Abstract
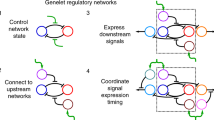
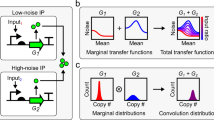
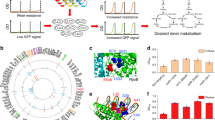
The ability to construct synthetic gene networks enables experimental investigations of deliberately simplified systems that can be compared to qualitative and quantitative models1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23. If simple, well-characterized modules can be coupled together into more complex networks with behaviour that can be predicted from that of the individual components, we may begin to build an understanding of cellular regulatory processes from the ‘bottom up’. Here we have engineered a promoter to allow simultaneous repression and activation of gene expression in Escherichia coli. We studied its behaviour in synthetic gene networks under increasingly complex conditions: unregulated, repressed, activated, and simultaneously repressed and activated. We develop a stochastic model that quantitatively captures the means and distributions of the expression from the engineered promoter of this modular system, and show that the model can be extended and used to accurately predict the in vivo behaviour of the network when it is expanded to include positive feedback. The model also reveals the counterintuitive prediction that noise in protein expression levels can increase upon arrest of cell growth and division, which we confirm experimentally. This work shows that the properties of regulatory subsystems can be used to predict the behaviour of larger, more complex regulatory networks, and that this bottom-up approach can provide insights into gene regulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000)
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000)
Becskei, A. & Serrano, L. Engineering stability in gene networks by autoregulation. Nature 405, 590–593 (2000)
Becskei, A., Séraphin, B. & Serrano, L. Positive feedback in eukaryotic gene networks: cell differentiation by graded to binary response conversion. EMBO J. 20, 2528–2535 (2001)
Elowitz, M., Levine, A., Siggia, E. & Swain, P. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002)
Ozbudak, E., Thattai, M., Kurtser, I., Grossman, A. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002)
Rosenfeld, N. Y., Elowitz, M. B. & Alon, U. Negative autoregulation speeds the response times of transcription networks. J. Mol. Biol. 323, 785–793 (2002)
Guet, C. C., Elowitz, M. B., Hsing, W. & Leibler, S. Combinatorial synthesis of genetic networks. Science 296, 1407–1470 (2002)
Isaacs, F. J., Hasty, J., Cantor, C. R. & Collins, J. J. Prediction and measurement of an autoregulatory genetic module. Proc. Natl. Acad. Sci. USA 100, 7714–7719 (2003)
Blake, W. J., Kaern, M., Cantor, C. R. & Collins, J. J. Noise in eukaryotic gene expression. Nature 422, 633–637 (2003)
Atkinson, M. R., Savageau, M. A., Myers, J. T. & Ninfa, A. J. Developement of genetic circuitry exhibiting toggle switch or oscillatory behaviour in Escherichia coli. Cell 113, 597–607 (2003)
Weiss, R. et al. Genetic circuit building blocks for cellular computation, communications, and signal processing. Natural Comput. 2, 47–84 (2003)
Basu, S., Mahreja, R., Thiberge, S., Chen, M. T. & Weiss, R. Spatiotemporal control of gene expression with pulse generating networks. Proc. Natl. Acad. Sci. 17, 6355–6360 (2004)
You, L., Cox, R. S. III, Weiss, R. & Arnold, F. H. Programmable population control cell-cell communication and regulated killing. Nature 428, 868–871 (2004)
Kramer, B. P. et al. An engineered epigenetic transgene switch in mammalian cells. Nature Biotechnol. 22, 867–870 (2004)
Kobayashi, H. et al. Programmable cells: interfacing natural and engineered gene networks. Proc. Natl. Acad. Sci. 101, 8414–8419 (2004)
Isaacs, F. J. et al. Engineered riboregulators enable post-transcriptional control of gene expression. Nature Biotechnol. 22, 841–847 (2004)
Fung, E. et al. A synthetic gene-metabolic oscillator. Nature 435, 118–122 (2005)
Hooshangi, S., Thiberge, S. & Weiss, R. Ultrasensitivity and noise propagation in a synthetic transcriptional cascade. Proc. Natl. Acad. Sci. USA 102, 3581–3586 (2005)
Pedraza, J. M. & van Oudenaarden, A. Noise propagation in gene networks. Science 307, 1965–1969 (2005)
Rosenfeld, N. Y., Young, J. W., Alon, U., Swain, P. S. & Elowitz, M. B. Gene regulation at the single cell level. Science 307, 1962–1965 (2005)
Isalan, M., Lemerle, C. & Serrano, L. Engineering gene networks to emulate Drosophila embryonic pattern formation. PLoS Biol. 3(3), e64 (2005)
Kramer, B. P. & Fussenegger, M. Hysteresis in a synthetic mammalian gene network. Proc. Natl. Acad. Sci. USA 102, 9517–9522 (2005)
Ptashne, M. A Genetic Switch: Phage λ and Higher Organisms (Cell Press & Blackwell Scientific, Cambridge, Massachusetts, 1992)
Kepler, T. & Elston, T. C. Stochasticity in transcriptional regulation: origins, consequences, and mathematical representations. Biophys. J. 81, 3116–3136 (2001)
Adalsteinsson, D., McMillen, D. & Elston, T. C. Biochemical Network Stochastic Simulator (BioNetS): software for stochastic modeling of biochemical networks. BMC Bioinform. 5, 24 (2004)
Paulsson, J. & Ehrenberg, M. Noise in a minimal regulatory network: plasmid copy number control. Q. Rev. Biophys. 34, 1–59 (2001)
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997)
Greenwood, D. & O'Grady, F. Comparison of the response of Escherichia coli and Proteus mirabilis to seven β-lactam antibiotics. J. Infect. Dis. 128, 211–222 (1973)
Sambrook, J., Fritsch, E. & Maniatis, T. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, Plainview, New York, 1989)
Acknowledgements
This work was supported by the NIH, NSF and DARPA. Author Contributions T.C.E. and J.J.C. are co-senior authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains text detailing modeling details and further experimental work omitted from the main text due to space restrictions. There are 13 Supplementary Figures and 3 Supplementary Tables. (PDF 811 kb)
Rights and permissions
About this article
Cite this article
Guido, N., Wang, X., Adalsteinsson, D. et al. A bottom-up approach to gene regulation. Nature 439, 856–860 (2006). https://doi.org/10.1038/nature04473
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04473
This article is cited by
-
Modulating gene regulation function by chemically controlled transcription factor clustering
Nature Communications (2022)
-
Enhanced regulation of prokaryotic gene expression by a eukaryotic transcriptional activator
Nature Communications (2021)
-
Bottom-up approaches in synthetic biology and biomaterials for tissue engineering applications
Journal of Industrial Microbiology and Biotechnology (2018)
-
Scaling up genetic circuit design for cellular computing: advances and prospects
Natural Computing (2018)
-
Re-using biological devices: a model-aided analysis of interconnected transcriptional cascades designed from the bottom-up
Journal of Biological Engineering (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.