): comparison of gene arrangement and tRNA genes with Halocynthia roretzi mitochondrial genome. J. Mol. Evol. 57, 574–587 (2003)
Weichert, C. K. Anatomy of the Chordates (McGraw-Hill, New York, 1965)
Westblad, E. Xenoturbella bocki n.g, n.sp, a peculiar, primitive turbellarian type. Ark. Zool. 1, 3–29 (1949)
Pedersen, K. J. & Pedersen, L. R. Ultrastructural observations on the epidermis of Xenoturbella bocki Westblad, 1949—with a discussion of epidermal cytoplasmic filament systems of invertebrates. Acta Zool. 69, 231–246 (1988)
Raikova, O. I., Reuter, M., Jondelius, U. & Gustafsson, M. K. S. An immunocytochemical and ultrastructural study of the nervous and muscular systems of Xenoturbella westbladi (Bilateria inc. sed.). Zoomorphology 120, 107–118 (2000)
Franzen, A. & Afzelius, B. A. The ciliated epidermis of Xenoturbella bocki (Platyhelminthes, Xenoturbellida) with some phylogenetic considerations. Zool. Scr. 16, 9–17 (1987)
Pardos, F. Fine structure and function of pharynx cilia in Glossobalanus minutus Kowalewsky (Entropneusta). Acta Zool. 69, 1–12 (1988)
Ruiz Trillo, I., Riutort, M., Littlewood, D. T. J., Herniou, E. A. & Baguñà, J. Acoel flatworms: earliest extant bilaterian metazoans, not members of Platyhelminthes. Science 283, 1919–1923 (1999)
Acknowledgements
R.R.C., T.J., M.J.T. and S.J.B. are supported by the EU Marie Curie RTN Zoonet. S.J.B. is also funded by the BBSRC. H.N. was supported by the Human Frontier Science Program Long-Term Fellowship. L.L.M. was supported by NIH and NSF grants as well as the McKnight Brain Research Foundation and the University of Florida opportunity funds. Author Contributions S.J.B. sequenced the mitochondrial genome. C.J.L., R.F., J.A., M.K. and E.S.L. generated the Saccoglossus and Solaster EST data. M.T., H.N., A.B.K., A.H. and L.L.M. generated the Xenoturbella EST data. R.R.C. and T.J. assembled the EST data. M.J.T. analysed the data and led the write-up.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The Xenoturbella mitochondrial genome sequence has GenBank Accession number DQ832701. Xenoturbella ESTs have accession numbers EC906293–EC907475 and novel Saccoglossus ESTs used in this project have accession numbers EE111315–EE122968. Novel Solaster ESTs used in this project have accession numbers EE122969–EE123339. Alignments used are available for download as Supplementary Information or on request from M.J.T. Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Materials
Details of additional Materials and Methods and Supplementary Figures 1 to 5. (PDF 282 kb)
Supplementary Data 1
Nexus format file of aligned and concatenated nuclear EST sequences. Further annotation within file. Part 1 (TXT 717 kb)
Supplementary Data 2
Nexus format file of aligned and concatenated nuclear EST sequences. Further annotation within file. Part 2 (TXT 518 kb)
Supplementary Data 3
Nexus format file of aligned and concatenated nuclear EST sequences. Further annotation within file. Part 3 (TXT 426 kb)
Supplementary Data 4
Nexus format file of aligned EST sequences. This is a concatenation of EST_PART1-3.dat after exclusion of the unreliably aligned positions as indicated in these sub files. Further annotation within file. (TXT 869 kb)
Supplementary Data 5
Nexus format file of aligned and concatenated mitochondrial protein coding gene nucleotide sequences. Further annotation within file. (TXT 795 kb)
Supplementary Data 6
Nexus format file of aligned and concatenated mitochondrial protein coding gene amino acid sequences after exclusion of the unreliably aligned positions as indicated in Supplementary Data 5 and subsequent translation of the remaining nucleotide sequences according to taxon specific genetic codes. Further annotation within file. (TXT 132 kb)
Rights and permissions
About this article
Cite this article
Bourlat, S., Juliusdottir, T., Lowe, C. et al. Deuterostome phylogeny reveals monophyletic chordates and the new phylum Xenoturbellida. Nature 444, 85–88 (2006). https://doi.org/10.1038/nature05241
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05241
This article is cited by
-
Differentially expressed chaperone genes reveal a stress response required for unidirectional regeneration in the basal chordate Ciona
BMC Biology (2023)
-
Cardiopharyngeal deconstruction and ancestral tunicate sessility
Nature (2021)
-
The ontology of the anatomy and development of the solitary ascidian Ciona: the swimming larva and its metamorphosis
Scientific Reports (2020)
-
Molecular dynamics simulation involved in expounding the activation of adrenoceptors by sympathetic nervous system signaling
Structural Chemistry (2020)
-
On the larva and the zooid of the pterobranch Rhabdopleura recondita Beli, Cameron and Piraino, 2018 (Hemichordata, Graptolithina)
Marine Biodiversity (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.