Abstract
Human chromosome 12 contains more than 1,400 coding genes1 and 487 loci that have been directly implicated in human disease2. The q arm of chromosome 12 contains one of the largest blocks of linkage disequilibrium found in the human genome3. Here we present the finished sequence of human chromosome 12, which has been finished to high quality and spans approximately 132 megabases, representing ∼4.5% of the human genome. Alignment of the human chromosome 12 sequence across vertebrates reveals the origin of individual segments in chicken, and a unique history of rearrangement through rodent and primate lineages. The rate of base substitutions in recent evolutionary history shows an overall slowing in hominids compared with primates and rodents.
Similar content being viewed by others
Main
Among the human chromosome sequencing projects, the chromosome 12 sequencing effort benefited most from an earlier advanced sequence tagged site (STS) physical map, which contained 5,300 large-insert clones and 3,100 markers with an average resolution of 44 kilobases (kb)4. After integration with the whole genome fingerprint map5, a final tiling path of 1,168 large-insert clones was chosen for sequencing using the clone-by-clone shotgun sequencing strategy. Each clone was finished to community standards (http://www.genome.gov/10001812) yielding 130,683,379 base pairs (bp) of nonoverlapping sequence, independently measured as greater than 99.99% accurate6. The features and annotations presented here may be viewed as user-specified tracks within the Genboree genome browser (http://www.genboree.org/Hs.chr12).
The finished sequence contains just five euchromatic gaps, estimated to total 380 kb by fibre fluorescence in situ hybridization (FISH) (C. Wagner-McPherson, personal communication) and by comparison to primate and rodent genome assemblies (Supplementary Table 1). The data extend to 16 and 60 kb from the telomere terminus repeats at the p and q arms, respectively (H. Riethman, personal communication; http://www.wistar.upenn.edu/Riethman/)7. The pericentromeric sequences on the p and q arms contain approximately 425 and 600 kb of tandem alpha-satellite repeats, respectively. The alpha satellites do not demonstrate the higher-order structure indicative of the ‘core centromere’ on either arm, but previously established markers8, which are present in BAC clones flanking the centromere, result in a calculated centromere length of 1.395 megabases (Mb) and an overall chromosome length of 132,449,811 bp.
Starting with an automated analysis of human build 33 using the Ensembl pipeline9, we manually annotated the chromosome 12 finished sequence using evidence from all publicly available protein and complementary DNA databases, as well as spliced expressed sequence tags (ESTs). We identified a further 282 gene structures beyond the automated output, resulting in a total of 1,435 loci. Using annotation standards developed by the Human Annotation Workshop (http://www.sanger.ac.uk/HGP/havana/hawk.shtml), the loci were categorized as 1,294 ‘known genes’, 12 ‘novel coding sequences (CDS)’, 34 ‘novel transcripts’, 2 ‘putative genes’ and 93 ‘pseudogenes’.
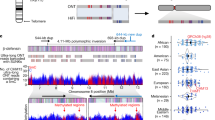
We found 4,427 paralogous pairings to genes on chromosome 12, of which 528 were intrachromosomal. Notably, the density of paralogues correlates well with the density of SINEs (short interspersed elements) and breakpoints observed between the human chromosome and its syntenic regions in avian, rodent and canine genomes (see Fig. 1). Excluding pseudogenes, the average gene density on chromosome 12 is 11.0 genes per Mb, which is relatively typical when compared to the gene densities of other chromosomes. A total of just 11 out of 1,264 RefSeq genes1 are completely or partially absent from genome build 33, while there are only three partially missing genes (NM_001733, C1R; NM_018711, SVOP; and NM_020993, BCL7A) from build 35.
Tracks are numbered on the left, and syntenic alignments across human chromosome 12 are shown in the top five tracks: 1, human–chicken; 2, human–rat; 3, human–mouse; 4, human–dog; and 5, human–chimpanzee. The inter- and intrachromosomal breakpoints are represented by red and blue gaps, respectively. Aqua gaps indicate regions without sequence alignment, and the centromere is denoted by the grey bar running through all tracks. Purple brackets portray sequence inversions. The density of recent segmental intra- and interchromosomal duplications from low-copy repeats are shown in tracks 6 and 7, respectively. The incidence of major interspersed (high-copy) repeats is depicted in tracks 8 (LINEs), 9 (LTRs) and 10 (SINEs). The variations in G + C content, and densities of CpG islands, genes and pseudogenes, appear in tracks 11, 12, 13 and 14, respectively, while gene paralogue density, gene density and gene variant density appear in tracks 15, 16 and 17, respectively. Gene density in track 13 is from UCSC ‘known genes’, and in track 16 is from the nonredundant locus annotations performed in this study. The ymax values in tracks 14–17 reflect the maximum y-axis values obtained for those tracks.
Approximately 58.3% of chromosome 12 genes expressed alternative transcripts, averaging 2.89 transcripts per gene, but ranging as high as 20 (for UBC). The majority of these produced altered protein products (2,923 different proteins from among 3,148 alternative transcripts). There were at least 677 partial transcripts, based primarily on spliced EST data, for which we could not identify the complete coding sequence. Therefore, these estimates represent a lower bound on the total alternative splicing activity on the chromosome.
The density of genes across chromosome 12 varies quite widely (see Fig. 1) and there are three large gene clusters: the natural killer cell gene cluster (9 genes) at 12p13.2–12p12.3, the type II keratin gene cluster (14 genes) at 12q13.13, and the homeobox C gene cluster (9 genes) at 12q13. There are also smaller clusters encoding the voltage-gated potassium channels (3 genes) at 12p13, the aquaporin gene cluster (3 genes) at 12q13.1, and a large cluster encoding salivary proline-rich proteins (7 genes) at 12p13.2.
There are 993 segmental duplications (defined as having greater than 90% identity and being >1 kb), which accounts for 2.66% of the chromosome (versus 5.37% for the entire genome), with particular activity in the pericentromeric region of the p arm, and the telomeres. These duplicated regions represent the fraction of the genomic sequence that was most improved in the finishing process, and are a rich resource for the study of gene clusters and large-scale human DNA polymorphism. The chromosome is typical of the remainder of the genome with respect to noncoding RNA (Supplementary Table 2), LINE (long interspersed elements) and SINE distribution, CpG island distribution, and overall G + C content.
A comparison of physical and genetic maps revealed wide variation in recombination activity, with a slightly higher overall recombination rate in females as expected, and an average of 1.3 centimorgans (cM) per Mb (see Supplementary Fig. 1). There are, however, no extensive recombination ‘deserts’ or ‘jungles’ (with ‘jungles’ defined as having a recombination rate of >3 cM per Mb) as previously described10.
One exceptional region of 12q showed a very large block of linkage disequilibrium spanning 987 kb (compared with a chromosomal average of 26 kb) in all three continental populations assayed. This is one of the largest structures of its kind in the genome, and is associated with evidence for recent positive selection of a pre-expansion ATXN2 gene CAG repeat allele in Americans of European ancestry3.
The history of individual human chromosomes can be reconstructed by tracing blocks of conserved synteny across species, and there is sufficient conservation in the chicken genome (> 300 million years (Myr) of evolutionary separation from humans) to identify 13 segments representing about 72% of human chromosome 12. Reconstruction of a more recent ancestral mammalian genome shows a double reciprocal rearrangement between two ancient acrocentric chromosomes that were comprised primarily of material syntenic to chromosomes 12 and 22, leading to the current structure (Fig. 2)11. The rearrangement occurred in the anthropoid line sometime after the divergence of prosimians. While the chromosome has not undergone any major subsequent rearrangements, pericentric inversions have occurred independently in the chimpanzee and gorilla orthologues12,13.
Numbers above the chromosome indicate the human orthologous chromosome, and numbers to the left of the chromosome refer to the chromosome of the compared organism. Regions orthologous to human chromosome 12 are shown in green. The rodent comparisons to human were performed using Pash28, and the ancestral mammal and chicken are adapted from GRIMM (Genome Rearrangements In Man and Mouse)-Synteny computations presented in ref. 30. On the basis of cytogenetic evidence, a double reciprocal translocation of human chromosomes 12 and 22 occurred after the divergence of prosimian primates, and is present in all anthropoid primates.
Alignment to rodent chromosomes (> 80 Myr evolutionary separation) shows that approximately 9 major rearrangements have occurred between human chromosome 12 and the hypothetical common ancestor with rat and mouse. The breakpoints identified between the human and rodent genomes are summarized in Fig. 1. An unexpected feature is the pattern of increased activity at the ends of the chromosome, with a substantial portion of the proximal q arm relatively unaffected by gross rearrangement across mammals. The detailed examination of breakpoints between the available genome sequences does not suggest any obvious sequence features that correlate with the evolutionary change, and the pattern of breaks otherwise generally conforms to that predicted from the known evolutionary relationships. There is an observed increase in the number of intra- and interchromosomal duplications in the regions where breaks occur, but a full analysis awaits a comprehensive description of duplication in the nonhuman species, which are all currently at draft coverage.
There are innumerable ‘small-scale’ differences between regions of the rodent and human chromosomes with conserved synteny. The data presented here confirm earlier studies14 showing that the rate of change of single base differences and small insertions was greater in the rodent than in hominid lineages, while the deletion rates were slightly lower. Several studies have attempted to correlate the frequency and distribution of these small-scale changes with other genome features. For example, the relative increase in insertions correlates with the active expansion of the overall size of human chromosomes, and a genome-wide burst of SINE insertions (mainly AluJ and AluS repeats) in the human lineage. Furthermore, the density of SINEs shows a modest positive correlation with the microinsertion rate (r = 0.66), a strong negative correlation with the substitution rate (r = -0.73), but no correlation with microdeletions. The SINE distribution correlations may be due to local G + C content, but the relationships are complicated. For example, G + C density shows modest-to-strong positive correlation with both SINEs and insertions. However, SINEs are known to prefer to insert in locally G + C-rich DNA, and this alone may govern the connection between G + C content and the other events. The human chromosomes have a lower G + C content than in rodents, leading to a generally lower rate of G or C gain substitutions (compared with A/T gains) (Supplementary Table 3) exacerbating the complexity of the effect of G + C SINE insertions14,15.
To better understand more recent small-scale changes leading to the current human sequence, we compared chromosome 12 to the available orthologous regions of the chimpanzee and rhesus macaque genomes. Although neither nonhuman assembly represents a ‘deep draft’, high sequence identity enabled alignment of about 87% of the human chromosome to both nonhuman primates. The availability of two additional primate genomes allowed reconstruction of the ancestral hominoid genome from the three-way alignment. Human- and chimpanzee-specific evolutionary changes, listed in Table 1, were defined by differences in each species compared with the ancestral hominoid.
Previous comparisons of substitution rates per year in neutral sites between human, rat and mouse14 revealed a lower rate in the lineage from the human–rodent ancestor to human (1.73 × 10-9) compared with the rate from the human–rodent ancestor to rat (4.95 × 10-9) and mouse (4.84 × 10-9). Other findings indicate hominoid substitution rates are lower than the rates in Old World monkeys and in other eutherian mammals16,17,18. Our analysis reveals further slow down in the branch from the hominoid ancestor to human (9.70 × 10-10) compared with chimpanzee (1.19 × 10-9). (See note added in proof.)
The pattern of insertion–deletions (indels) in human chromosome 12 relative to the hominoid ancestor is summarized in Table 2. Microsatellite expansion events, predominantly A/T mononucleotide and CA dinucleotide expansions, comprise 19% of insertion events in the 1–100-bp range. Trinucleotide expansions are more common in coding (1.62 per million base pairs (Mbp)) than noncoding regions (0.727 per Mbp). Comparisons of insertions in the 101–8,000-bp range to the “Retroposed Genes” track of the University of California at Santa Cruz (UCSC) Genome Browser (http://genome.ucsc.edu/; ref. 19) revealed 12 human-specific pseudogenes. Comparisons to known segmental duplications20 revealed nine human-specific duplications. Human-specific retrotransposon insertions in the 101–8,000-bp size range are classified by type in Table 3. Consistent with previous results21, SINE insertion rates in the human lineage are twofold higher than in the chimpanzee lineage. All retrotransposon insertions were intergenic or intronic, except for an AluY insertion in the 3′ untranslated region (UTR) of RAB21, a member of the RAS oncogene family. The overall ratio of inserted to deleted base pairs was 1.67, lengthening the human chromosome by 2.69% relative to the ancestral hominoid and consistent with previously detected evolutionary increases in the size of the human genome22.
Chromosome 12 is rich in disease-associated loci, and a total of 487 disease genes have been assigned to this chromosome2, accounting for 5.2% of all disease genes. In Supplementary Tables 4 and 5, we present the most cited genes on chromosome 12 together with the medically relevant genes sorted by function, respectively.
With regard to disease, chromosome 12 is best known for its links with cancer. Three genes mapping to the chromosome have been associated with cancer-related chromosome translocations. The most prominent of these is the ETV6 (TEL1) gene at 12p13, encoding an ETS-like transcription factor, which has a central role in haematopoietic malignancies including acute lymphoblastic leukaemia (ALL), acute myeloblastic leukaemia (AML) and myelodysplastic syndromes (MDS). Furthermore, 5% of children with ALL have 12p13–p12 deletions23. The other two genes on this chromosome involved in oncogenic gene fusions are DDIT3 (or CHOP; 12q13.1), often rearranged in myxoid liposarcoma, and HMGA2 (12q15), which has been found fused to various genes in lipoma, salivary adenoma, uterine leiomyoma and multiple lipomatosis. In addition, several genes mapped to this chromosome have been directly or indirectly associated with cancer, including BCL7A (12q24.13) in B-cell non-Hodgkin lymphoma, P2RX7 (12q24) in susceptibility to chronic lymphatic leukaemia (CLL), YEATS4 (12q13–q15) in glioma, CDK4 (12q14) in melanoma, ACVR1B (12q13) in pancreatic cancer, and KRAS (12p12.1) in colorectal adenoma.
There are 16 genes associated with movement disorders mapped to chromosome 12, as well as two diabetes-related genes, genes associated with four separate mood disorders, and four genes involved in heart disease. Of the 28 members of the keratin gene family mapped to this chromosome, 12 have been linked to skin and hair disorders. Chromosome 12 also harbours the human CD4 locus, which encodes the main receptor for the human immunodeficiency virus, at 12p3.1. This locus is within one of the most dense gene clusters in the genome, and has been the focus of many functional and population genetic studies24.
A primary goal in generating a high quality, finished reference sequence for chromosome 12 is to provide the research community with the resources to accelerate the search for additional disease-causing genes. As an example, Li and co-workers25 were recently able to demonstrate an association between sequence variants of the GAPDH gene on chromosome 12 and late-onset Alzheimer's disease (LOAD).
The finished sequence of chromosome 12 reported here marks the beginning of more extensive studies aimed at understanding our evolutionary history, together with the variation in genome structure and sequence that defines us as individuals.
Note added in proof: Recent data31 showed that of the nine human chromosomes assayed, chromosome 12 demonstrated the greatest slow down in single nucleotide substitution rate.
Methods
Mapping and sequencing
Gap closure, sequencing and finishing strategies, and clone integrity assays, are described in Supplementary Methods. BAC clone overlaps were verified by BLAST (http://www.ncbi.nlm.nih.gov/BLAST/) in-house using a locally installed copy of the software, and polymorphic regions within overlaps were confirmed by polymerase chain reaction (PCR) using a bi-gender, multi-ethnic pool of genomic DNA isolated from eight individuals (J. Belmont, personal communication). Coverage, integrity and clone order were analysed using available genetic and radiation hybrid map markers (see Supplementary Methods and Supplementary Fig. 2). Additionally, we aligned the unique paired-end fosmid reads to the finished chromosome sequence, and concluded that the limited set of clustered size discrepancies reflected probable polymorphisms between clone libraries. The unique paired fosmid end-sequence (Broad Institute) analysis was performed using sequences downloaded from the UCSC Genome Browser that were checked for both paired end-sequence orientation and resulting insert size.
Annotation
Manual curation identified each known gene, novel gene and novel transcript locus, defined as a set of one or more transcripts that share at least one exon of coding sequence (in frame) and supported by full-length and partial human or vertebrate cDNA sequences having a best-in-genome BLAST or blat (http://genome.ucsc.edu/cgi-bin/hgBlat) hit (≥ 98% identity). All analyses were performed in-house using locally installed copies of the software. The cDNA/RefSeq sequences were compared to the genomic sequence to place exons. All intron/exon splice sites for all predicted exons were examined for canonical splice motifs. Coding regions were examined for a best-fit open reading frame (ORF). The 5′ and 3′ UTRs were annotated and extended using available EST and cDNA evidence, and poly(A) sites and poly(A) signals were annotated on each gene where identified. Alternative splice variants were identified from cDNA, EST and protein evidence. The translation product for each coding sequence was verified using Swissprot. Pseudogenes were defined as sequences with no direct evidence for expression but that match with a high score to a spliced messenger RNA or spliced EST from elsewhere in the genome. This is a more stringent definition than has been applied by others in broad genomic screens of pseudogenes, and results in a fivefold-lower count across chromosome 12 than previously reported26. For paralogue analysis, protein sequences corresponding to the ‘known genes’ track of the UCSC Genome Browser27 were compared in an all-against-all BLAST search. Two loci were defined as paralogues if there was a match of any of their transcript variants with the following criteria: expect value cutoff of 10-10 or less, the lengths of the matching transcripts were within 20% of each other, and the match length extended over 70% of the average length of the two sequences. The complete set of annotations has been submitted to the Vega database (http://vega.sanger.ac.uk/Homo_Sapiens/).
Landscape features
For CpG analysis, a CpG island has been defined as an expanse of >200 nucleotides in which the G + C content is greater than 50% and the ratio of observed CG dinucleotides to expected in the segment is >0.6. We developed databases of known microRNAs, small nuclear (sn) RNAs and small nucleolar (sno) RNAs, and used tRNAscan-SE v.1.23 (http://www.genetics.wustl.edu/eddy/tRNAscan-SE; installed locally) and BLAST to search for noncoding RNA sequences. We identified recent intra- and interchromosomal segmental duplications by BLAST searching the repeat-masked chromosome sequence against itself and the rest of the human genome. The duplication densities were calculated by averaging the duplications of each base over nonoverlapping 100-kb windows after filtering low identity matches (< 90%). The densities of SINEs, LINEs and long terminal repeats (LTRs) were calculated from repeat-masked data using 100-kb windows. The G + C density was calculated by counting the G + C content over nonoverlapping 100-kb windows. The densities of CpG islands (UCSC), genes (Baylor College of Medicine Human Genome Sequencing Center annotations), and pseudogenes (as defined above) were counted and displayed using 1-Mb windows. Markers were placed on the genomic sequence using a combination of locally installed e-PCR (http://www.ncbi.nlm.nih.gov/sutils/e-pcr/) and BLAST software packages.
Comparative analysis
The multiple alignments of human, chimp (panTro1), dog (canFam1), mouse (mm5), rat (rn3), chicken (galGal2), zebrafish (danRer1) and fugu (fr1) were downloaded from the UCSC Genome Browser (http://hgdownload.cse.ucsc.edu/; May 2004/hg17-Build35). The pairwise synteny blocks between human and other species were parsed out with Synteny-Parser (X. Song & G. Weinstock, unpublished perl script), which was tuned to include all visible chromosome rearrangements in the dot plot. Rhesus scaffolds from the Mmul_0.1 preliminary assembly were mapped to human using Pash28. Rhesus scaffolds that mapped to human chromosome 12 by both Pash and the human–rhesus Alignment Net (UCSC) were aligned with orthologous human regions and chimpanzee regions from the Human–Chimpanzee Reciprocal-Best Chain alignments (UCSC) using MLAGAN29.
References
Pruitt, K. D., Tatusova, T. & Maglott, D. R. NCBI Reference Sequence (RefSeq): A curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 33, D501–D504 (2005)
Online Mendelian Inheritance in Man (OMIM). McKusick-Nathans Institute for Genetic Medicine, Johns Hopkins University (Baltimore, Maryland) and National Center for Biotechnology Information, National Library of Medicine (Bethesda, Maryland) http://www.ncbi.nlm.nih.gov/omim/ (2000).
Yu, F. et al. Positive selection of a pre-expansion CAG repeat of the human SCA2 gene. PLoS Genet. 1, e41 (2005)
Montgomery, K. T. et al. A high-resolution map of human chromosome 12. Nature 409, 945–946 (2001)
McPherson, J. D. et al. A physical map of the human genome. Nature 409, 934–941 (2001)
Schmutz, J. et al. Quality assessment of the human genome sequence. Nature 429, 365–368 (2004)
Knight, S. J. et al. An optimized set of human telomere clones for studying telomere integrity and architecture. Am. J. Hum. Genet. 67, 320–332 (2000)
Vermeesch, J. R. et al. A physical map of the chromosome 12 centromere. Cytogenet. Genome Res. 103, 63–73 (2003)
Curwen, V. et al. The Ensembl automatic gene annotation system. Genome Res. 14, 942–950 (2004)
Yu, A. et al. Comparison of human genetic and sequence-based physical maps. Nature 409, 951–953 (2001)
Wienberg, J. Fluorescence in situ hybridization to chromosomes as a tool to understand human and primate genome evolution. Cytogenet. Genome Res. 108, 139–160 (2005)
Nickerson, E. & Nelson, D. L. Molecular definition of pericentric inversion breakpoints occurring during the evolution of humans and chimpanzees. Genomics 50, 368–372 (1998)
Kehrer-Sawatzki, H., Sandig, C. A., Goidts, V. & Hameister, H. Breakpoint analysis of the pericentric inversion between chimpanzee chromosome 10 and the homologous chromosome 12 in humans. Cytogenet. Genome Res. 108, 91–97 (2005)
Rat Genome Sequencing Project Consortium. Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428, 493–521 (2004)
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002)
Li, W. H. & Tanimura, M. The molecular clock runs more slowly in man than in apes and monkeys. Nature 326, 93–96 (1987)
Steiper, M. E., Young, N. M. & Sukarna, T. Y. Genomic data support the hominoid slowdown and an Early Oligocene estimate for the hominoid–cercopithecoid divergence. Proc. Natl Acad. Sci. USA 101, 17021–17026 (2004)
Yi, S., Ellsworth, D. L. & Li, W. H. Slow molecular clocks in Old World monkeys, apes, and humans. Mol. Biol. Evol. 19, 2191–2198 (2002)
Kent, W. J., Baertsch, R., Hinrichs, A., Miller, W. & Haussler, D. Evolution's cauldron: Duplication, deletion, and rearrangement in the mouse and human genomes. Proc. Natl Acad. Sci. USA 100, 11484–11489 (2003)
Bailey, J. A., Yavor, A. M., Massa, H. F., Trask, B. J. & Eichler, E. E. Segmental duplications: Organization and impact within the current human genome project assembly. Genome Res. 11, 1005–1017 (2001)
Hedges, D. J. et al. Differential Alu mobilization and polymorphism among the human and chimpanzee lineages. Genome Res. 14, 1068–1075 (2004)
Liu, G. et al. Analysis of primate genomic variation reveals a repeat-driven expansion of the human genome. Genome Res. 13, 358–368 (2003)
Stegmaier, K. et al. Frequent loss of heterozygosity at the TEL gene locus in acute lymphoblastic leukemia of childhood. Blood 86, 38–44 (1995)
Ansari-Lari, M. A. et al. A gene-rich cluster between the CD4 and triosephosphate isomerase genes at human chromosome 12p13. Genome Res. 6, 314–326 (1996)
Li, Y. et al. Association of late-onset Alzheimer's disease with genetic variation in multiple members of the GAPD gene family. Proc. Natl Acad. Sci. USA 101, 15688–15693 (2004)
Zhang, Z., Harrison, P. M., Liu, Y. & Gerstein, M. Millions of years of evolution preserved: A comprehensive catalog of the processed pseudogenes in the human genome. Genome Res. 13, 2541–2558 (2003)
Hsu, F. et al. The UCSC Proteome Browser. Nucleic Acids Res. 33 (suppl. 1), D454–D458 (2005)
Kalafus, K. J., Jackson, A. R. & Milosavljevic, A. Pash: Efficient genome-scale sequence anchoring by Positional Hashing. Genome Res. 14, 672–678 (2004)
Brudno, M. et al. LAGAN and Multi-LAGAN: Efficient tools for large-scale multiple alignment of genomic DNA. Genome Res. 13, 721–731 (2003)
Bourque, G., Zdobnov, E. M., Bork, P., Pevzner, P. A. & Tesler, G. Comparative architectures of mammalian and chicken genomes reveal highly variable rates of genomic rearrangements across different lineages. Genome Res. 15, 98–110 (2005)
Elango, N., Thomas, J. W., NISC Comparative Sequencing Program & Soojin, V. Y. Variable molecular clocks in hominoids. Proc. Natl Acad. Sci. USA 103, 1370–1375 (2006)
Acknowledgements
This work was supported by NIH grants to R.K. and to R.A.G. We acknowledge and thank the genome sequencing community for generating the data sets used in our comparative analysis. We also acknowledge the following members of the HUGO Gene Nomenclature Committee: S. Povey (chair), E. A. Bruford, V. K. Khodiyar, M. J. Lush, K. M. B. Sneddon, T. P. Sneddon, C. C. Talbot Jr and M. W. Wright.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The chromosome 12 sequence has been deposited into GenBank under the accession number NC_000012. Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Additional information
*A list of participants and affiliations appears at the end of the paper
Supplementary information
Supplementary Figure 1
Plot of deCODE recombination distance in males, females and sex-averaged compared to the physical sequence of chromosome 12 from ptel to q-tel. (PDF 45 kb)
Supplementary Figure 2
Plot of genetic marker order using markers from five maps across chromosome 12 from p-tel to q-tel. (PDF 94 kb)
Supplementary Figure Legends
Legends describing Supplementary Figures 1 and 2. (PDF 79 kb)
Supplementary Table 1
Table of euchromatic gap sizes Supplementary Table 2 (pdf; 48 kb): Table of non-coding RNA loci across the chromosome from p-tel to q-tel. (PDF 55 kb)
Supplementary Table 2
Chromosome 12 non-coding RNA genes (PDF 65 kb)
Supplementary Table 3
G+C composition and rate of change between human and rodent. (PDF 50 kb)
Supplementary Table 4
Chromosome 12 disease loci listed by number of PubMed references. (PDF 58 kb)
Supplementary Table 5
Chromosome 12 disease loci listed by functional category. (PDF 61 kb)
Supplementary Methods
Additional methods to what is presented in the text together with their supplementary references. (PDF 81 kb)
Rights and permissions
About this article
Cite this article
Scherer, S., Muzny, D., Buhay, C. et al. The finished DNA sequence of human chromosome 12. Nature 440, 346–351 (2006). https://doi.org/10.1038/nature04569
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04569
This article is cited by
-
A Review of Functional Characterization of Single Amino Acid Change Mutations in HNF Transcription Factors in MODY Pathogenesis
The Protein Journal (2021)
-
Involvement of the P2X7-NLRP3 axis in leukemic cell proliferation and death
Scientific Reports (2016)
-
In-depth LC-MS/MS analysis of the chicken ovarian cancer proteome reveals conserved and novel differentially regulated proteins in humans
Analytical and Bioanalytical Chemistry (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.