Abstract
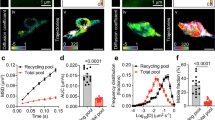
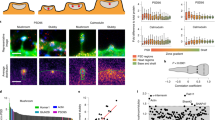
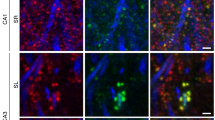
The regulated degradation of proteins by the ubiquitin proteasome pathway is emerging as an important modulator of synaptic function and plasticity1,2,3,4,5,6,7,8,9,10,11,12,13,14,15. The proteasome is a large, multi-subunit cellular machine that recognizes, unfolds and degrades target polyubiquitinated proteins. Here we report NMDA (N-methyl-d-aspartate) receptor-dependent redistribution of proteasomes from dendritic shafts to synaptic spines upon synaptic stimulation, providing a mechanism for local protein degradation. Using a proteasome-activity reporter and local perfusion, we show that synaptic stimulation regulates proteasome activity locally in the dendrites. We used restricted photobleaching of individual spines and dendritic shafts to reveal the dynamics that underlie proteasome sequestration, and show that activity modestly enhances the entry rate of proteasomes into spines while dramatically reducing their exit rate. Proteasome sequestration is persistent, reflecting an association with the actin-based cytoskeleton. Together, our data indicate that synaptic activity can promote the recruitment and sequestration of proteasomes to locally remodel the protein composition of synapses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hegde, A. N. et al. Ubiquitin C-terminal hydrolase is an immediate-early gene essential for long-term facilitation in Aplysia. Cell 89, 115–126 (1997)
Campbell, D. S. & Holt, C. E. Chemotropic responses of retinal growth cones mediated by rapid local protein synthesis and degradation. Neuron 32, 1013–1026 (2001)
Burbea, M., Dreier, L., Dittman, J. S., Grunwald, M. E. & Kaplan, J. M. Ubiquitin and AP180 regulate the abundance of GLR-1 glutamate receptors at postsynaptic elements in C. elegans. Neuron 35, 107–120 (2002)
Ehlers, M. D. Activity level controls postsynaptic composition and signaling via the ubiquitin-proteasome system. Nature Neurosci. 6, 231–242 (2003)
Zhao, Y., Hegde, A. N. & Martin, K. C. The ubiquitin proteasome system functions as an inhibitory constraint on synaptic strengthening. Curr. Biol. 13, 887–898 (2003)
Pak, D. T. & Sheng, M. Targeted protein degradation and synapse remodeling by an inducible protein kinase. Science 302, 1368–1373 (2003)
Patrick, G. N., Bingol, B., Weld, H. A. & Schuman, E. M. Ubiquitin-mediated proteasome activity is required for agonist-induced endocytosis of GluRs. Curr. Biol. 13, 2073–2081 (2003)
Colledge, M. et al. Ubiquitination regulates PSD-95 degradation and AMPA receptor surface expression. Neuron 40, 595–607 (2003)
Bingol, B. & Schuman, E. M. A proteasome-sensitive connection between PSD-95 and GluR1 endocytosis. Neuropharmacology 47, 755–763 (2004)
van Roessel, P., Elliott, D. A., Robinson, I. M., Prokop, A. & Brand, A. H. Independent regulation of synaptic size and activity by the anaphase-promoting complex. Cell 119, 707–718 (2004)
Juo, P. & Kaplan, J. M. The anaphase-promoting complex regulates the abundance of GLR-1 glutamate receptors in the ventral nerve cord of C. elegans. Curr. Biol. 14, 2057–2062 (2004)
Dreier, L., Burbea, M. & Kaplan, J. M. LIN-23-mediated degradation of β-catenin regulates the abundance of GLR-1 glutamate receptors in the ventral nerve cord of C. elegans. Neuron 46, 51–64 (2005)
Steward, O. & Schuman, E. M. Compartmentalized synthesis and degradation of proteins in neurons. Neuron 40, 347–359 (2003)
Bingol, B. & Schuman, E. M. Synaptic protein degradation by the ubiquitin proteasome system. Curr. Opin. Neurobiol. 15, 536–541 (2005)
Yi, J. J. & Ehlers, M. D. Ubiquitin and protein turnover in synapse function. Neuron 47, 629–632 (2005)
Enenkel, C., Lehmann, A. & Kloetzel, P. M. GFP-labelling of 26S proteasomes in living yeast: insight into proteasomal functions at the nuclear envelope/rough ER. Mol. Biol. Rep. 26, 131–135 (1999)
Isaac, J. T. Postsynaptic silent synapses: evidence and mechanisms. Neuropharmacology 45, 450–460 (2003)
Fujimuro, M., Sawada, H. & Yokosawa, H. Production and characterization of monoclonal antibodies specific to multi-ubiquitin chains of polyubiquitinated proteins. FEBS Lett. 349, 173–180 (1994)
Lindsten, K., Menendez-Benito, V., Masucci, M. G. & Dantuma, N. P. A transgenic mouse model of the ubiquitin/proteasome system. Nature Biotechnol. 21, 897–902 (2003)
Dantuma, N. P., Lindsten, K., Glas, R., Jellne, M. & Masucci, M. G. Short-lived green fluorescent proteins for quantifying ubiquitin/proteasome-dependent proteolysis in living cells. Nature Biotechnol. 18, 538–543 (2000)
Allison, D. W., Gelfand, V. I., Spector, I. & Craig, A. M. Role of actin in anchoring postsynaptic receptors in cultured hippocampal neurons: differential attachment of NMDA versus AMPA receptors. J. Neurosci. 18, 2423–2436 (1998)
Kurz-Isler, G. Induction of paracrystalline arrays by vincristine in the synaptic formations of the teleost retina. Cell Tissue Res. 191, 75–82 (1978)
Setou, M. et al. Glutamate-receptor-interacting protein GRIP1 directly steers kinesin to dendrites. Nature 417, 83–87 (2002)
Fang, S. & Weissman, A. M. A field guide to ubiquitylation. Cell. Mol. Life Sci. 61, 1546–1561 (2004)
Miller, J. & Gordon, C. The regulation of proteasome degradation by multi-ubiquitin chain binding proteins. FEBS Lett. 579, 3224–3230 (2005)
Glickman, M. H. & Raveh, D. Proteasome plasticity. FEBS Lett. 579, 3214–3223 (2005)
Gordon, C. The intracellular localization of the proteasome. Curr. Top. Microbiol. Immunol. 268, 175–184 (2002)
Ferrell, K., Wilkinson, C. R., Dubiel, W. & Gordon, C. Regulatory subunit interactions of the 26S proteasome, a complex problem. Trends Biochem. Sci. 25, 83–88 (2000)
Ostroff, L. E., Fiala, J. C., Allwardt, B. & Harris, K. M. Polyribosomes redistribute from dendritic shafts into spines with enlarged synapses during LTP in developing rat hippocampal slices. Neuron 35, 535–545 (2002)
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nature Biotechnol. 20, 87–90 (2002)
Acknowledgements
We thank the Kloetzel, Masucci and Kennedy laboratories for providing the Rpt1(CIM5)–GFP, UbG76V–GFP and mRFP clones, respectively. We also thank members of the Schuman laboratory, especially C.-Y. Tai and S. Kim, and former member G. Patrick for discussions. E.M.S. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains Supplementary Figures 1–6 and Supplementary Legends 1–6, Supplementary Methods, Supplementary Notes (additional references pertaining to supplementary methods) and Supplementary Table 1. (PDF 7302 kb)
Rights and permissions
About this article
Cite this article
Bingol, B., Schuman, E. Activity-dependent dynamics and sequestration of proteasomes in dendritic spines. Nature 441, 1144–1148 (2006). https://doi.org/10.1038/nature04769
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04769
This article is cited by
-
Post-ischemic ubiquitination at the postsynaptic density reversibly influences the activity of ischemia-relevant kinases
Communications Biology (2024)
-
Interactions Between the Ubiquitin–Proteasome System, Nrf2, and the Cannabinoidome as Protective Strategies to Combat Neurodegeneration: Review on Experimental Evidence
Neurotoxicity Research (2024)
-
Synaptic proteasome is inhibited in Alzheimer’s disease models and associates with memory impairment in mice
Communications Biology (2023)
-
RNA transport and local translation in neurodevelopmental and neurodegenerative disease
Nature Neuroscience (2021)
-
Traceable stimulus-dependent rapid molecular changes in dendritic spines in the brain
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.