Abstract
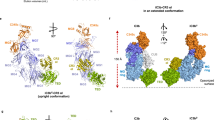
Resistance to infection and clearance of cell debris in mammals depend on the activation of the complement system, which is an important component of innate and adaptive immunity1,2. Central to the complement system is the activated form of C3, called C3b, which attaches covalently to target surfaces3 to amplify complement response, label cells for phagocytosis and stimulate the adaptive immune response. C3b consists of 1,560 amino-acid residues and has 12 domains. It binds various proteins and receptors to effect its functions4. However, it is not known how C3 changes its conformation into C3b and thereby exposes its many binding sites. Here we present the crystal structure at 4-Å resolution of the activated complement protein C3b and describe the conformational rearrangements of the 12 domains that take place upon proteolytic activation. In the activated form the thioester is fully exposed for covalent attachment to target surfaces and is more than 85 Å away from the buried site in native C3 (ref. 5). Marked domain rearrangements in the α-chain present an altered molecular surface, exposing hidden and cryptic sites that are consistent with known putative binding sites of factor B and several complement regulators. The structural data indicate that the large conformational changes in the proteolytic activation and regulation of C3 take place mainly in the first conversion step, from C3 to C3b. These insights are important for the development of strategies to treat immune disorders that involve complement-mediated inflammation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Walport, M. J. Complement. First of two parts. N. Engl. J. Med. 344, 1058–1066 (2001)
Carroll, M. C. The complement system in regulation of adaptive immunity. Nature Immunol. 5, 981–986 (2004)
Law, S. K. & Levine, R. P. Interaction between the third complement protein and cell surface macromolecules. Proc. Natl Acad. Sci. USA 74, 2701–2705 (1977)
Lambris, J. D. The multifunctional role of C3, the third component of complement. Immunol. Today 9, 387–393 (1988)
Janssen, B. J. et al. Structures of complement component C3 provide insights into the function and evolution of immunity. Nature 437, 505–511 (2005)
Mastellos, D. & Lambris, J. D. Complement: more than a ‘guard’ against invading pathogens?. Trends Immunol. 23, 485–491 (2002)
Tack, B. F., Harrison, R. A., Janatova, J., Thomas, M. L. & Prahl, J. W. Evidence for presence of an internal thiolester bond in third component of human complement. Proc. Natl Acad. Sci. USA 77, 5764–5768 (1980)
Fishelson, Z., Pangburn, M. K. & Muller-Eberhard, H. J. Characterization of the initial C3 convertase of the alternative pathway of human complement. J. Immunol. 132, 1430–1434 (1984)
Helmy, K. Y. et al. CRIg: a macrophage complement receptor required for phagocytosis of circulating pathogens. Cell 124, 915–927 (2006)
Dempsey, P. W., Allison, M. E., Akkaraju, S., Goodnow, C. C. & Fearon, D. T. C3d of complement as a molecular adjuvant: bridging innate and acquired immunity. Science 271, 348–350 (1996)
Ross, G. D., Lambris, J. D., Cain, J. A. & Newman, S. L. Generation of three different fragments of bound C3 with purified factor I or serum. I. Requirements for factor H vs CR1 cofactor activity. J. Immunol. 129, 2051–2060 (1982)
Harrison, R. A. & Lachmann, P. J. Novel cleavage products of the third component of human complement. Mol. Immunol. 17, 219–228 (1980)
Nagar, B., Jones, R. G., Diefenbach, R. J., Isenman, D. E. & Rini, J. M. X-ray crystal structure of C3d: a C3 fragment and ligand for complement receptor 2. Science 280, 1277–1281 (1998)
Brown, K. M. et al. Influence of donor C3 allotype on late renal-transplantation outcome. N. Engl. J. Med. 354, 2014–2023 (2006)
Dodds, A. W., Ren, X. D., Willis, A. C. & Law, S. K. The reaction mechanism of the internal thioester in the human complement component C4. Nature 379, 177–179 (1996)
Pangburn, M. K., Schreiber, R. D. & Muller-Eberhard, H. J. C3b deposition during activation of the alternative complement pathway and the effect of deposition on the activating surface. J. Immunol. 131, 1930–1935 (1983)
Sim, R. B., Twose, T. M., Paterson, D. S. & Sim, E. The covalent-binding reaction of complement component C3. Biochem. J. 193, 115–127 (1981)
Taniguchi-Sidle, A. & Isenman, D. E. Interactions of human complement component C3 with factor B and with complement receptors type 1 (CR1, CD35) and type 3 (CR3, CD11b/CD18) involve an acidic sequence at the N-terminus of C3 alpha′-chain. J. Immunol. 153, 5285–5302 (1994)
O′Keefe, M. C., Caporale, L. H. & Vogel, C. W. A novel cleavage product of human complement component C3 with structural and functional properties of cobra venom factor. J. Biol. Chem. 263, 12690–12697 (1988)
Kolln, J., Spillner, E., Andra, J., Klensang, K. & Bredehorst, R. Complement inactivation by recombinant human C3 derivatives. J. Immunol. 173, 5540–5545 (2004)
Inal, J. M. & Schifferli, J. A. Complement C2 receptor inhibitor trispanning and the beta-chain of C4 share a binding site for complement C2. J. Immunol. 168, 5213–5221 (2002)
Daoudaki, M. E., Becherer, J. D. & Lambris, J. D. A 34-amino acid peptide of the third component of complement mediates properdin binding. J. Immunol. 140, 1577–1580 (1988)
Oran, A. E. & Isenman, D. E. Identification of residues within the 727–767 segment of human complement component C3 important for its interaction with factor H and with complement receptor 1 (CR1, CD35). J. Biol. Chem. 274, 5120–5130 (1999)
Lambris, J. D., Avila, D., Becherer, J. D. & Muller-Eberhard, H. J. A discontinuous factor H binding site in the third component of complement as delineated by synthetic peptides. J. Biol. Chem. 263, 12147–12150 (1988)
Clemenza, L. & Isenman, D. E. Structure-guided identification of C3d residues essential for its binding to complement receptor 2 (CD21). J. Immunol. 165, 3839–3848 (2000)
Lambris, J. D., Dobson, N. J. & Ross, G. D. Release of endogenous C3b inactivator from lymphocytes in response to triggering membrane receptors for beta 1H globulin. J. Exp. Med. 152, 1625–1644 (1980)
The C. C. P. 4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994)
Storoni, L. C., McCoy, A. J. & Read, R. J. Likelihood-enhanced fast rotation functions. Acta Crystallogr. D Biol. Crystallogr. 60, 432–438 (2004)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004)
Nicholls, A., Bharadwaj, R. & Honig, B. GRASP: Graphical representation and analysis of surface properties. Biophys. J. 64, 166–170 (1993)
Acknowledgements
We acknowledge the European Synchrotron Radiation Facility for provision of synchrotron radiation facilities; and S.J. Kolodziej and J.K. Stoops (Houston) for providing the native α2-macroglobulin and methylamine-treated α2-macroglobulin EM-reconstruction maps. We thank B. Nilsson (Uppsala), R.B.G. Ravelli (Grenoble) and E.G. Huizinga (Utrecht) for reading the manuscript. This work was supported by a ‘Pioneer’ programme grant to P.G. by the Council for Chemical Sciences of the Netherlands Organization for Scientific Research (NWO-CW) and an NIH grant to J.D.L.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Coordinates and structure factors of the C3b structure have been deposited in the Protein Data Bank with accession number 2I07. Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Figure
Figure captions for Supplementary Tables 1 and 2, Supplementary Figures 1 to 3 and Supplementary Movie. (DOC 71 kb)
Supplementary Table 1
Diffraction data, refinement statistics and molecular replacement scores for C3b (PDF 28 kb)
Supplementary Table 2
Domain rotation and translation between C3b, C3 and C3c. (PDF 11 kb)
Supplementary Figure 1
Electron density for the individual domains of C3b. (PDF 1581 kb)
Supplementary Figure 2
Proteolytic cleavage sites of factor I mapped on the structure of C3b. (PDF 233 kb)
Supplementary Figure 3
Modeling of native and methylamine α2-macroglobulin based on 25-Å resolution EM reconstruction maps. (PDF 1578 kb)
Supplementary Movie
Conformational transition of C3 into C3b. (AVI 4920 kb)
Rights and permissions
About this article
Cite this article
Janssen, B., Christodoulidou, A., McCarthy, A. et al. Structure of C3b reveals conformational changes that underlie complement activity. Nature 444, 213–216 (2006). https://doi.org/10.1038/nature05172
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05172
This article is cited by
-
Current Understanding of Complement Proteins as Therapeutic Targets for the Treatment of Immunoglobulin A Nephropathy
Drugs (2023)
-
Insight into mode-of-action and structural determinants of the compstatin family of clinical complement inhibitors
Nature Communications (2022)
-
Invariant surface glycoprotein 65 of Trypanosoma brucei is a complement C3 receptor
Nature Communications (2022)
-
The crystal structure of iC3b-CR3 αI reveals a modular recognition of the main opsonin iC3b by the CR3 integrin receptor
Nature Communications (2022)
-
Cryo-EM structures of human A2ML1 elucidate the protease-inhibitory mechanism of the A2M family
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.