Abstract
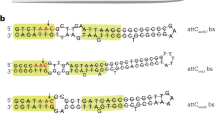
Lateral DNA transfer—the movement of genetic traits between bacteria—has a profound impact on genomic evolution and speciation. The efficiency with which bacteria incorporate genetic information reflects their capacity to adapt to changing environmental conditions. Integron integrases are proteins that mediate site-specific DNA recombination between a proximal primary site (attI) and a secondary target site (attC) found within mobile gene cassettes encoding resistance or virulence factors. The lack of sequence conservation among attC sites has led to the hypothesis that a sequence-independent structural recognition determinant must exist within attC. Here we report the crystal structure of an integron integrase bound to an attC substrate. The structure shows that DNA target site recognition and high-order synaptic assembly are not dependent on canonical DNA but on the position of two flipped-out bases that interact in cis and in trans with the integrase. These extrahelical bases, one of which is required for recombination in vivo, originate from folding of the bottom strand of attC owing to its imperfect internal dyad symmetry. The mechanism reported here supports a new paradigm for how sequence-degenerate single-stranded genetic material is recognized and exchanged between bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kazazian, H. H. Jr. Mobile elements: drivers of genome evolution. Science 303, 1626–1632 (2004)
Craig, N. L., Craigie, R., Gellert, M. & Lambowitz, A. M. (eds) Mobile DNA II (ASM Press, Washington DC, 2002)
Messier, N. & Roy, P. H. Integron integrases possess a unique additional domain necessary for activity. J. Bacteriol. 183, 6699–6706 (2001)
Chen, Y. & Rice, P. A. New insight into site-specific recombination from Flp recombinase–DNA structures. Annu. Rev. Biophys. Biomol. Struct. 32, 135–159 (2003)
Van Duyne, G. A structural view of tyrosine recombinase site-specific recombination. In Mobile DNA II (eds Craig, N. L., Craigie, R., Gellert, M. & Lambowitz, A. M.) (ASM Press, Washington DC, 2002)
Azaro, M. A. & Landy, A. Lambda integrase and the lambda Int family. In Mobile DNA II (eds Craig, N. L., Craigie, R., Gellert, M. & Lambowitz, A. M.) (ASM Press, Washington DC, 2002)
Stokes, H. W., O'Gorman, D. B., Recchia, G. D., Parsekhian, M. & Hall, R. M. Structure and function of 59-base element recombination sites associated with mobile gene cassettes. Mol. Microbiol. 26, 731–745 (1997)
Gravel, A., Fournier, B. & Roy, P. H. DNA complexes obtained with the integron integrase IntI1 at the attI1 site. Nucleic Acids Res. 26, 4347–4355 (1998)
Stark, W. M., Boocock, M. R. & Sherratt, D. J. Site-specific recombination by Tn3 resolvase. Trends Genet. 5, 304–309 (1989)
Guo, F., Gopaul, D. N. & van Duyne, G. D. Structure of Cre recombinase complexed with DNA in a site-specific recombination synapse. Nature 389, 40–46 (1997)
Hansson, K., Skold, O. & Sundstrom, L. Non-palindromic attI sites of integrons are capable of site-specific recombination with one another and with secondary targets. Mol. Microbiol. 26, 441–453 (1997)
Bouvier, M., Demarre, G. & Mazel, D. Integron cassette insertion: a recombination process involving a folded single strand substrate. EMBO J. 24, 4356–4367 (2005)
Biskri, L., Bouvier, M., Guerout, A. M., Boisnard, S. & Mazel, D. Comparative study of class 1 integron and Vibrio cholerae superintegron integrase activities. J. Bacteriol. 187, 1740–1750 (2005)
Rowe-Magnus, D. A., Guerout, A. M., Biskri, L., Bouige, P. & Mazel, D. Comparative analysis of superintegrons: engineering extensive genetic diversity in the Vibrionaceae. Genome Res. 13, 428–442 (2003)
Johansson, C., Kamali-Moghaddam, M. & Sundstrom, L. Integron integrase binds to bulged hairpin DNA. Nucleic Acids Res. 32, 4033–4043 (2004)
Francia, M. V., Zabala, J. C., de la Cruz, F. & Garcia Lobo, J. M. The IntI1 integron integrase preferentially binds single-stranded DNA of the attC site. J. Bacteriol. 181, 6844–6849 (1999)
Mazel, D., Dychinco, B., Webb, V. A. & Davies, J. A distinctive class of integron in the Vibrio cholerae genome. Science 280, 605–608 (1998)
Subramanya, H. S. et al. Crystal structure of the site-specific recombinase, XerD. EMBO J. 16, 5178–5187 (1997)
Aihara, H., Kwon, H. J., Nunes-Duby, S. E., Landy, A. & Ellenberger, T. A conformational switch controls the DNA cleavage activity of λ integrase. Mol. Cell 12, 187–198 (2003)
Chen, Y., Narendra, U., Iype, L. E., Cox, M. M. & Rice, P. A. Crystal structure of a Flp recombinase–Holliday junction complex: assembly of an active oligomer by helix swapping. Mol. Cell 6, 885–897 (2000)
Kwon, H. J., Tirumalai, R., Landy, A. & Ellenberger, T. Flexibility in DNA recombination: structure of the lambda integrase catalytic core. Science 276, 126–131 (1997)
Hickman, A. B., Waninger, S., Scocca, J. J. & Dyda, F. Molecular organization in site-specific recombination: the catalytic domain of bacteriophage HP1 integrase at 2.7 Å resolution. Cell 89, 227–237 (1997)
Krogh, B. O. & Shuman, S. Catalytic mechanism of DNA topoisomerase IB. Mol. Cell 5, 1035–1041 (2000)
Rafferty, J. B. et al. Crystal structure of DNA recombination protein RuvA and a model for its binding to the Holliday junction. Science 274, 415–421 (1996)
Gopaul, D. N. & Van Duyne, G. D. Structure and mechanism in site-specific recombination. Curr. Opin. Struct. Biol. 9, 14–20 (1999)
Arciszewska, L. K., Grainge, I. & Sherratt, D. J. Action of site-specific recombinases XerC and XerD on tethered Holliday junctions. EMBO J. 16, 3731–3743 (1997)
Biswas, T. et al. A structural basis for allosteric control of DNA recombination by λ integrase. Nature 435, 1059–1066 (2005)
Nunes-Duby, S. E., Azaro, M. A. & Landy, A. Swapping DNA strands and sensing homology without branch migration in λ site-specific recombination. Curr. Biol. 5, 139–148 (1995)
Gopaul, D. N., Guo, F. & Van Duyne, G. D. Structure of the Holliday junction intermediate in Cre–loxP site-specific recombination. EMBO J. 17, 4175–4187 (1998)
Meyer-Leon, L., Inman, R. B. & Cox, M. M. Characterization of Holliday structures in FLP protein-promoted site-specific recombination. Mol. Cell. Biol. 10, 235–242 (1990)
Ochman, H., Lawrence, J. G. & Groisman, E. A. Lateral gene transfer and the nature of bacterial innovation. Nature 405, 299–304 (2000)
Klimasauskas, S., Kumar, S., Roberts, R. J. & Cheng, X. HhaI methyltransferase flips its target base out of the DNA helix. Cell 76, 357–369 (1994)
Val, M. E. et al. The single-stranded genome of phage CTX is the form used for integration into the genome of Vibrio cholerae. Mol. Cell 19, 559–566 (2005)
Ronning, D. R. et al. Active site sharing and subterminal hairpin recognition in a new class of DNA transposases. Mol. Cell 20, 143–154 (2005)
Llosa, M., Gomis-Ruth, F. X., Coll, M. & de la Cruz, F. Bacterial conjugation: a two-step mechanism for DNA transport. Mol. Microbiol. 45, 1–8 (2002)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Cryst. 26, 795–800 (1993)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Weeks, C. M. et al. Towards automated protein structure determination: BnP, the SnB–PHASES interface. Z. Kristallogr. 217, 686–693 (2002)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Brunger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Acknowledgements
This work was supported by laboratory start-up funds from the Pasteur Institute to D.N.G., research grants from the CNRS to D.N.G., research grants from the Institut Pasteur and from the CNRS on transposable elements to D. Mazel, and a French Ministry of Research (MENESR) grant to D. Mazel and D.N.G. D. MacDonald is a Florence Gould Scholar funded by the Pasteur Foundation of New York. G.D. and M.B. are doctoral fellows from the Fondation pour la Recherche Médicale and MENESR, respectively. We also thank T. Ellenberger and G. Liou for critical review of this manuscript. Author Contributions D. MacDonald carried out the in vitro assays, structural determination and wrote the paper with editing from D.N.G. D. Mazel provided the conceptual framework for the substrate and invaluable background. G.D. and M.B. performed the in vivo excision frequency assays. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Coordinates and structure factors are deposited in the Protein Data Bank under accession code 2A3V. Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figure 1
This figure contains the predicted secondary structures of attC bottom strands. (PDF 94 kb)
Supplementary Figure 2
Supplementary Figure 2 nature04643-s02.pdf This figure contains stereo views of the VchIntIA–VCRbs active sites. (PDF 941 kb)
Supplementary Figure 3
This figure contains a stereo model of a VchIntIA attacking subunit and the sequence alignment of VchIntIA and Cre recombinase with their corresponding secondary structure elements. (PDF 259 kb)
Supplementary Figure 4
This figure contains a schematic representation of the protein-DNA contacts between VchIntIA and VCRbs. (PDF 225 kb)
Supplementary Figure 5
This figure contains a model of the proposed movement of the β 4,5 hairpin. (PDF 841 kb)
Supplementary Figure 6
This figure contains data from EMSA and in vivo excision assays. (PDF 124 kb)
Supplementary Figure 7
This figure contains proposed IntI double strand exchange pathway via a single-stranded DNA substrate. (PDF 141 kb)
Supplementary Figure 8
This figure contains the rotation of the C-terminal domain within the non-attacking subunits. (PDF 633 kb)
Supplementary Table 1
Data collection and refinement statistics (PDF 125 kb)
Supplementary Movie 1
This movie shows the final model of the VchIntIA–VCRbs synaptic complex. (WMV 4614 kb)
Supplementary Notes
This file contains Supplementary Methods, Supplementary Figure Legends and additional references. (DOC 56 kb)
Rights and permissions
About this article
Cite this article
MacDonald, D., Demarre, G., Bouvier, M. et al. Structural basis for broad DNA-specificity in integron recombination. Nature 440, 1157–1162 (2006). https://doi.org/10.1038/nature04643
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04643
This article is cited by
-
Influence of class 2 integron integrase concentration on gene cassette insertion and excision in vivo
Brazilian Journal of Microbiology (2023)
-
The association of group IIB intron with integrons in hypersaline environments
Mobile DNA (2021)
-
Comment on “Conserved phylogenetic distribution and limited antibiotic resistance of class 1 integrons revealed by assessing the bacterial genome and plasmid collection” by A.N. Zhang et al.
Microbiome (2021)
-
Comparison of Class 2 Integron Integrase Activities
Current Microbiology (2021)
-
A comprehensive survey of integron-associated genes present in metagenomes
BMC Genomics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.