Abstract
After the completion of a draft human genome sequence1, the International Human Genome Sequencing Consortium has proceeded to finish2 and annotate each of the 24 chromosomes comprising the human genome. Here we describe the sequencing and analysis of human chromosome 3, one of the largest human chromosomes. Chromosome 3 comprises just four contigs, one of which currently represents the longest unbroken stretch of finished DNA sequence known so far. The chromosome is remarkable in having the lowest rate of segmental duplication in the genome. It also includes a chemokine receptor gene cluster as well as numerous loci involved in multiple human cancers such as the gene encoding FHIT, which contains the most common constitutive fragile site in the genome, FRA3B3. Using genomic sequence from chimpanzee and rhesus macaque, we were able to characterize the breakpoints defining a large pericentric inversion that occurred some time after the split of Homininae from Ponginae, and propose an evolutionary history of the inversion.
Similar content being viewed by others
Main
The physical map of chromosome 3 was generated using a combination of STS-derived probe screening of bacterial artificial chromosome (BAC) clone libraries and the fingerprint map4 and then used to pick concomitantly a tiling path of 1,710 overlapping BAC and P1-derived artificial chromosome (PAC) clones for sequencing. The two remaining euchromatic gaps have proved recalcitrant to screening BAC libraries (http://bacpac.chori.org/) consisting of better than an 80-fold representation of the human genome. Gap sizes were estimated by a combination of fibre-fluorescence in situ hybridization (FISH; C. Wagner-McPherson, personal communication) and homologous gap flank mapping to the chimpanzee and/or rhesus macaque assembly, and total an estimated 137 kilobases (kb). A more thorough cross-species analysis of gap size can be found in Supplementary Table 1. The data extend to within 35 kb of the (TTAGGG)n telomeric repeat motif on the p-arm and 55 kb on the q-arm of chromosome 3 (H. Riethman, personal communication; see also http://www.wistar.upenn.edu/Riethman/). The p-arm pericentromeric sequence contains 147.5 kb of monomeric alpha-satellite repeats, whereas the q-arm sequence extends 8.6 kb into these repeats. The chromosome is characterized by a highly polymorphic heterochromatin block at 3q11.2—similar to, but far shorter than, those present on chromosomes 1, 9, 16 and Y—that ranges in size from 0.2 to 2.0 megabases (Mb)5 and is thought to consist primarily of satellite 1 repeat sequence6. We have assumed a 1.5-Mb block and a core centromere size of 2.9 Mb to arrive at an overall chromosome length of 199,344,050 base pairs (bp).
The sequence was generated using a clone-by-clone random shotgun sequencing and finishing strategy2 (see Methods). Each tiling path BAC clone was finished to community standards (http://genomeold.wustl.edu/Overview/g16stand.php). We finished 194,944,050 bp of euchromatic sequence to an independently measured accuracy of at least 99.99%7 and have covered more than 99.99% of the euchromatic chromosome. Each of the landscape features and annotations outlined here may be viewed as user-specified tracks on the Genboree Browser (http://www.genboree.org/Hs.chr3).
In the current assembly of the genome (NCBI build 35) all RefSeq8 genes are entirely accounted for with partial sequence available for SLC25A26 (NM_173471 and splice variants; bases encoding the first exon are now accounted for in GenBank, accession AC170165) and RYBP (NM_012234; bases 1–218 may be polymorphic in the population). As can be seen in Supplementary Fig. 1, there is strong concordance in marker order and content between the finished sequence and various genetic maps (see Supplementary Methods). We analysed the recombination rate across the chromosome using the deCODE9 markers and found the statistics to be in line with the other human chromosomes, yielding a sex-average rate of 1.14 cM Mb-1. The female and male recombination rates were found to be 1.43 cM Mb-1 and 0.85 cM Mb-1 respectively, with maximum rates of 3.77 cM Mb-1 in females and 5.77 cM Mb-1 in males (Supplementary Fig. 2). Although there are no recombination deserts as previously defined10, there is a recombination jungle at the tip of the p-arm (3p26.3-26.1).
As the beginning of what is inherently a dynamic process, we used manual curation of the automated Ensembl annotation output of NCBI Human Assembly build 33 to characterize fully the gene content of chromosome 3. Using all publicly available human protein, complementary DNA and spliced expressed sequence tag (EST) databases together with selected gene prediction algorithms and UCSC cDNA resources, we characterized each locus using the standards established by the Human Annotation Working Group (http://www.sanger.ac.uk/HGP/havana/hawk.shtml). Starting with 1,249 loci and 1,697 variants, we annotated 1,585 gene loci (Fig. 1). Among these were 1,425 known coding genes, 8 novel genes, 27 novel transcripts, 3 putative genes and 122 pseudogenes. We found 4,857 paralogous gene pairs, just 361 of which were intrachromosomal—a further reflection of the low segmental duplication (greater than 90% similarity over at least 1 kb) rate on this chromosome. However, this paralogous set reflects a number of ancient duplications, including one that contains genes encoding the developmentally important nuclear receptors (see Supplementary Fig. 3). Excluding the pseudogenes, the average gene density is 8.8 genes per Mb, making it one of the more gene-poor chromosomes. However, although the average gene density is low, as with other gene-poor chromosomes such as chromosome 13, the genes are larger than the genome average and cover 98.3 Mb or 49% of the chromosome. Chromosome 3 contains two gene-dense clusters on the p-arm between base coordinates 10–17 Mb and 41–55 Mb (18.9 and 21.1 genes per Mb, respectively). These two regions alone account for 26% of the genes on the chromosome. Relatively gene-poor tracts are confined to the pericentromeric regions.
Tracks are numbered on the left and syntenic alignments across the human chromosome are shown in the top five tracks: (1) human–chicken; (2) human–rat; (3) human–mouse; (4) human–dog; and (5) human–chimpanzee. The inter- and intrachromosomal breakpoints are represented by red and blue gaps, respectively. Cyan gaps indicate regions without sequence alignment and the centromere is located in the cyan gap that is common to all species. Purple brackets indicate sequence inversions. The density of recent segmental intra- and interchromosomal duplications from low-copy repeats is shown in tracks (6) and (7). The incidence of major interspersed (high-copy) repeats are depicted in tracks (8), (9) and (10) for LINEs, LTRs and SINEs, respectively. The variations in G + C content, and densities of CpG islands, genes and pseudogenes appear in tracks (11), (12), (13) and (14), respectively, whereas gene paralogue density, gene density and gene variant density appear in tracks (15), (16) and (17), respectively. Gene density in track 13 is from UCSC ‘known genes’, whereas track 16 reflects the non-redundant locus annotations detailed in this study.
Approximately 57% of chromosome 3 genes expressed alternative transcripts with an average of 2.86 transcripts per gene. The IFRD2 gene had the highest number of alternative transcripts at 21 annotated variants. Most of these could produce altered protein products (3,163 different proteins from among 4,096 alternative transcripts). There were at least 681 partial transcripts in the database for which we could not identify the complete coding sequence.
We analysed the chromosome for the presence of distinguishing features including CpG islands, G + C content, segmental duplications, repeat content (see Fig. 1) and non-coding RNA. Of the 1,575 genetic loci analysed (including their variants), 56–57% were associated with a CpG island. The G + C content was found to correlate well with gene density, as expected, and repeat content is unremarkable. Chromosome 3 is relatively devoid of segmental duplications, having just 1.7% of its bases composed of duplicated sequence compared to the whole-genome average of 5.3%. This is the lowest percentage for any chromosome in the genome.
We analysed the known and predicted non-coding RNA gene content on chromosome 3 as a prelude to future annotation of regulatory regions. Using three different strategies (see Methods), we were able to find 703 putative, non-redundant non-coding RNAs (ncRNAs). The most abundant ncRNA candidates found (68%) were mRNA-like ncRNAs, whereas the remainder consisted of smaller ncRNAs of various types including small nuclear RNAs, Y RNAs, small nucleolar RNAs, microRNAs, SRP RNAs, a telomerase RNA, 7SK RNAs, small Cajal body-specific RNAs (scaRNAs), a small non-messenger RNA (snmRNA) and a small group of ribosomal RNAs and transfer RNAs (see Supplementary Tables 2 and 3). Further characterization of the genomic landscape from 3pter to D3S3397 is also available11.
Cytogenetic studies using chromosome painting and comparative mapping analysis suggest that a fission event in the largest ancestral eutherian chromosome gave rise to human chromosomes 3 and 21 (ref. 12). These observations were extended by a study13 using gene or genome sequence anchors and the chicken genome sequence as an out-group to reconstruct the ancestral mammalian genome. These analyses paint a more complicated evolutionary picture requiring six or seven recombination events to account for human chromosome 3. The pattern gets more complicated in comparison to the rodent genomes due to their well-characterized higher rates of interchromosomal rearrangement. Nevertheless, consistent syntenic blocks are observed in both mouse and rat, particularly at each end of the chromosome and along most of the q-arm (see Fig. 1).
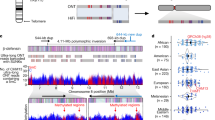
Further comparative FISH analysis revealed that a large-scale pericentric inversion occurred in the ancestor of the African apes and is present in modern human chromosome 3 as well as the chimpanzee and gorilla orthologues, but not in orang-utan or Old World monkeys14. Two scaffolds from the Baylor College of Medicine Human Genome Sequencing Center (BCM-HGSC) rhesus macaque Mmul_0.1 assembly were found to span both breakpoints of the human inversion (Fig. 2; see Supplementary Table 4 for breakpoint details). The macaque 5′ breakpoint is characterized by a short homologous GTGG track (Fig. 2b) and by a mammalian interspersed repeat (MIR) that was split by a segmental duplication before the inversion resulting in one part, designated MIR A, present in boundary I and a second part, designated MIR B, present in boundary III (see Fig. 2a). The MIR at the 3′ end of boundary III was present in the segmental duplication and may have been involved in the insertion event. A number of simple repeats and low complexity regions were found within 1 kb of the breakpoint (see Supplementary Table 5). Each of these elements, including retrotransposons15, short homologous sequence and alternating purine-pyrimidine tracks16 have been reported for many other breakpoints.
a, Inversion breakpoint regions in human chromosome 3 compared to chimpanzee and rhesus macaque. Insets of breakpoint boundaries are designated by Roman numerals. Figure adapted from the UCSC genome browser and ref. 16. b, Rhesus breakpoint sequence aligned to human. The sequences homologous to both breakpoints are underlined.
The inversion breakpoint regions on human chromosome 3 are characterized by segmental duplications. Both breakpoints contain segmental duplications that are at least partially repeated at numerous intra- and interchromosomal locations on chromosomes 3, 4, 7, 8, 11, 12 and 16. The entire 5′ segmental duplication maps to a location on 11q13, suggesting its probable origin, and the 3′ segmental duplication maps to an adjacent block of 11q13. The 5′ and 3′ segmental duplications do not align to one another. These results suggest that a single segmental duplication occurred at the 5′ breakpoint followed by the inversion break within the segmental duplication, splitting a long terminal repeat from the human endogenous retrovirus K family (LTR5B) into two parts. The two parts are designated LTR5B A and LTR5B B in Fig. 2a, and the dot plots in Supplementary Fig. 4 can be placed together to form a complete LTR5B element. The 5′ and 3′ segmental duplications are aligned to the same chromosome 11 region in this figure, showing their adjacency and also that the duplication inserted in the reverse orientation on chromosome 3 compared with chromosome 11.
It is unclear whether segmental duplications are the cause or result of rearrangements15. The segmental duplication is not present in macaque, and the MIR element spanning the 5′ breakpoint in macaque seems to have been split by the segmental duplication before the inversion. LTR5B elements are found in human, chimpanzee and gorilla but not orang-utan17, suggesting that the duplication occurred after the African ape–orang-utan divergence. Indeed, the 3′ duplication is not present in orang-utan or gibbon based on comparative FISH studies18. The LTR5B element was present in the segmental duplication, so the splitting of LTR5B by the inversion most probably occurred after the duplication.
Regions of segmental duplications involved in evolutionary rearrangements can also be involved in rearrangements associated with human disease19,20. The q-arm pericentromeric breakpoint undergoes t(3:11)(q21:q13) translocations in head and neck squamous cell carcinomas21 and acute myeloid leukaemia22. Perhaps the most interesting—because it involves the same regions as the evolutionary inversion—are inv(3)(p25:q21) pericentric inversions, along with other accompanying chromosomal abnormalities, which cause severe developmental abnormalities23,24.
At least 505 disease loci have been mapped to chromosome 3 (see http://www.ncbi.nlm.nih.gov/Omim/mimstats.html and Supplementary Tables 6 and 7). These include simple repeat expansions such as spinocerebellar ataxia 7 (ATXN7) involving an expanded CAG repeat (38–150 copies in the mutant allele compared with 7–17 copies in the normal allele) and myotonic dystrophy 2 (DM2), caused by the expansion of a CCTG repeat in the zinc finger gene ZNF9. Genes involved in DNA repair are encoded on chromosome 3, including XPC (xeroderma pigmentosum complementation group C), MLH1, a gene involved in DNA mismatch repair and mutated in hereditary non-polyposis coli, and Fanconi anaemia complementation group D2 (FANCD2), mapped to 3p26.
Among the most interesting medically relevant regions on the chromosome is the cluster of chemokine receptor genes mapping to 3p21. Within this group, the gene encoding CCR5 has been shown to be a critical cofactor for HIV-1 virus entry into cells, as defective alleles have been associated with HIV infection resistance. The clustering of both the chemokine and chemokine receptor genes suggests a relatively recent and rapid evolution of both gene families by local duplications.
Finally, a large number of cancer lesions have been mapped to chromosome 3 and cancer breakpoints seem to correlate with the four known fragile sites on the chromosome (see Supplementary Fig. 5). The cancer loci include the VHL gene, mutated in von Hippel–Lindau syndrome and linked to kidney cancer susceptibility, β-catenin, mutated in a number of colon tumours, and mutations in the FHIT gene, which encompasses the most common fragile site in the human genome (FRA3B) and for which aberrant transcripts have been found in about half of all oesophageal, stomach and colon carcinomas. The complete chromosome 3 sequence presented here provides a rich resource for future studies aimed at understanding our evolutionary history and the molecular basis of human variation and disease.
Methods
Mapping and sequencing
BAC clone screening, sequencing and finishing strategies are described in Supplementary Methods. Sequence overlaps between BAC clones were verified by BLAST (http://www.ncbi.nlm.nih.gov/BLAST/) and polymorphic regions within overlaps were confirmed by polymerase chain reaction using a bi-gender, multi-ethnic pool of genomic DNA isolated from eight individuals (J. Belmont, personal communication). The quality of the assembly was assessed using restriction digests of individual BAC clones together with genetic and radiation hybrid map marker content and order (see Supplementary Methods and Supplementary Fig. 1), gene content and large-insert paired ends. Unique fosmid end sequences (Broad Institute) were downloaded from the UCSC Genome Browser, aligned to the genomic sequence and checked for both pair orientation and resulting insert size.
Annotation
We manually curated each known, novel CDS and novel transcript locus, defined as a set of one or more transcripts that share at least one exon of in-frame coding sequence and supported by full-length and partial human cDNAs or vertebrate cDNAs having a best in genome BLAST/Blat (http://genome.ucsc.edu/cgi-bin/hgBlat) hit with >98% identity. The cDNAs/reference sequences (RefSeqs)8 were compared to the genomic sequence to place exons, and all splice sites were examined for canonical sequence. Coding regions were examined for a best-fit open reading frame. The 5′ and 3′ untranslated regions were annotated and extended using available EST and cDNA evidence; poly-A sites and poly-A signals were annotated on each gene where identified. Alternative splice variants were identified from cDNA, EST and protein evidence and the translation product for each CDS was verified using SwissProt. Pseudogenes were defined as sequences with no direct evidence for expression while having a match with high score to a spliced mRNA or spliced EST from elsewhere in the genome. This is a more stringent definition than has been applied by others in broad genomic screens of pseudogenes and results in a fivefold lower count across chromosome 3 than previously reported25. For paralogue analysis, protein sequences corresponding to the ‘KnownGenes’ track of the UCSC Browser were compared in an all-against-all BLAST search. Two loci were defined as paralogues if there was a match of any of their transcript variants with the following criteria: expect value cutoff of 10-10 or less, the lengths of the matching transcripts are within 20% of each other, and the match length extends over 70% of the average length of the two sequences. The complete set of annotations has been submitted to the Vega database (http://vega.sanger.ac.uk/Homo_Sapiens/).
Landscape features
CpG islands were defined as an expanse of greater than 200 nucleotides in which the G + C content is >50% and the ratio of the observed CG dinucleotides to expected in the segment is >0.6. We scanned the chromosome for ncRNAs as detailed in Supplementary Methods. We identified recent intra- and interchromosomal segmental duplications by using BLAST to align the repeat-masked chromosome sequence against itself and the rest of the human genome. The duplication densities were calculated by averaging the duplications of each base over non-overlapping 100-kb windows after filtering low-identity matches (< 90%). The densities of short interspersed elements (SINEs), long interspersed elements (LINEs) and long terminal repeats (LTRs) were calculated from repeat-masked data using 100-kb windows. The G + C density was calculated by counting the G + C content over non-overlapping 100-kb windows. The densities of CpG islands, genes (BCM-HGSC annotations) and pseudogenes were counted and displayed using 1-Mb windows.
Comparative analysis
The multiple alignments of human, chimpanzee (panTro1), dog (canFam1), mouse (mm5), rat (rn3), chicken (galGal2), zebrafish (danRer1) and Fugu (fr1) were downloaded from the UCSC Genome Browser (http://hgdownload.cse.ucsc.edu/) (see Supplementary Methods). The pairwise synteny blocks between human and other species were parsed with Synteny-Parser (X. Song and G. Weinstock, unpublished perl script), which was tuned to include all visible chromosome rearrangements in the dot plot. Rhesus scaffolds from the Mmul_0.1 preliminary assembly were mapped to human chromosome 3 using Pash26. Rhesus scaffolds mapped by both Pash and human–rhesus net alignments (UCSC) were aligned with orthologous human regions and chimpanzee regions from the human–chimpanzee reciprocal best chain alignments (UCSC) using MLAGAN27.
References
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004)
Markkanen, A., Heinonen, K., Knuutila, S. & de la Chapelle, A. Methotrexate-induced increase in gap formation in human chromosome band 3p14. Hereditas 96, 317–319 (1982)
McPherson, J. D. et al. A physical map of the human genome. Nature 409, 934–941 (2001)
Kalitsis, P., Earle, E., Vissel, B., Shaffer, L. G. & Choo, K. H. A chromosome 13-specific human satellite I DNA subfamily with minor presence on chromosome 21: further studies on Robertsonian translocations. Genomics 16, 104–112 (1993)
Tagarro, I., Fernandez-Peralta, A. M. & Gonzalez-Aguilera, J. J. Digestion of centromeric DNA from each human metaphase chromosome by the 6 bp-restriction enzyme StuI. Histochemistry 99, 453–456 (1993)
Schmutz, J. et al. Quality assessment of the human genome sequence. Nature 429, 365–368 (2004)
Pruitt, K. D., Tatusova, T. & Maglott, D. R. NCBI Reference Sequence (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 33, D501–D504 (2005)
Kong, A. et al. A high-resolution recombination map of the human genome. Nature Genet. 31, 241–247 (2002)
Yu, A. et al. Comparison of human genetic and sequence-based physical maps. Nature 409, 951–953 (2001)
The Chinese Human Genome Sequencing Consortium. “Beijing Region” (3pter-D3S3397) of the human genome: complete sequence and analysis. Sci. China C Life Sci. 48, 311–329 (2005)
Wienberg, J. The evolution of eutherian chromosomes. Curr. Opin. Genet. Dev. 14, 657–666 (2004)
Bourque, G., Zdobnov, E. M., Bork, P., Pevzner, P. A. & Tesler, G. Comparative architectures of mammalian and chicken genomes reveal highly variable rates of genomic rearrangements across different lineages. Genome Res. 15, 98–110 (2005)
Ventura, M. et al. Recurrent sites for new centromere seeding. Genome Res. 14, 1696–1703 (2004)
Eichler, E. E. & Sankoff, D. Structural dynamics of eukaryotic chromosome evolution. Science 301, 793–797 (2003)
Bacolla, A. et al. Breakpoints of gross deletions coincide with non-B DNA conformations. Proc. Natl Acad. Sci. USA 101, 14162–14167 (2004)
Hughes, J. F. & Coffin, J. M. Evidence for genomic rearrangements mediated by human endogenous retroviruses during primate evolution. Nature Genet. 29, 487–489 (2001)
Yue, Y., Grossmann, B., Ferguson-Smith, M., Yang, F. & Haaf, T. Comparative cytogenetics of human chromosome 3q21.3 reveals a hot spot for ectopic recombination in hominoid evolution. Genomics 85, 36–47 (2005)
Eichler, E. E. Recent duplication, domain accretion and the dynamic mutation of the human genome. Trends Genet. 17, 661–669 (2001)
Stankiewicz, P. & Lupski, J. R. Genome architecture, rearrangements and genomic disorders. Trends Genet. 18, 74–82 (2002)
Jin, Y., Jin, C., Wennerberg, J., Hoglund, M. & Mertens, F. Cyclin D1 amplification in chromosomal band 11q13 is associated with overrepresentation of 3q21-q29 in head and neck carcinomas. Int. J. Cancer 98, 475–479 (2002)
Cigudosa, J. C. et al. A recurrent translocation, t(3;11)(q21;q13), found in two distinct cases of acute myeloid leukemia. Cancer Genet. Cytogenet. 83, 119–120 (1995)
Allderdice, P. W., Browne, N. & Murphy, D. P. Chromosome 3 duplication q21 leads to qter deletion p25 leads to pter syndrome in children of carriers of a pericentric inversion inv(3) (p25q21). Am. J. Hum. Genet. 27, 699–718 (1975)
Stine, S. B., Clark, C. E., Telfer, M. A., Casey, P. A. & Cowell, H. R. Ullrich-Turner syndrome (45,X/46,X,i[Xq]) in a child with a familial inversion of chromosome 3. Am. J. Med. Genet. 12, 57–62 (1982)
Zhang, Z., Harrison, P. M., Liu, Y. & Gerstein, M. Millions of years of evolution preserved: a comprehensive catalog of the processed pseudogenes in the human genome. Genome Res. 13, 2541–2558 (2003)
Kalafus, K. J., Jackson, A. R. & Milosavljevic, A. Pash: efficient genome-scale sequence anchoring by Positional Hashing. Genome Res. 14, 672–678 (2004)
Brudno, M. et al. LAGAN and Multi-LAGAN: efficient tools for large-scale multiple alignment of genomic DNA. Genome Res. 13, 721–731 (2003)
Acknowledgements
We wish to acknowledge T. Taylor and the rest of the Riken Genome Center for substitute BAC clones added to the tiling path. The authors acknowledge and thank the genome sequencing community for generating the data sets used in our comparative analysis. We also acknowledge the following authors from the HUGO Gene Nomenclature Committee: S. Povey (chair), E. A. Bruford, V. K. Khodiyar, R. C. Lovering, M. J. Lush, K. M. B. Sneddon, T. P. Sneddon and C. C. Talbot Jr. This work was supported by NIH grants to M. Olson and R. Gibbs. The Chinese Human Genome Sequencing Consortium is sponsored by the Ministry of Science and Technology, Chinese Academy of Sciences, National Natural Science Foundation of China, Beijing Municipal Government, Zhejiang Provincial Government, Hangzhou Municipal Government and Yueqing Municipal Government. Funding was also supplied by the Federal German Ministry of Education and Research and The Max Planck Society.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The chromosome 3 sequence has been deposited in GenBank under accession number NC_000003. Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1–5. (PDF 997 kb)
Supplementary Tables
This file contains Supplementary Tables 1–7. (PDF 382 kb)
Supplementary Methods
This file contains core methodologies used to sequence BAC clones, assess BAC clone coverage, order and integrity together with identification of ncRNA and comparative sequence analysis sources. (PDF 86 kb)
Supplementary Notes
This file contains a list of further authors contributing to this manuscript. (DOC 36 kb)
Rights and permissions
About this article
Cite this article
Muzny, D., Scherer, S., Kaul, R. et al. The DNA sequence, annotation and analysis of human chromosome 3. Nature 440, 1194–1198 (2006). https://doi.org/10.1038/nature04728
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04728
This article is cited by
-
Odontogene Keratozyste – Ätiologie, Pathogenese und Therapie
wissen kompakt (2019)
-
The international Human Genome Project (HGP) and China’s contribution
Protein & Cell (2017)
-
Novel Biomarkers for Non-functioning Invasive Pituitary Adenomas were Identified by Using Analysis of microRNAs Expression Profile
Biochemical Genetics (2017)
-
ARMc8 indicates aggressive colon cancers and promotes invasiveness and migration of colon cancer cells
Tumor Biology (2015)
-
Cloning and expression of the two new variants of Nav1.5/SCN5A in rat brain
Molecular and Cellular Biochemistry (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.