Abstract
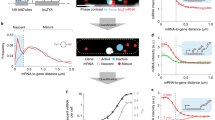
In a living cell, gene expression—the transcription of DNA to messenger RNA followed by translation to protein—occurs stochastically, as a consequence of the low copy number of DNA and mRNA molecules involved1,2,3,4,5,6. These stochastic events of protein production are difficult to observe directly with measurements on large ensembles of cells owing to lack of synchronization among cells. Measurements so far on single cells lack the sensitivity to resolve individual events of protein production. Here we demonstrate a microfluidic-based assay that allows real-time observation of the expression of β-galactosidase in living Escherichia coli cells with single molecule sensitivity. We observe that protein production occurs in bursts, with the number of molecules per burst following an exponential distribution. We show that the two key parameters of protein expression—the burst size and frequency—can be either determined directly from real-time monitoring of protein production or extracted from a measurement of the steady-state copy number distribution in a population of cells. Application of this assay to probe gene expression in individual budding yeast and mouse embryonic stem cells demonstrates its generality. Many important proteins are expressed at low levels7,8, and are thus inaccessible by current genomic and proteomic techniques. This microfluidic single cell assay opens up possibilities for system-wide characterization of the expression of these low copy number proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
McAdams, H. H. & Arkin, A. Stochastic mechanisms in gene expression. Proc. Natl Acad. Sci. USA 94, 814–819 (1997)
Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002)
Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002)
Blake, W. J., Kærn, M., Cantor, C. R. & Collins, J. J. Noise in eukaryotic gene expression. Nature 422, 633–637 (2003)
Raser, J. M. & O'Shea, E. K. Control of stochasticity in eukaryotic gene expression. Science 304, 1811–1814 (2004)
Rosenfeld, N., Young, J. W., Alon, U., Swain, P. S. & Elowitz, M. B. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005)
Guptasarma, P. Does replication-induced transcription regulate synthesis of the myriad low copy number proteins of Escherichia coli? Bioessays 17, 987–997 (1995)
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003)
Miller, J. H. Experiments in Molecular Genetics (Cold Spring Harbor Laboratory, New York, 1972)
Russo-Marie, F., Roederer, M., Sager, B., Herzenberg, L. A. & Kaiser, D. β-galactosidase activity in single differentiating bacterial cells. Proc. Natl Acad. Sci. USA 90, 8194–8198 (1993)
Novick, A. & Weiner, M. Enzyme induction as an all-or-none phenomenon. Proc. Natl Acad. Sci. USA 43, 553–566 (1957)
Nolan, G. P., Fiering, S., Nicolas, J. F. & Herzenberg, L. A. Fluorescence-activated cell analysis and sorting of viable mammalian cells based on β-D-galactosidase activity after transduction of Escherichia coli lacZ. Proc. Natl Acad. Sci. USA 85, 2603–2607 (1988)
Rotman, B. Measurement of activity of single molecules of β-D-galactosidase. Proc. Natl Acad. Sci. USA 47, 1981–1991 (1961)
Nikaido, H. Prevention of drug access to bacterial targets: permeability barriers and active efflux. Science 264, 382–388 (1994)
Unger, M. et al. Monolithic microfabricated valves and pumps by multilayer soft lithography. Science 288, 113–116 (2000)
Wu, H., Wheeler, A. & Zare, R. N. Chemical cytometry on a picoliter-scale integrated microfluidic chip. Proc. Natl Acad. Sci. USA 101, 12809–12813 (2004)
McDonald, J. C. et al. Fabrication of microfluidic systems in poly(dimethylsiloxane). Electrophoresis 21, 27–40 (2000)
Balagadde, F. K., You, L., Hansen, C. L., Arnold, F. H. & Quake, S. R. Long-term monitoring of bacteria undergoing programmed population control in a microchemostat. Science 309, 137–140 (2005)
Rondelez, Y. et al. Microfabricated arrays of femtoliter chambers allow single molecule enzymology. Nature Biotechnol. 23, 361–365 (2005)
Monod, J., Pappenheimer, A. M. Jr & Cohen-Bazire, G. The kinetics of the biosynthesis of β-galactosidase in Escherichia coli as a function of growth. Biochim. Biophys. Acta 9, 648–660 (1952)
Kennell, D. & Riezman, H. Transcription and translation initiation frequencies of the Escherichia coli lac operon. J. Mol. Biol. 114, 1–21 (1977)
Sorensen, M. & Pedersen, S. Absolute in vivo translation rates of individual codons in Escherichia coli. The two glutamic acid codons GAA and GAG are translated with a threefold difference in rate. J. Mol. Biol. 222, 265–280 (1991)
Berg, O. G. A model for the statistical fluctuations of protein number in a microbial population. J. Theor. Biol. 71, 587–603 (1975)
Yarchuk, O., Jacques, N., Guillerez, J. & Dreyfus, M. Interdependence of translation, transcription and mRNA degradation in the lacZ gene. J. Mol. Biol. 226, 581–596 (1992)
Paulsson, J., Berg, O. G. & Ehrenberg, M. Stochastic focusing: fluctuation-enhanced sensitivity of intracellular regulation. Proc. Natl Acad. Sci. USA 97, 7148–7153 (2000)
Ozbudak, E. M., Thattai, M., Lim, H. N., Shraiman, B. I. & Van Oudenaarden, A. Multistability in the lactose utilization network of Escherichia coli. Nature 427, 737–740 (2004)
Levsky, J. M., Shenoy, S. M., Pezo, R. C. & Singer, R. H. Single-cell gene expression profiling. Science 297, 836–840 (2002)
Golding, I., Paulsson, J., Zawilski, S. M. & Cox, E. C. Real-time kinetics of gene activity in individual bacteria. Cell 123, 1025–1036 (2005)
Yu, J. et al. Probing gene expression in live cells, one protein molecule at a time. Science 311, 1600–1603 (2006)
Efron, B. & Tibshirani, R. J. An Introduction to the Bootstrap (Chapman & Hall, New York, 1993)
Acknowledgements
We acknowledge J. Xiao, K. Gudiksen and J. Markson for early involvement in the project, and J. Yu, P. Choi and J. Elf for discussions and careful reading of the manuscript. We are grateful to Q. Xu, H. Wu, members of G. M. Whitesides and R. N. Zare groups, and the Harvard Center for Nanoscale Systems for help with microfluidic fabrication. We thank K. Thorn for yeast strains, and A. and J. McMahon for stem cell strains. This work was funded by Department of Energy, Office of Biological and Environmental Research, Genomics: GTL Program, and in part by National Institute of Health (NIH) Director's Pioneer Award to X.S.X., and an NIH grant. L.C. is supported by a National Science Foundation Graduate Research Fellowship and N.F. by a Human Frontiers Science Program Long Term Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains the Supplementary Methods, Supplementary Figures 1–12 and additional references.(DOC 280 kb)
Rights and permissions
About this article
Cite this article
Cai, L., Friedman, N. & Xie, X. Stochastic protein expression in individual cells at the single molecule level. Nature 440, 358–362 (2006). https://doi.org/10.1038/nature04599
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04599
This article is cited by
-
Frequency modulation of a bacterial quorum sensing response
Nature Communications (2022)
-
Independent control of mean and noise by convolution of gene expression distributions
Nature Communications (2021)
-
Measuring expression heterogeneity of single-cell cytoskeletal protein complexes
Nature Communications (2021)
-
Probing low-copy-number proteins in single living cells using single-cell plasmonic immunosandwich assays
Nature Protocols (2021)
-
Neural network aided approximation and parameter inference of non-Markovian models of gene expression
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.