Abstract
Arising from: J. K. Taubenberger et al. Nature 437, 889–893 (2005); see also communication from Antonovics et al.; Taubenberger et al. reply
Taubenberger et al.1 have sequenced the polymerase genes of the pandemic ‘Spanish’ influenza A virus of 1918, thereby completing the decoding of the genome of this virus2,3,4,5,6. The authors conclude from these sequences that the virus jumped from birds to humans shortly before the start of the pandemic and that it was not derived from earlier viruses by gene shuffling, a process called reassortment. However, we believe that their evidence does not convincingly support these conclusions and that some of their results even indicate that, on the contrary, the virus evolved in mammals before the pandemic began and that it was a reassortant. In light of this alternative interpretation, we suggest that the current intense surveillance of influenza viruses should be broadened to include mammalian sources.
Similar content being viewed by others
Main
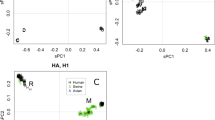
The inferences by Taubenberger et al.1 are based on two lines of evidence. First, they report phylogenetic trees for nine of the eleven genes of the virus. The trees would have supported the conclusions if, in all or at least in most of them, the 1918 virus linked the bird influenza viruses with the later human and swine influenza viruses: that is, if it had been placed on the root of the cluster of mammalian viruses (Fig. 1a). That topology would indicate that the 1918 virus was the ancestor of all post-1918 human influenza viruses, as Taubenberger et al.1 assume.
a, Phylogenetic tree that would be expected if the 1918 virus had come directly from birds. The avian influenza-virus source would be linked to the later influenzas isolated from mammals, namely on the branch linking the avian viruses to the mammalian viruses. b, Summary of the trees found by Taubenberger and co-workers1,2,3,4,5,6 for five of the influenza-virus genes (PA, PB1, HA, M1, M2; see ref. 1 for details), which placed the 1918 virus next to the later human influenzas: that is, they are sister groups. c, Summary of the remaining four influenza virus gene trees found by Taubenberger and co-workers1,2,3,4,5,6 (PB2, NP, NS1, NA; see ref. 1 for details), which placed the 1918 virus next to the later swine viruses. The avian-virus source cluster in most of the gene trees also includes virus isolates from mammals.
However, none of the nine phylogenetic trees described by Taubenberger and his co-workers, including those of the polymerases, shows that topology1,2,3,4,5,6. In five of the published trees, the 1918 gene is placed next to the main cluster of human influenza viruses, and the classical swine influenzas are linked to the tree between the branches of the human and avian viruses (Fig. 1b). The other four trees have the reverse topology; the 1918 gene lies next to the classical swine viruses, and the main cluster of human viruses is between them and the avian viruses (Fig. 1c). The combined mammalian cluster is linked to the bird cluster directly in five of the trees, but in the other four trees it is linked first to other lineages from mammals (that is, those of pigs and horses).
Taubenberger and colleagues1,2,3,4,5,6 report that the 1918 virus was placed with the bird influenza viruses in some unpublished trees and that in all the trees, both published and unpublished, the 1918 virus lies close to the root of the mammalian lineages of influenza viruses. However, the 1918 virus does not lie on the root, and therefore the phylogenetic analyses of Taubenberger et al. do not support their conclusions: their results indicate instead that the virus evolved in people or pigs for an unknown period of time before the pandemic started and that the virus may have been a reassortant.
The second line of evidence used by Taubenberger et al.1 to support their conclusions comes from comparing similarities in polymerase amino-acid sequences. Phylogenetic analysis is usually preferred to this approach for studying origins, because similarities can sometimes occur by coincidence through parallel evolution. Indeed, Taubenberger et al.1 discovered examples of identical amino-acid residues in bird and human influenza virus polymerases that resulted from parallel evolution, and we contend that those discoveries undermine their conclusions.
Although the 1918 virus polymerase proteins are placed closest to the typical sequences of some bird influenza viruses, the ranking depends on very few residues; parallel evolution could therefore have affected the ranking. In contrast, when nucleotide similarity was measured, rather than amino-acid similarity, the authors found the 1918 virus to be significantly closer to influenza viruses from mammals. The evidence of sequence similarity uncovered by Taubenberger et al.1 is important for understanding how influenza viruses adapt to humans, but it does not prove their conclusions.
The events that led to the emergence of the 1918 virus are unclear and will probably remain so, at least until the immediate ancestors of the virus have been characterized and the mutation rate of the virus lineage is known. Data from other influenzas show that the mutation rate varies from close to 0% to almost 1% per year7, introducing uncertainty about timing inferred from influenza-virus sequences. The haemagglutinin gene from an influenza virus from a goose captured in 1917 has already been sequenced8: perhaps unluckily, the 1917 virus was not an ancestor of the 1918 pandemic virus, but it was similar to contemporary H1N1 avian influenzas.
References
Taubenberger, J. K. et al. Nature 437, 889–893 (2005).
Basler, C. F. et al. Proc. Natl Acad. Sci. USA 98, 2746–2751 (2001).
Reid, A. H., Fanning, T. G., Hultin, J. V. & Taubenberger, J. K. Proc. Natl Acad. Sci. USA 96, 1651–1656 (1999).
Reid, A. H., Fanning, T. G., Janczewski, T. A. & Taubenberger, J. K. Proc. Natl Acad. Sci. USA 97, 6785–6790 (2000).
Reid, A. H., Fanning, T. G., Janczewski, T. A., McCall, S. & Taubenberger, J. K. J. Virol. 76, 10717–10723 (2002).
Reid, A. H., Fanning, T. G., Janczewski, T. A., Lourens, R. M. & Taubenberger, J. K. J. Virol. 78, 12462–12470 (2004).
Webster, R. G., Bean, W. G., Gorman, O. T., Chambers, T. M. & Kawaoka, Y. Microbiol. Rev. 56, 152–179 (1992).
Fanning, T. G. et al. J. Virol. 76, 7860–7862 (2002).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gibbs, M., Gibbs, A. Was the 1918 pandemic caused by a bird flu?. Nature 440, E8 (2006). https://doi.org/10.1038/nature04823
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04823
This article is cited by
-
Cross-species virus transmission and its pandemic potential
Bulletin of the National Research Centre (2022)
-
Long-term RNA persistence in postmortem contexts
Investigative Genetics (2013)
-
Detecting transmission and reassortment events for influenza A viruses with genotype profile method
Virology Journal (2011)
-
Identifying genetic markers of adaptation for surveillance of viral host jumps
Nature Reviews Microbiology (2010)
-
Using Non-Homogeneous Models of Nucleotide Substitution to Identify Host Shift Events: Application to the Origin of the 1918 ‘Spanish’ Influenza Pandemic Virus
Journal of Molecular Evolution (2009)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.