Abstract
Mutations in ClC-7, a late endosomal/lysosomal member of the CLC family of chloride channels and transporters1,2, cause osteopetrosis3 and lysosomal storage disease4 in humans and mice. Severe osteopetrosis is also observed with mutations in the OSTM1 gene, which encodes a membrane protein of unknown function5. Here we show that both ClC-7 and Ostm1 proteins co-localize in late endosomes and lysosomes of various tissues, as well as in the ruffled border of bone-resorbing osteoclasts. Co-immunoprecipitations show that ClC-7 and Ostm1 form a molecular complex and suggest that Ostm1 is a β–subunit of ClC-7. ClC-7 is required for Ostm1 to reach lysosomes, where the highly glycosylated Ostm1 luminal domain is cleaved. Protein but not RNA levels of ClC-7 are greatly reduced in grey-lethal mice, which lack Ostm1, suggesting that the ClC-7–Ostm1 interaction is important for protein stability. As ClC-7 protein levels in Ostm1-deficient tissues and cells, including osteoclasts, are decreased below 10% of normal levels, Ostm1 mutations probably cause osteopetrosis by impairing the acidification of the osteoclast resorption lacuna, which depends on ClC-7 (ref. 3). The finding that grey-lethal mice, just like ClC-7-deficient mice4, show lysosomal storage and neurodegeneration in addition to osteopetrosis implies a more general importance for ClC-7–Ostm1 complexes.
Similar content being viewed by others
Main
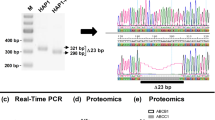
In western blots of brain membranes, an antibody against the carboxy terminus of mouse Ostm1 recognized an ∼80 kDa band and a doublet at ∼35–45 kDa (Fig. 1c). These bands were absent in grey-lethal (gl) mice, an osteopetrotic mouse mutant6 carrying a deletion of the Ostm1 promoter and exon 1 (ref. 5). Incubating cells with protease inhibitors increased the proportion of the large band at the expense of the smaller ones (Fig. 1d), indicating that the small forms of Ostm1 are produced by proteolytic cleavage of the ∼80 kDa protein. The apparent sizes of the Ostm1 species suggest cleavage roughly in the middle of the protein (Fig. 1b). As the large and the small forms (Fig. 1c) ran together in a single large band under non-reducing conditions (Fig. 1e), the cleaved fragments might be linked by disulphide bonds between some of the cysteine residues that abound in the luminal domain of Ostm1 (Fig. 1a).
a, Schematic of Ostm1 protein. Black box, hydrophobic stretch; dashed box, proposed RING-finger domain7; Y, consensus site for N-linked glycosylation. Asterisks indicate cysteine residues; arrow indicates predicted signal peptide cleavage site; AB indicates antibody binding site. Lines below show the truncated proteins predicted from human OSTM1 mutations5,20. b, Topology model derived from our work. Arrow shows the approximate cleavage site in lysosomal Ostm1. c–f, Western blots of Ostm1. WT, wild type. c, The band at ∼80 kDa (filled arrowhead) and the doublet at ∼35–45 kDa detected in wild-type brain membranes (open arrowhead) were absent from the grey-lethal brain. Asterisk indicates a non-specific band. d, Incubating fibroblasts with the protease inhibitor leupeptin increased the proportion of the large Ostm1 species. Similar results were obtained with E64, which inhibits several cathepsins (not shown). e, Western blots of brain proteins separated by non-reducing SDS–PAGE showed a single band of ∼80 kDa Ostm1. f, Deglycosylation with PNGaseF reduced the sizes of all Ostm1 species detected under reducing conditions. g, Western blot of cells (lanes 1 and 2) and supernatants (sup., lanes 3 and 4) from HEK293 cells expressing Ostm1 (lanes 1 and 3) or a truncated form of Ostm1 (lanes 2 and 4) that mimics a human mutation5. Both proteins carried a C-terminal HA-epitope.
Recent work has proposed an E3 ubiquitin ligase function for Ostm1 (cloned independently from rat as GIPN)7. The stretch between the amino- and carboxy-terminal hydrophobic regions of Ostm1 (Fig. 1a) shows weak homology to RING-finger proteins and was suggested to be cytosolic7. However, this stretch (and no other part of the protein) contains several consensus sites for N-linked glycosylation. Several or all of these sites are used, because deglycosylation with PNGaseF (peptide N-glycosidase F) greatly reduced the apparent size of all Ostm1 species (Fig. 1f). The observed glycosylation places the hypothetical RING-finger domain7 in the lumen of the endoplasmic reticulum (ER), a localization difficult to reconcile with the cytosolic/nucleoplasmic activity of ubiquitin ligases8. The disappearance in transfected cells of a haemagglutinin (HA)-epitope added to the N terminus indicates the presence of a cleavable signal peptide (data not shown). We also investigated an Ostm1 mutant truncated before the second hydrophobic stretch. This mutant, but not wild-type Ostm1, was secreted into the supernatant of transfected cells (Fig. 1g). Hence, the first and second hydrophobic domains serve as a cleavable signal peptide and transmembrane domain, respectively, in agreement with the type I transmembrane protein model proposed in ref. 5 (Fig. 1b). Figure 1f also showed that the apparent molecular mass of the largest deglycosylated band agreed roughly with the prediction from the Ostm1 reading frame (∼37 kDa). As deglycosylation of the small Ostm1 species yielded a single band, the doublet is attributable to non-uniform glycosylation.
In cultured primary fibroblasts, Ostm1 co-localized with Lamp-1, a marker for late endosomes and lysosomes (Fig. 2a). This localization resembled that of ClC-7, the loss of which also causes osteopetrosis3. However, when fibroblasts were transiently transfected with Ostm1 (Fig. 2b), it showed an ER-like distribution and no significant co-localization with Lamp-1 was observed. Co-transfection with ClC-7 restored a punctate Ostm1 staining pattern that largely co-stained with Lamp-1 (Fig. 2c) and ClC-7 (Fig. 2d). The effect of ClC-7 was specific, as co-transfection with neither ClC-3 (not shown) nor ClC-6 (Supplementary Fig. S1), which are both expressed in the endosomal/lysosomal pathway2,9, resulted in such changes.
a, Mouse fibroblast stained for Ostm1 and Lamp-1. Right panels show the overlay, insets show higher magnification. b, Ostm1-transfected fibroblasts showed reticular and perinuclear Ostm1 staining that co-localized poorly with Lamp-1. c, d, In fibroblasts co-transfected with Ostm1 and ClC-7, Ostm1 co-localized with Lamp-1 (c) and ClC-7 (d). In b–d, Clcn7-/- fibroblasts were used to avoid effects of endogenous ClC-7, but similar results were seen in HeLa cells. Scale bar in a represents 8.5 µm (a), 10 µm (b–d), 1.7 µm (insets).
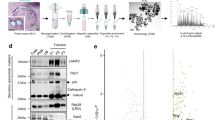
As ClC-7 changed Ostm1 localization, we studied Ostm1 in ClC-7-knockout (Clcn7-/-) mice. Western blots indicated exclusive loss of the 35–45 kDa Ostm1 doublet in brain tissue from Clcn7-/- mice (Fig. 3a). Subcellular fractionation of wild-type brain tissue revealed that the small Ostm1 form was co-enriched with lysosomal markers (bottom fractions 1–2), whereas the 80 kDa form was only detectable in fractions 9–12, which contain markers for endosomes and the ER (Fig. 3b). Such experiments yielded samples containing only the small (wild-type fractions containing lysosomes) or large (Clcn7-/- fractions containing endosomes/ER) forms of Ostm1, which we used for deglycosylation experiments. PNGaseF reduced the size of both species (Fig. 3c, lanes 2 and 4), and only the 35–45 kDa form was partially resistant to EndoH (endoglycosidase H) (Fig. 3c, compare lanes 2 and 6). This indicates that the small, but not the large species of Ostm1 had left the ER. Our results thus suggest that ClC-7 is required for trafficking Ostm1 from the ER to lysosomes, and that the luminal domain of Ostm1 is cleaved on its way to, or in, this final compartment.
a, Western blot of Ostm1 in brain from wild-type, Clcn7-/- and gl mice. b, Subcellular distribution of Ostm1 in wild-type and Clcn7-/- mice. Brain membranes (P2) and Percoll gradient fractions were analysed. Bottom fractions (fr. 1–2) contained lysosomal markers (cathepsin D, ClC-7), top fractions (9–12) contained ER (calnexin, PDI) and endosomal (Rab4) markers. Small forms of Ostm1 were enriched in lysosomal fractions, and the large form in ER/endosomal fractions. c, Deglycosylation of wild-type lysosomal and Clcn7-/- ER/endosomal fractions, using EndoH or PNGaseF. d, Co-immunoprecipitation reveals a ClC-7–Ostm1 complex. Solubilized brain membranes were directly loaded on the gel (input, lanes 1–3), or first immunoprecipitated (IP) with ClC-7 (lanes 4, 5) or Ostm1 antibodies (lanes 6, 7). Western blots were probed for the proteins indicated on the right. Arrowheads show specific Ostm1 bands; asterisk shows non-specific bands. Equivalent amounts of lysates and precipitates were loaded.
ClC-7 was efficiently co-immunoprecipitated from the brain with Ostm1, and vice versa (Fig. 3d). This interaction was specific, as it was observed neither with the related endosomal proteins ClC-3 and ClC-6, nor with Lamp-2. As expected from the lysosomal localization of ClC-7 (ref. 4), antibodies against ClC-7 almost exclusively precipitated the cleaved, lysosomal Ostm1 fragment from brain. Co-immunoprecipitation performed with transfected cells in which only the large form of Ostm1 could be detected showed that this putative ER form also interacted with ClC-7 (Supplementary Fig. S2). Notably, Fig. 3d also shows that ClC-7 levels are greatly reduced in brain extracts from gl mice (lane 3).
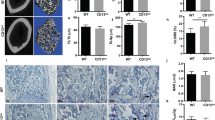
Both ClC-7 and Ostm1 are expressed in several tissues, including brain, liver, kidney and osteoclasts3,5,7,10 (see Supplementary Fig. S3). Immunohistochemistry of brain sections revealed that both proteins co-localize in neuronal cell bodies in structures that most likely represent lysosomes4 (Fig. 4a). Both proteins also co-localize in osteoclasts in a pattern that represents the ‘ruffled border’ (Fig. 4c). This acid-secreting plasma membrane domain was identified by co-staining for the a3 proton pump subunit11,12 (Fig. 4d), mutations of which also underlie osteopetrosis11,13,14,15. Consistent with our western blot analyses (Fig. 3a, b, d), Ostm1 staining was very weak in Clcn7-/- mice (Supplementary Fig. S4), and ClC-7 labelling was greatly reduced in grey-lethal cells. These cells included neurons (Fig. 4b) and osteoclasts (Fig. 4e), in which the remaining ClC-7 still localized to the ruffled border. Western blot analysis of brain, kidney, liver and bone revealed that ClC-7 levels were reduced to less than 10% in gl compared to wild-type mice (Fig. 4f, g). The transcript levels of both Clcn7 and Ostm1 genes were unchanged (Supplementary Table).
a, b, Immunofluorescence of cerebellar Purkinje cells. a, Ostm1 co-localized with ClC-7 in late endosomes/lysosomes4 of wild-type cells. b, In gl neurons, both proteins were undetectable. c–e, Immunofluorescence of osteoclasts in situ. Ostm1 and ClC-7 co-localized in the ruffled border (c), as did ClC-7 and the a3 proton-pump subunit (d). e, In gl osteoclasts, ClC-7 levels were greatly reduced. In a–e, nucleic acids were counterstained using TOTO (blue). Scale bar, 10 µm (a–e). f, Western blots show a decrease in ClC-7, but not in ClC-3, ClC-6, Lamp-1 or Lamp-2 in gl brain. g, Quantification of western blots from various tissues shows a decrease in ClC-7 in gl mice at postnatal day (P)11–33, but no decrease in other late endosomal/lysosomal proteins. The moderate decrease in control proteins in gl bone might be explained by the osteopetrosis. Error bars indicate s.e.m.; n = 3–9 mouse pairs, except for ClC-3 in bone (n = 2). h, i, Retinal degeneration is visible in gl (h) but not in wild-type (i) mice at P31. j, NeuN (neuronal nuclear antigen) staining revealed neurodegeneration in the CA3 region (arrows) of a gl hippocampus at P47. Scale bar, 100 µm.
ClC-7 might support bone resorption by electrically shunting the H+-ATPase that acidifies the osteoclast resorption lacuna3, or similarly by facilitating the insertion of proton-pump-containing vesicles into the ruffled border, which is underdeveloped in both Clcn7-/- and gl osteoclasts3,6. This mechanism would also be feasible if ClC-7 were not a Cl- channel, as believed so far3,10, but an electrogenic Cl-/H+-exchanger such as ClC-ec1 (ref. 16), ClC-4 or ClC-5 (refs 17, 18). As the intracellular localization of ClC-7 precluded biophysical analysis, these alternatives remain untested. Our work suggests that loss of OSTM1 causes osteopetrosis5,19,20 by decreasing the amount of ClC-7 to pathogenic levels. A 75% decrease in ClC-7 function as a result of dominant-negative mutations causes mild osteopetrosis in humans21,22. The even lower levels of ClC-7 in gl mice may be sufficient to cause the severe osteopetrosis observed with OSTM1 mutations5,19,20.
Known disease-causing mutations in the human OSTM1 gene5,20 introduce frame-shifts that replace the OSTM1 polypeptide with short unrelated sequences 143 or 21 residues5,20 before the transmembrane domain. When an epitope-tagged construct modelled after the latter mutation5 (Fig. 1a) was transfected into cells, co-expression with ClC-7 failed to direct the truncated Ostm1 to lysosomes (data not shown) and the truncated protein was secreted into the supernatant (Fig. 1g). We suggest that this mutant may lead to disease because Ostm1 no longer interacts with ClC-7.
The phenotypes of Clcn7-/- and gl mice are strikingly similar. On an agouti background, the fur of Clcn7-/- mice is grey (data not shown), just like the coat colour of grey-lethal mice5,23. The disruption of ClC-7 leads not only to osteopetrosis3, but also to retinal3 and central nervous system4 degeneration, which are related to lysosomal storage disease4. We detected similar retinal and hippocampal degeneration in grey-lethal mice (Fig. 4h–j). Like Clcn7-/- mice4, they showed electron-dense storage material in neurons and renal proximal tubular cells (Supplementary Fig. S5). Again similar to Clcn7-/- mice4, the pH of gl lysosomes seemed normal (Supplementary Fig. S6), possibly pointing to altered acidification during lysosome formation, or to a role for lysosomal chloride4,18. Taken together, we suggest that the ClC-7–Ostm1 complex is also important for the function of melanocytes and lysosomes. Patients with OSTM1 mutations may develop lysosomal storage disease in addition to osteopetrosis.
Our work has identified Ostm1 as a hitherto unknown ancillary β-subunit of ClC-7. Ostm1 requires ClC-7 to travel to lysosomes, whereas ClC-7 can reach its destination without Ostm1. The stability of ClC-7 depends on its association with Ostm1. As pronounced glycosylation is thought to protect lysosomal membrane proteins from degradation24,25, one might speculate that the highly glycosylated Ostm1 protein shields ClC-7, the only mammalian CLC protein lacking N-linked glycosylation sites, from lysosomal proteases. The osteopetrosis, lysosomal storage and neurodegeneration observed upon loss of Ostm1 may be entirely explained by a large reduction in ClC-7 protein levels.
Methods
Please refer to Supplementary Methods for full details.
Mice
Grey-lethal mice5,26 obtained from Jackson Laboratory as double-heterozygous GL/Le Edardl-J + / + Ostm1gl/J mice were crossed once to C57BL/6 mice and further bred to isolate the gl allele and produce homozygous gl/gl mice. Details of Clcn7-/- mice have been published3.
Antibodies
Antibodies against the a3 subunit of the V-type H+-ATPase (peptide PDASTLENSWSPDEEK) or Ostm1 (LKSSTSFANIQENAT) were raised in guinea pigs and rabbits, affinity purified and tested for specificity on oc and gl knockout tissue, respectively5,11. Antibodies against ClC-3 and ClC-7 have been described3,9. See Supplementary Information for details of other antibodies.
DNA constructs
The Ostm1 cDNA was amplified by polymerase chain reaction with reverse transcription (RT–PCR) from mouse kidney, and cloned into an eukaryotic expression vector. A mutant mimicking the OSTM1 fragment remaining in an osteopetrosis patient5 was generated by replacing the sequence encoding Ostm1 amino acids 266–338 with an HA-epitope (resulting sequence VEDA-VD-YPYDVPDYA-stop).
Cell culture
Fibroblasts from wild-type or Clcn7-/- mice, HEK293 or HeLa cells were prepared and cultured as described4. For protease inhibition, fibroblasts were cultured for 24 h in the presence of 20 µM leupeptin (Roche) or 10 µM E64 (Sigma).
Immunohistochemistry and histology
Immunostaining of cryo- and paraffin sections and histology were done as described3,4.
Biochemical methods
Cellular membranes from mouse brain were fractionated by sedimentation velocity centrifugation as detailed in the Supplementary Methods. Membrane preparations or gradient fractions were deglycosylated using PNGaseF or EndoH (Roche). For immunoprecipitation experiments, Ostm1 or ClC-7 antibodies were crosslinked to protein A sepharose by dimethylpimelimidate. Brain membranes were pelleted and solubilized in lysis buffer containing 1% Triton X-100. Non-solubilized material was removed by a 70,000g spin. After incubation with protein A sepharose–antibody complexes for 2 h at 4 °C and washing, samples were eluted at pH 2.8, neutralized, and denatured in SDS sample buffer.
References
Jentsch, T. J., Neagoe, I. & Scheel, O. CLC chloride channels and transporters. Curr. Opin. Neurobiol. 15, 319–325 (2005)
Jentsch, T. J., Poët, M., Fuhrmann, J. C. & Zdebik, A. A. Physiological functions of CLC Cl- channels gleaned from human genetic disease and mouse models. Annu. Rev. Physiol. 67, 779–807 (2005)
Kornak, U. et al. Loss of the ClC-7 chloride channel leads to osteopetrosis in mice and man. Cell 104, 205–215 (2001)
Kasper, D. et al. Loss of the chloride channel ClC-7 leads to lysosomal storage disease and neurodegeneration. EMBO J. 24, 1079–1091 (2005)
Chalhoub, N. et al. Grey-lethal mutation induces severe malignant autosomal recessive osteopetrosis in mouse and human. Nature Med. 9, 399–406 (2003)
Rajapurohitam, V. et al. The mouse osteopetrotic grey-lethal mutation induces a defect in osteoclast maturation/function. Bone 28, 513–523 (2001)
Fischer, T., De Vries, L., Meerloo, T. & Farquhar, M. G. Promotion of Gαi3 subunit down-regulation by GIPN, a putative E3 ubiquitin ligase that interacts with RGS-GAIP. Proc. Natl Acad. Sci. USA 100, 8270–8275 (2003)
Pickart, C. M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001)
Stobrawa, S. M. et al. Disruption of ClC-3, a chloride channel expressed on synaptic vesicles, leads to a loss of the hippocampus. Neuron 29, 185–196 (2001)
Brandt, S. & Jentsch, T. J. ClC-6 and ClC-7 are two novel broadly expressed members of the CLC chloride channel family. FEBS Lett. 377, 15–20 (1995)
Scimeca, J. C. et al. The gene encoding the mouse homologue of the human osteoclast-specific 116-kDa V-ATPase subunit bears a deletion in osteosclerotic (oc/oc) mutants. Bone 26, 207–213 (2000)
Nishi, T. & Forgac, M. Molecular cloning and expression of three isoforms of the 100-kDa a subunit of the mouse vacuolar proton-translocating ATPase. J. Biol. Chem. 275, 6824–6830 (2000)
Li, Y. P., Chen, W., Liang, Y., Li, E. & Stashenko, P. Atp6i-deficient mice exhibit severe osteopetrosis due to loss of osteoclast-mediated extracellular acidification. Nature Genet. 23, 447–451 (1999)
Frattini, A. et al. Defects in TCIRG1 subunit of the vacuolar proton pump are responsible for a subset of human autosomal recessive osteopetrosis. Nature Genet. 25, 343–346 (2000)
Kornak, U. et al. Mutations in the a3 subunit of the vacuolar H+-ATPase cause infantile malignant osteopetrosis. Hum. Mol. Genet. 9, 2059–2063 (2000)
Accardi, A. & Miller, C. Secondary active transport mediated by a prokaryotic homologue of ClC Cl- channels. Nature 427, 803–807 (2004)
Picollo, A. & Pusch, M. Chloride/proton antiporter activity of mammalian CLC proteins ClC-4 and ClC-5. Nature 436, 420–423 (2005)
Scheel, O., Zdebik, A., Lourdel, S. & Jentsch, T. J. Voltage-dependent electrogenic chloride proton exchange by endosomal CLC proteins. Nature 436, 424–427 (2005)
Quarello, P. et al. Severe malignant osteopetrosis caused by a GL gene mutation. J. Bone Miner. Res. 19, 1194–1199 (2004)
Ramírez, A. et al. Identification of a novel mutation in the coding region of the grey-lethal gene OSTM1 in human malignant infantile osteopetrosis. Hum. Mutat. 23, 471–476 (2004)
Cleiren, E. et al. Albers-Schönberg disease (autosomal dominant osteopetrosis, type II) results from mutations in the ClCN7 chloride channel gene. Hum. Mol. Genet. 10, 2861–2867 (2001)
Frattini, A. et al. Chloride channel ClCN7 mutations are responsible for severe recessive, dominant, and intermediate osteopetrosis. J. Bone Miner. Res. 18, 1740–1747 (2003)
Boyce, B. F. Bad bones, grey hair, one mutation. Nature Med. 9, 395–396 (2003)
Fukuda, M. Lysosomal membrane glycoproteins. Structure, biosynthesis, and intracellular trafficking. J. Biol. Chem. 266, 21327–21330 (1991)
Eskelinen, E. L., Tanaka, Y. & Saftig, P. At the acidic edge: emerging functions for lysosomal membrane proteins. Trends Cell Biol. 13, 137–145 (2003)
Vacher, J. & Bernard, H. Genetic localization and transmission of the mouse osteopetrotic grey-lethal mutation. Mamm. Genome 10, 239–243 (1999)
Acknowledgements
We thank M. Schweizer for electron microscopy and retina morphology, S. Bauer, N. Krönke and E. Orthey for technical assistance, and R. Pohlmann for cathepsin D antiserum. This work was supported by the Deutsche Forschungsgemeinschaft, and by a fellowship from the Boehringer Ingelheim Fonds to L. Wartosch.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains Supplementary Figures 1–6, Supplementary Table 1 and Supplementary Methods. This file also contains additional references. (PDF 3034 kb)
Supplementary Data
This file contains details of the key genes and proteins used in this study. (PDF 2 kb)
Rights and permissions
About this article
Cite this article
Lange, P., Wartosch, L., Jentsch, T. et al. ClC-7 requires Ostm1 as a β-subunit to support bone resorption and lysosomal function. Nature 440, 220–223 (2006). https://doi.org/10.1038/nature04535
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04535
This article is cited by
-
A gain-of-function TPC2 variant R210C increases affinity to PI(3,5)P2 and causes lysosome acidification and hypopigmentation
Nature Communications (2023)
-
Structure and function of the membrane microdomains in osteoclasts
Bone Research (2023)
-
Proton-gated anion transport governs macropinosome shrinkage
Nature Cell Biology (2022)
-
Genetic analysis of osteopetrosis in Pakistani families identifies novel and known sequence variants
BMC Medical Genomics (2021)
-
In silico analysis of the effects of disease-associated mutations of β-hexosaminidase A in Tay‒Sachs disease
Journal of Genetics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.