Abstract
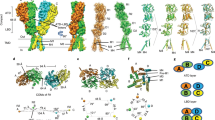
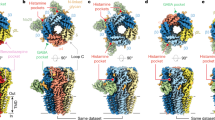
At synapses throughout the brain and spinal cord, the amino-acid glutamate is the major excitatory neurotransmitter. During evolution, a family of glutamate-receptor ion channels seems to have been assembled from a kit consisting of discrete ligand-binding, ion-channel, modulatory and cytoplasmic domains. Crystallographic studies that exploit this unique architecture have greatly aided structural analysis of the ligand-binding core, but the results also pose a formidable challenge, namely that of resolving the allosteric mechanisms by which individual domains communicate and function in an intact receptor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Watkins, J. C. & Evans, R. H. Excitatory amino acid transmitters. Annu. Rev. Pharmacol. Toxicol. 21, 165–204 (1981).
Mayer, M. L. & Westbrook, G. L. The physiology of excitatory amino acids in the vertebrate central nervous system. Prog. Neurobiol. 28, 197–276 (1987).
Collingridge, G. L. & Lester, R. A. J. Excitatory amino acid receptors in the vertebrate central nervous system. Pharmacol. Rev. 41, 143–210 (1989).
Collingridge, G. L. & Bliss, T. V. Memories of NMDA receptors and LTP. Trends Neurosci. 18, 54–56 (1995).
Geiger, J. R. P. et al. Relative abundance of subunit mRNAs determines gating and Ca2+ permeability of AMPA receptors in principle neurons and interneurons in rat CNS. Neuron 15, 193–204 (1995).
Lerma, J. Roles and rules of kainate receptors in synaptic transmission. Nature Rev. Neurosci. 4, 481–495 (2003).
Perl, T. M. et al. An outbreak of toxic encephalopathy caused by eating mussels contaminated with domoic acid. N. Engl. J. Med. 322, 1775–1780 (1990).
Watkins, J. C. & Jane, D. E. The glutamate story. Br. J. Pharmacol. 147 Suppl 1, S100–S108 (2006).
Hollmann, M. in Ionotropic Glutamate Receptors in the CNS (eds Jonas, P. & Monyer, H.) 3–98 (Springer, Berlin, 1999).
Sommer, B. et al. Flip and flop: a cell-specific functional switch in glutamate-operated channels of the CNS. Science 249, 1580–1585 (1990).
Sommer, B., Köhler, M., Sprengel, R. & Seeburg, P. H. RNA editing in brain controls a determinant of ion flow in glutamate-gated channels. Cell 67, 11–19 (1991).
Puchalski, R. B. et al. Selective RNA editing and subunit assembly of native glutamate receptors. Neuron 13, 131–147 (1994).
Ayalon, G. & Stern-Bach, Y. Functional assembly of AMPA and kainate receptors is mediated by several discrete protein-protein interactions. Neuron 31, 103–113 (2001).
Ayalon, G., Segev, E., Elgavish, S. & Stern-Bach, Y. Two regions in the N-terminal domain of ionotropic glutamate receptor 3 form the subunit oligomerization interfaces that control subtype-specific receptor assembly. J. Biol. Chem. 280, 15053–15060 (2005).
Paoletti, P., Ascher, P. & Neyton, J. High-affinity zinc inhibition of NMDA NR1-NR2A receptors. J. Neurosci. 17, 5711–5725 (1997).
Perin-Dureau, F., Rachline, J., Neyton, J. & Paoletti, P. Mapping the binding site of the neuroprotectant ifenprodil on NMDA receptors. J. Neurosci. 22, 5955–5965 (2002).
Masuko, T. et al. A regulatory domain (R1-R2) in the amino terminus of the N-methyl-D-aspartate receptor: effects of spermine, protons, and ifenprodil, and structural similarity to bacterial leucine/isoleucine/valine binding protein. Mol. Pharmacol. 55, 957–969 (1999).
Paoletti, P. et al. Molecular organization of a zinc binding N-terminal modulatory domain in a NMDA receptor subunit. Neuron 28, 911–925 (2000).
Pasternack, A. et al. Alpha-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptor channels lacking the N-terminal domain. J. Biol. Chem. 277, 49662–49667 (2002).
Stern-Bach, Y. et al. Agonist-selectivity of glutamate receptors is specified by two domains structurally related to bacterial amino acid binding proteins. Neuron 13, 1345–1357 (1994).
Kuusinen, A., Arvola, M. & Keinanen, K. Molecular dissection of the agonist binding site of an AMPA receptor. EMBO J. 14, 6327–6332 (1995).
Chen, G. Q., Sun, Y., Jin, R. & Gouaux, E. Probing the ligand binding domain of the GluR2 receptor by proteolysis and deletion mutagenesis defines domain boundaries and yields a crystallizable construct. Protein Sci. 7, 2623–2630 (1998).
Panchenko, V. A., Glasser, C. R. & Mayer, M. L. Structural similarities between glutamate receptor channels and K+ channels examined by scanning mutagenesis. J. Gen. Physiol. 117, 345–360 (2001).
Kuner, T., Seeburg, P. H. & Guy, H. R. A common architecture for K+ channels and ionotropic glutamate receptors? Trends Neurosci. 26, 27–32 (2003).
Chen, G. Q., Cui, C., Mayer, M. L. & Gouaux, E. Functional characterization of a potassium-selective prokaryotic glutamate receptor. Nature 402, 817–821 (1999).
Sobolevsky, A. I., Yelshansky, M. V. & Wollmuth, L. P. The outer pore of the glutamate receptor channel has 2-fold rotational symmetry. Neuron 41, 367–378 (2004).
Soderling, T. R. & Derkach, V. A. Postsynaptic protein phosphorylation and LTP. Trends Neurosci. 23, 75–80 (2000).
Tavalin, S. J. et al. Regulation of GluR1 by the A-kinase anchoring protein 79 (AKAP79) signaling complex shares properties with long-term depression. J. Neurosci. 22, 3044–3051 (2002).
Vissel, B., Krupp, J. J., Heinemann, S. F. & Westbrook, G. L. Intracellular domains of NR2 alter calcium-dependent inactivation of N-methyl-D-aspartate receptors. Mol. Pharmacol. 61, 595–605 (2002).
Peng, J. et al. Semiquantitative proteomic analysis of rat forebrain postsynaptic density fractions by mass spectrometry. J. Biol. Chem. 279, 21003–21011 (2004).
Mano, I., Lamed, Y. & Teichberg, V. I. A venus flytrap mechanism for activation and desensitization of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionic acid receptors. J. Biol. Chem. 271, 15299–15302 (1996).
Quiocho, F. A. & Ledvina, P. S. Atomic structure and specificity of bacterial periplasmic receptors for active transport and chemotaxis: variation of common themes. Mol. Microbiol. 20, 17–25 (1996).
Mayer, M. L., Olson, R. & Gouaux, E. Mechanisms for ligand binding to GluR0 ion channels: crystal structures of the glutamate and serine complexes and a closed apo state. J. Mol. Biol. 311, 815–836 (2001).
Watkins, J. C., Krogsgaard-Larsen, P. & Honoré, T. Structure-activity relationships in the development of excitatory amino acid receptor agonists and competitive antagonists. Trends Pharmacol. Sci. 11, 25–33 (1990).
Furukawa, H. & Gouaux, E. Mechanisms of activation, inhibition and specificity: crystal structures of NR1 ligand-binding core. EMBO J. 22, 1–13 (2003).
Mayer, M. L. Crystal structures of the GluR5 and GluR6 ligand binding cores: molecular mechanisms underlying kainate receptor selectivity. Neuron 45, 539–552 (2005).
Armstrong, N. & Gouaux, E. Mechanisms for activation and antagonism of an AMPA-sensitive glutamate receptor: Crystal structures of the GluR2 ligand binding core. Neuron 28, 165–181 (2000).
Jin, R., Banke, T. G., Mayer, M. L., Traynelis, S. F. & Gouaux, E. Structural basis for partial agonist action at ionotropic glutamate receptors. Nature Neurosci. 6, 803–810 (2003).
Hogner, A. et al. Structural basis for AMPA receptor activation and ligand selectivity: Crystal structures of five agonist complexes with the GluR2 ligand-binding core. J. Mol. Biol. 322, 93–109 (2002).
Mayer, M. L., Ghosal, A. K., Dolman, N. P. & Jane, D. E. Crystal structures of the kainate receptor GluR5 ligand binding core dimer with novel GluR5 selective antagonists. J. Neurosci. 26, 2852–2861 (2006).
Colquhoun, D. & Ogden, D. C. Activation of ion channels in the frog end-plate by high concentrations of acetylcholine. J. Physiol. (Lond.) 395, 131–159 (1988).
Rosenmund, C., Stern-Bach, Y. & Stevens, C. F. The tetrameric structure of a glutamate receptor channel. Science 280, 1596–1599 (1998).
Armstrong, N., Mayer, M. & Gouaux, E. Tuning activation of the AMPA-sensitive GluR2 ion channel by genetic adjustment of agonist-induced conformational changes. Proc. Natl Acad. Sci. USA 100, 5736–5741 (2003).
Inanobe, A., Furukawa, H. & Gouaux, E. Mechanism of partial agonist action at the NR1 subunit of NMDA receptors. Neuron 47, 71–84 (2005).
Erreger, K. et al. Mechanism of partial agonism at NMDA receptors for a conformationally restricted glutamate analog. J. Neurosci. 25, 7858–7866 (2005).
Chen, P. E. et al. Structural features of the glutamate binding site in recombinant NR1/NR2A N-methyl-D-aspartate receptors determined by site-directed mutagenesis and molecular modeling. Mol. Pharmacol. 67, 1470–1484 (2005).
Furukawa, H., Singh, S. K., Mancusso, R. & Gouaux, E. Subunit arrangement and function in NMDA receptors. Nature 438, 185–192 (2005).
Mamonova, T., Hespenheide, B., Straub, R., Thorpe, M. F. & Kurnikova, M. Protein flexibility using constraints from molecular dynamics simulations. Phys. Biol. 2, S137–S147 (2005).
Arinaminpathy, Y., Sansom, M. S. & Biggin, P. C. Binding site flexibility: Molecular simulation of partial and full agonists with a glutamate receptor. Mol. Pharmacol. 69, 5–12 (2006).
Jiang, Y. et al. The open pore conformation of potassium channels. Nature 417, 523–526 (2002).
Banke, T. G. & Traynelis, S. F. Activation of NR1/NR2B NMDA receptors. Nature Neurosci. 6, 144–152 (2003).
Schorge, S., Elenes, S. & Colquhoun, D. Maximum likelihood fitting of single channel NMDA activity with a mechanism composed of independent dimers of subunits. J. Physiol. 569, 395–418 (2005).
Sun, Y. et al. Mechanism of glutamate receptor desensitization. Nature 417, 245–253 (2002).
Horning, M. S. & Mayer, M. L. Regulation of AMPA receptor gating by ligand binding core dimers. Neuron 41, 379–388 (2004).
Partin, K. M., Fleck, M. W. & Mayer, M. L. AMPA receptor flip/flop mutants affecting deactivation, desensitization, and modulation by cyclothiazide, aniracetam, and thiocyanate. J. Neurosci. 16, 6634–6647 (1996).
Jin, R. et al. Mechanism of positive allosteric modulators acting on AMPA receptors. J. Neurosci. 25, 9027–9036 (2005).
Yelshansky, M. V., Sobolevsky, A. I., Jatzke, C. & Wollmuth, L. P. Block of AMPA receptor desensitization by a point mutation outside the ligand-binding domain. J. Neurosci. 24, 4728–4736 (2004).
Holm, M. M. et al. A binding site tyrosine shapes desensitization kinetics and agonist potency at GluR2. A mutagenic, kinetic, and crystallographic study. J. Biol. Chem. 280, 35469–35476 (2005).
Lester, R. A. J., Clements, J. D., Westbrook, G. L. & Jahr, C. E. Channel kinetics determine the time course of NMDA receptor-mediated synaptic currents. Nature 346, 565–567 (1990).
Monyer, H. et al. Heteromeric NMDA receptors: molecular and functional distinction of subtypes. Science 256, 1217–1221 (1992).
Laurie, D. J. & Seeburg, P. H. Ligand affinities at recombinant N-methyl-D-aspartate receptors depend on subunit composition. Eur. J. Pharmacol. 268, 335–345 (1994).
Kuner, T., Wollmuth, L. P., Karlin, A., Seeburg, P. H. & Sakmann, B. Structure of the NMDA receptor channel M2 segment inferred from the accessibility of substituted cysteines. Neuron 17, 343–352 (1996).
Beck, C., Wollmuth, L. P., Seeburg, P. H., Sakmann, B. & Kuner, T. NMDAR channel segments forming the extracellular vestibule inferred from the accessibility of substituted cysteines. Neuron 22, 559–570 (1999).
Sobolevsky, A. I., Yelshansky, M. V. & Wollmuth, L. P. State-dependent changes in the electrostatic potential in the pore of a GluR channel. Biophys. J. 88, 235–242 (2005).
Wollmuth, L. P. & Sobolevsky, A. I. Structure and gating of the glutamate receptor ion channel. Trends Neurosci. 27, 321–328 (2004).
Watanabe, J., Beck, C., Kuner, T., Premkumar, L. S. & Wollmuth, L. P. DRPEER: a motif in the extracellular vestibule conferring high Ca2+ flux rates in NMDA receptor channels. J. Neurosci. 22, 10209–10216 (2002).
Schorge, S. & Colquhoun, D. Studies of NMDA receptor function and stoichiometry with truncated and tandem subunits. J. Neurosci. 23, 1151–1158 (2003).
Balannik, V., Menniti, F. S., Paternain, A. V., Lerma, J. & Stern-Bach, Y. Molecular mechanism of AMPA receptor noncompetitive antagonism. Neuron 48, 279–288 (2005).
Lipton, S. A. Paradigm shift in neuroprotection by NMDA receptor blockade: Memantine and beyond. Nature Rev. Drug Discov. 5, 160–170 (2006).
Zuo, J. et al. Neurodegeneration in Lurcher mice caused by mutation in delta2 glutamate receptor gene. Nature 388, 769–773 (1997).
Kohda, K., Wang, Y. & Yuzaki, M. Mutation of a glutamate receptor motif reveals its role in gating and delta2 receptor channel properties. Nature Neurosci. 3, 315–322 (2000).
Klein, R. M. & Howe, J. R. Effects of the lurcher mutation on GluR1 desensitization and activation kinetics. J. Neurosci. 24, 4941–4951 (2004).
Tomita, S. et al. Stargazin modulates AMPA receptor gating and trafficking by distinct domains. Nature 435, 1052–1058 (2005).
Turetsky, D., Garringer, E. & Patneau, D. K. Stargazin modulates native AMPA receptor functional properties by two distinct mechanisms. J. Neurosci. 25, 7438–7448 (2005).
Priel, A. et al. Stargazin reduces desensitization and slows deactivation of the AMPA-type glutamate receptors. J. Neurosci. 25, 2682–2686 (2005).
Acknowledgements
Work in the author's laboratory is supported by the intramural research programme of NICHD, NIH, DHHS. Synchotron diffraction data were collected at Southeast Regional Collaborative Access Team (SER-CAT) 22-ID beamline at the Advanced Photon Source, Argonne National Laboratory. Use of the Advanced Photon Source was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Additional information
Author Information Reprints and permissions information is available at npg.nature.com/reprints and permissions.
Rights and permissions
About this article
Cite this article
Mayer, M. Glutamate receptors at atomic resolution. Nature 440, 456–462 (2006). https://doi.org/10.1038/nature04709
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04709
This article is cited by
-
Association of NOS1 gene polymorphisms with cerebral palsy in a Han Chinese population: a case-control study
BMC Medical Genomics (2018)
-
Mutations of N-Methyl-D-Aspartate Receptor Subunits in Epilepsy
Neuroscience Bulletin (2018)
-
Synaptic AMPA receptor composition in development, plasticity and disease
Nature Reviews Neuroscience (2016)
-
Structural basis of kainate subtype glutamate receptor desensitization
Nature (2016)
-
Association of genetic variants of GRIN2B with autism
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.