Abstract
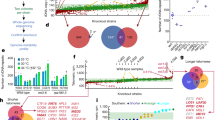
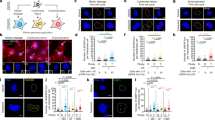
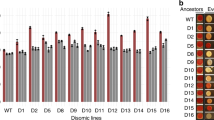
Polyploidy, increased sets of chromosomes, occurs during development, cellular stress, disease and evolution. Despite its prevalence, little is known about the physiological alterations that accompany polyploidy. We previously described ‘ploidy-specific lethality’, where a gene deletion that is not lethal in haploid or diploid budding yeast causes lethality in triploids or tetraploids. Here we report a genome-wide screen to identify ploidy-specific lethal functions. Only 39 out of 3,740 mutations screened exhibited ploidy-specific lethality. Almost all of these mutations affect genomic stability by impairing homologous recombination, sister chromatid cohesion, or mitotic spindle function. We uncovered defects in wild-type tetraploids predicted by the screen, and identified mechanisms by which tetraploidization affects genomic stability. We show that tetraploids have a high incidence of syntelic/monopolar kinetochore attachments to the spindle pole. We suggest that this defect can be explained by mismatches in the ability to scale the size of the spindle pole body, spindle and kinetochores. Thus, geometric constraints may have profound effects on genome stability; the phenomenon described here may be relevant in a variety of biological contexts, including disease states such as cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Comai, L. The advantages and disadvantages of being polyploid. Nature Rev. Genet. 6, 836–846 (2005)
Storchova, Z. & Pellman, D. From polyploidy to aneuploidy, genome instability and cancer. Nature Rev. Mol. Cell Biol. 5, 45–54 (2004)
Otto, S. P. & Whitton, J. Polyploid incidence and evolution. Annu. Rev. Genet. 34, 401–437 (2000)
Taylor, J. S. & Raes, J. Duplication and divergence: The evolution of new genes and old ideas. Annu. Rev. Genet. 38, 615–643 (2004)
Ohno, S., Wolf, U. & Atkin, N. B. Evolution from fish to mammals by gene duplication. Hereditas 59, 169–187 (1968)
Edgar, B. A. & Orr-Weaver, T. L. Endoreplication cell cycles: more for less. Cell 105, 297–306 (2001)
Guidotti, J. E. et al. Liver cell polyploidization: a pivotal role for binuclear hepatocytes. J. Biol. Chem. 278, 19095–19101 (2003)
Akerlund, T., Nordstrom, K. & Bernander, R. Analysis of cell size and DNA content in exponentially growing and stationary-phase batch cultures of Escherichia coli. J. Bacteriol. 177, 6791–6797 (1995)
Mortimer, R. K. Radiobiological and genetic studies on a polyploid series (haploid to hexaploid) of Saccharomyces cerevisiae. Radiat. Res. 9, 312–326 (1958)
Mable, B. K. & Otto, S. P. Masking and purging mutations following EMS treatment in haploid, diploid and tetraploid yeast (Saccharomyces cerevisiae). Genet. Res. 77, 9–26 (2001)
Andalis, A. A. et al. Defects arising from whole-genome duplications in Saccharomyces cerevisiae. Genetics 167, 1109–1121 (2004)
Mayer, V. W. & Aguilera, A. High levels of chromosome instability in polyploids of Saccharomyces cerevisiae. Mutat. Res. 231, 177–186 (1990)
Mayer, V. W., Goin, C. J., Arras, C. A. & Taylor-Mayer, R. E. Comparison of chemically induced chromosome loss in a diploid, triploid, and tetraploid strain of Saccharomyces cerevisiae. Mutat. Res. 279, 41–48 (1992)
Fujiwara, T. et al. Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature 437, 1043–1047 (2005)
Nigg, E. A. Centrosome aberrations: cause or consequence of cancer progression? Nature Rev. Cancer 2, 815–825 (2002)
Lin, H. et al. Polyploids require Bik1 for kinetochore-microtubule attachment. J. Cell Biol. 155, 1173–1184 (2001)
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002)
Paques, F. & Haber, J. E. Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 63, 349–404 (1999)
Tong, A. H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001)
Huang, D. & Koshland, D. Chromosome integrity in Saccharomyces cerevisiae: the interplay of DNA replication initiation factors, elongation factors, and origins. Genes Dev. 17, 1741–1754 (2003)
Kadyk, L. C. & Hartwell, L. H. Sister chromatids are preferred over homologs as substrates for recombinational repair in Saccharomyces cerevisiae. Genetics 132, 387–402 (1992)
Klein, H. L. RDH54, a RAD54 homologue in Saccharomyces cerevisiae, is required for mitotic diploid-specific recombination and repair and for meiosis. Genetics 147, 1533–1543 (1997)
Whelan, W. L., Gocke, E. & Manney, T. R. The CAN1 locus of Saccharomyces cerevisiae: fine-structure analysis and forward mutation rates. Genetics 91, 35–51 (1979)
Krogh, B. O. & Symington, L. S. Recombination proteins in yeast. Annu. Rev. Genet. 38, 233–271 (2004)
Pan, X. et al. A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell 124, 1069–1081 (2006)
Melo, J. A., Cohen, J. & Toczyski, D. P. Two checkpoint complexes are independently recruited to sites of DNA damage in vivo. Genes Dev. 15, 2809–2821 (2001)
Lisby, M., Barlow, J. H., Burgess, R. C. & Rothstein, R. Choreography of the DNA damage response: spatiotemporal relationships among checkpoint and repair proteins. Cell 118, 699–713 (2004)
Sonoda, E. et al. Rad51-deficient vertebrate cells accumulate chromosomal breaks prior to cell death. EMBO J. 17, 598–608 (1998)
Michaelis, C., Ciosk, R. & Nasmyth, K. Cohesins: chromosomal proteins that prevent premature separation of sister chromatids. Cell 91, 35–45 (1997)
Mayer, M. L. et al. Identification of protein complexes required for efficient sister chromatid cohesion. Mol. Biol. Cell 15, 1736–1745 (2004)
Tanaka, T. U. et al. Evidence that the Ipl1-Sli15 (Aurora kinase-INCENP) complex promotes chromosome bi-orientation by altering kinetochore-spindle pole connections. Cell 108, 317–329 (2002)
Biggins, S. & Murray, A. W. The budding yeast protein kinase Ipl1/Aurora allows the absence of tension to activate the spindle checkpoint. Genes Dev. 15, 3118–3129 (2001)
Indjeian, V. B., Stern, B. M. & Murray, A. W. The centromeric protein Sgo1 is required to sense lack of tension on mitotic chromosomes. Science 307, 130–133 (2005)
Tanaka, T. U., Stark, M. J. & Tanaka, K. Kinetochore capture and bi-orientation on the mitotic spindle. Nature Rev. Mol. Cell Biol. 6, 929–942 (2005)
Dewar, H., Tanaka, K., Nasmyth, K. & Tanaka, T. U. Tension between two kinetochores suffices for their bi-orientation on the mitotic spindle. Nature 428, 93–97 (2004)
Cheeseman, I. M. et al. Implication of a novel multiprotein Dam1p complex in outer kinetochore function. J. Cell Biol. 155, 1137–1145 (2001)
Hsu, J. Y. et al. Mitotic phosphorylation of histone H3 is governed by Ipl1/aurora kinase and Glc7/PP1 phosphatase in budding yeast and nematodes. Cell 102, 279–291 (2000)
Yoder, T. J., Pearson, C. G., Bloom, K. & Davis, T. N. The Saccharomyces cerevisiae spindle pole body is a dynamic structure. Mol. Biol. Cell 14, 3494–3505 (2003)
Byers, B. & Goetsch, L. Duplication of spindle plaques and integration of the yeast cell cycle. Cold Spring Harb. Symp. Quant. Biol. 38, 123–131 (1974)
Tanaka, T., Cosma, M. P., Wirth, K. & Nasmyth, K. Identification of cohesin association sites at centromeres and along chromosome arms. Cell 98, 847–858 (1999)
Akhmanova, A. et al. The microtubule plus-end-tracking protein CLIP-170 associates with the spermatid manchette and is essential for spermatogenesis. Genes Dev. 19, 2501–2515 (2005)
Green, R. A., Wollman, R. & Kaplan, K. B. APC and EB1 function together in mitosis to regulate spindle dynamics and chromosome alignment. Mol. Biol. Cell 16, 4609–4622 (2005)
Raderschall, E. et al. Elevated levels of Rad51 recombination protein in tumor cells. Cancer Res. 62, 219–225 (2002)
Flygare, J. et al. Effects of HsRad51 overexpression on cell proliferation, cell cycle progression, and apoptosis. Exp. Cell Res. 268, 61–69 (2001)
Keen, N. & Taylor, S. Aurora-kinase inhibitors as anticancer agents. Nature Rev. Cancer 4, 927–936 (2004)
Dong, Z. & Fasullo, M. Multiple recombination pathways for sister chromatid exchange in Saccharomyces cerevisiae: role of RAD1 and the RAD52 epistasis group genes. Nucleic Acids Res. 31, 2576–2585 (2003)
Lea, D. E. & Coulson, C. A. The distribution of the numbers of mutants in bacterial populations. J. Genet. 49, 264–285 (1949)
Acknowledgements
We are grateful to many colleagues from the yeast community for providing the reagents. We thank A. Amon, G. Fink, J. Haber, R. Rothstein, M. McLaughlin, M. Raschle and members of the Pellman laboratory for discussions; G. Fink, S. Elledge, J. Haber, A. Van Oudenaarden, M. McLaughlin, R. Rothstein and J. Walter for comments on the manuscript; C. Glavin for Supplementary Fig. 9; and M. Lenburg for guidance on the analysis of the expression profile data. D.P. was supported by an NIH grant and a gift from the G. Harold and Leila Y. Mathers Foundation. The Boulder Laboratory for 3D Electron Microscopy of Cells is supported by an NIH grant to J. R. McIntosh.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The transcriptional profiling data are available at MIAMExpress database (http://www.ebi.ac.uk/miamexpress) under accession number E-MEXP-822. Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file provides detailed descriptions of the genome-wide strategy for tetraploid formation, the plasmid shuffle strategy for tetraploid formation, Nuf2-GFP fluorescence intensity measurements, high-voltage EM tomography of yeast and description and explanation of expression profiling analysis. This file also contains Supplementary Figures 1–9 and Supplementary Tables 1–3. (PDF 9627 kb)
Supplementary Data 1
Complete results of the genome-wide screen for genes specifically required in yeast tetraploid cells. (XLS 633 kb)
Supplementary Data 2
Complete results of the expression profiling. (XLS 1693 kb)
Supplementary Movie
3D reconstruction of a tetraploid forming spindle. (MOV 132380 kb)
Rights and permissions
About this article
Cite this article
Storchová, Z., Breneman, A., Cande, J. et al. Genome-wide genetic analysis of polyploidy in yeast. Nature 443, 541–547 (2006). https://doi.org/10.1038/nature05178
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature05178
This article is cited by
-
Genome doubling causes double trouble
Nature (2022)
-
Oncogenic BRAF induces whole-genome doubling through suppression of cytokinesis
Nature Communications (2022)
-
Sublinear scaling of the cellular proteome with ploidy
Nature Communications (2022)
-
Comparative regulomics supports pervasive selection on gene dosage following whole genome duplication
Genome Biology (2021)
-
Whole-genome doubling confers unique genetic vulnerabilities on tumour cells
Nature (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.