Abstract
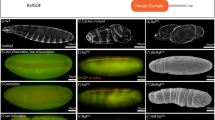
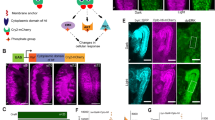
Breaking left–right symmetry in Bilateria embryos is a major event in body plan organization that leads to polarized adult morphology, directional organ looping, and heart and brain function1,2,3,4. However, the molecular nature of the determinant(s) responsible for the invariant orientation of the left–right axis (situs choice) remains largely unknown. Mutations producing a complete reversal of left–right asymmetry (situs inversus) are instrumental for identifying mechanisms controlling handedness, yet only one such mutation has been found in mice (inversin)5 and snails6,7. Here we identify the conserved type ID unconventional myosin 31DF gene (Myo31DF) as a unique situs inversus locus in Drosophila. Myo31DF mutations reverse the dextral looping of genitalia, a prominent left–right marker in adult flies. Genetic mosaic analysis pinpoints the A8 segment of the genital disc as a left–right organizer and reveals an anterior–posterior compartmentalization of Myo31DF function that directs dextral development and represses a sinistral default state. As expected of a determinant, Myo31DF has a trigger-like function and is expressed symmetrically in the organizer, and its symmetrical overexpression does not impair left–right asymmetry. Thus Myo31DF is a dextral gene with actin-based motor activity controlling situs choice. Like mouse inversin8, Myo31DF interacts and colocalizes with β-catenin, suggesting that situs inversus genes can direct left–right development through the adherens junction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Bisgrove, B. W., Morelli, S. H. & Yost, H. J. Genetics of human laterality disorders: insights from vertebrate model systems. Annu. Rev. Genomics Hum. Genet. 4, 1–32 (2003)
Levin, M. Left–right asymmetry in embryonic development: a comprehensive review. Mech. Dev. 122, 3–25 (2005)
Hamada, H., Meno, C., Watanabe, D. & Saijoh, Y. Establishment of vertebrate left–right asymmetry. Nature Rev. Genet. 3, 103–113 (2002)
Raya, A. & Belmonte, J. C. Sequential transfer of left–right information during vertebrate embryo development. Curr. Opin. Genet. Dev. 14, 575–581 (2004)
Morgan, D. et al. Inversin, a novel gene in the vertebrate left–right axis pathway, is partially deleted in the inv mouse. Nature Genet. 20, 149–156 (1998)
Ueshima, R. & Asami, T. Evolution: single-gene speciation by left–right reversal. Nature 425, 679 (2003)
Freeman, G. & Lundelius, J. The developmental genetics of dextrality and sinistrality in the gastropod Lymnaea peregra. Wilhelm Roux's Archives 191, 69–83 (1982)
Nurnberger, J., Bacallao, R. L. & Phillips, C. L. Inversin forms a complex with catenins and N-cadherin in polarized epithelial cells. Mol. Biol. Cell 13, 3096–3106 (2002)
Adam, G., Perrimon, N. & Noselli, S. The retinoic-like juvenile hormone controls the looping of left–right asymmetric organs in Drosophila. Development 130, 2397–2406 (2003)
Gleichauf, R. Anatomie und variabilität des geschlechtsapparates von Drosophila melanogaster (Meigen). Z. Wiss. Zool. 148, 1–66 (1936)
Morgan, N. S., Heintzelman, M. B. & Mooseker, M. S. Characterization of myosin-IA and myosin-IB, two unconventional myosins associated with the Drosophila brush border cytoskeleton. Dev. Biol. 172, 51–71 (1995)
Morgan, D. et al. The left–right determinant inversin has highly conserved ankyrin repeat and IQ domains and interacts with calmodulin. Hum. Genet. 110, 377–384 (2002)
Eley, L. et al. A perspective on inversin. Cell Biol. Int. 28, 119–124 (2004)
Bahler, M. & Rhoads, A. Calmodulin signaling via the IQ motif. FEBS Lett. 513, 107–113 (2002)
Casares, F., Sanchez, L., Guerrero, I. & Sanchez-Herrero, E. The genital disc of Drosophila melanogaster. 1. Segmental and compartmental organization. Dev. Genes Evol. 207, 216–228 (1997)
Chen, E. H. & Baker, B. S. Compartmental organization of the Drosophila genital imaginal discs. Development 124, 205–218 (1997)
Freeland, D. E. & Kuhn, D. T. Expression patterns of developmental genes reveal segment and parasegment organization of D. melanogaster genital discs. Mech. Dev. 56, 61–72 (1996)
Sanchez, L., Casares, F., Gorfinkiel, N. & Guerrero, I. The genital disc of Drosophila melanogaster. 2. Role of the genes hedgehog, decapentaplegic and wingless. Dev. Genes Evol. 207, 229–241 (1997)
Watanabe, D. et al. The left–right determinant inversin is a component of node monocilia and other 9 + 0 cilia. Development 130, 1725–1734 (2003)
Bobinnec, Y., Marcaillou, C. & Debec, A. Microtubule polyglutamylation in Drosophila melanogaster brain and testis. Eur. J. Cell Biol. 78, 671–674 (1999)
Dubruille, R. et al. Drosophila regulatory factor X is necessary for ciliated sensory neuron differentiation. Development 129, 5487–5498 (2002)
Bonnafe, E. et al. The transcription factor RFX3 directs nodal cilium development and left–right asymmetry specification. Mol. Cell. Biol. 24, 4417–4427 (2004)
Macias, A. et al. PVF1/PVR signaling and apoptosis promotes the rotation and dorsal closure of the Drosophila male terminalia. Int. J. Dev. Biol. 48, 1087–1094 (2004)
Shibazaki, Y., Shimizu, M. & Kuroda, R. Body handedness is directed by genetically determined cytoskeletal dynamics in the early embryo. Curr. Biol. 14, 1462–1467 (2004)
Garcia-Castro, M. I., Vielmetter, E. & Bronner-Fraser, M. N-Cadherin, a cell adhesion molecule involved in establishment of embryonic left–right asymmetry. Science 288, 1047–1051 (2000)
Huber, L. A. et al. Both calmodulin and the unconventional myosin Myr4 regulate membrane trafficking along the recycling pathway of MDCK cells. Traffic 1, 494–503 (2000)
Bertet, C., Sulak, L. & Lecuit, T. Myosin-dependent junction remodelling controls planar cell intercalation and axis elongation. Nature 429, 667–671 (2004)
Strutt, H. & Strutt, D. EGF signaling and ommatidial rotation in the Drosophila eye. Curr. Biol. 13, 1451–1457 (2003)
Acknowledgements
We thank the Bloomington Stock Center, B. Durand, B. Edde, A. Laurençon, V. van de Bor, J. P. Vincent and M. L. Cariou for materials and fly lines; E. Sanchez-Herrero for AbdB–Gal4; Y. Bellaiche and M. Morgan for GST–Arm protein; C. Mionnet for help with GST-pulldown assays; and C. Featherstone, P. Follette, E. Sanchez-Herrero, M. Suzanne, L. Wolpert and laboratory members for critically reading the manuscript. This work was supported by the Centre National de la Recherche Scientifique (CNRS), the Ministère de l'éducation et de la Recherche (ACI), the Hungarian National Scientific Research Fund (OTKA), the Association pour la Recherche contre le Cancer (ARC) and the EMBO Young Investigator Programme.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Data
This file contains figure legends for Supplementary Figures 1–6 as well as Supplementary Methods (DOC 25 kb)
Supplementary Table 1
Direction of genitalia rotation in several Drosophila species. (DOC 28 kb)
Supplementary Figure 1
Myo31DF is a conserved type ID unconventional myosin that colocalizes with F-actin. (PDF 2562 kb)
Supplementary Figure 2
Characterization of anti-Myo31DF antibodies. (PDF 215 kb)
Supplementary Figure 3
Developmental expression pattern of Myo31DF in the genital disc. (PDF 381 kb)
Supplementary Figure 4
Expression pattern of Ptc-GAL4 in the A8 segment. (PDF 410 kb)
Supplementary Figure 5
Schematic representation of the non-rotated phenotype. (PDF 105 kb)
Supplementary Figure 6
Interaction between Myo31DF and Armadillo. (PDF 156 kb)
Rights and permissions
About this article
Cite this article
Spéder, P., Ádám, G. & Noselli, S. Type ID unconventional myosin controls left–right asymmetry in Drosophila. Nature 440, 803–807 (2006). https://doi.org/10.1038/nature04623
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04623
This article is cited by
-
Asymmetric activity of NetrinB controls laterality of the Drosophila brain
Nature Communications (2023)
-
Actin polymerisation and crosslinking drive left-right asymmetry in single cell and cell collectives
Nature Communications (2023)
-
Proper direction of male genitalia is prerequisite for copulation in Drosophila, implying cooperative evolution between genitalia rotation and mating behavior
Scientific Reports (2019)
-
Vertebrate myosin 1d regulates left–right organizer morphogenesis and laterality
Nature Communications (2018)
-
Myosin1D is an evolutionarily conserved regulator of animal left–right asymmetry
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.