Abstract
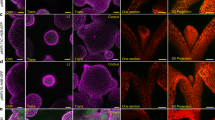
Many types of plant cell retain their developmental plasticity and have the capacity to switch fate when exposed to a new source of positional information. In the root epidermis of Arabidopsis, cells differentiate in alternating files of hair cells and non-hair cells1,2, in response to positional information and the activity of the homoeodomain transcription factor GLABRA2 (GL2) in future non-hair cells3,4,5,6. Here we show by three-dimensional fluorescence in situ hybridization on intact root epidermal tissue that alternative states of chromatin organization around the GL2 locus are required to control position-dependent cell-type specification. When, as a result of an atypical cell division, a cell is displaced from a hair file into a non-hair file, it switches fate6. We show that during this event the chromatin state around the GL2 locus is not inherited, but is reorganized in the G1 phase of the cell cycle in response to local positional information. This ability to remodel chromatin organization may provide the basis for the plasticity in plant cell fate changes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dolan, L. et al. Clonal relationships and patterning in the root epidermis of Arabidopsis. Development 120, 2465–2474 (1994)
Galway, M. E. et al. The TTG gene is required to specify epidermal cell fate and cell patterning in the Arabidopsis root. Dev. Biol. 166, 740–754 (1994)
Masucci, J. D. et al. The homeobox gene GLABRA2 is required for position-dependent cell differentiation in the root epidermis of Arabidopsis thaliana. Development 122, 1253–1260 (1996)
Di Cristina, M. et al. The Arabidopsis Athb-10 (GLABRA2) is an HD-Zip protein required for regulation of root hair development. Plant J. 10, 393–402 (1996)
Hung, C. Y. et al. A common position-dependent mechanism controls cell-type patterning and GLABRA2 regulation in the root and hypocotyl epidermis of Arabidopsis. Plant Physiol. 117, 73–84 (1998)
Berger, F., Haseloff, J., Schiefelbein, J. & Dolan, L. Positional information in the root epidermis is defined during embryogenesis and acts in domains with strict boundaries. Curr. Biol. 8, 421–430 (1998)
Lin, Y. & Schiefelbein, J. Embryonic control of epidermal cell patterning in the root and hypocotyl of Arabidopsis. Development 128, 3697–3705 (2001)
Costa, S. & Dolan, L. Epidermal patterning genes are active during embryogenesis in Arabidopsis. Development 130, 2839–2901 (2003)
Kwak, S. H., Shen, R. & Schiefelbein, J. Positional signalling mediated by a receptor-like kinase in Arabidopsis. Science 307, 1111–1113 (2005)
Lee, M. M. & Schiefelbein, J. WEREWOLF, a MYB-related protein in Arabidopsis, is a positional-dependent regulator of epidermal cell patterning. Cell 99, 473–483 (1999)
Lee, M. M. & Schiefelbein, J. Cell pattern in the Arabidopsis root epidermis determined by lateral inhibition with feedback. Plant Cell 14, 611–618 (2002)
Wada, T., Tachibana, T., Shimura, Y. & Okada, K. Epidermal cell differentiation in Arabidopsis determined by a Myb homolog, CPC. Science 277, 1113–1116 (1997)
Wada, T. et al. Role of a positive regulator of root hair development, CAPRICE, in Arabidopsis root epidermal cell differentiation. Development 129, 5409–5419 (2002)
Kaya, H. et al. FASCIATA genes for chromatin assembly factor-1 in Arabidopsis maintain the cellular organization of apical meristems. Cell 104, 131–142 (2001)
Wolpert, L. Cell boundaries: knowing who to mix with and what to shout or whisper. Development 130, 4497–4500 (2003)
Szymkowiak, E. J. & Sussex, I. M. What chimeras can tell us about plant development. Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 351–376 (1996)
Dolan, L. & Okada, K. Signalling in cell specification. Semin. Cell Dev. Biol. 10, 149–156 (1999)
Kidner, C., Sundaresan, V., Roberts, K. & Dolan, L. Clonal analysis of the Arabidopsis root confirms that position, not lineage, determines cell fate. Planta 211, 191–199 (2000)
Laux, T. The stem cell concept in plants: a matter of debate. Cell 113, 281–283 (2003)
Acknowledgements
We thank L. Dolan, G. Moore, C. Dean and M. Towers for discussion and advice. This work was funded by the BBSRC and the Gatsby Charitable Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figure 1
Schematic representation of the organisation of the Arabidopsis root epidermis. (DOC 192 kb)
Supplementary Figure 2
Close-ups of chromatin organisation at the GL2 locus in wild-type and cpc nuclei and scatter plots of FISH signal intensities measured on both. (DOC 394 kb)
Supplementary Figure 3
In wer mutant every cell of the meristematic root epidermis is in an open chromatin state in the GL2 region. (DOC 640 kb)
Supplementary Figure 4
The number of FISH signals detected indicates the cell cycle stage. (DOC 486 kb)
Supplementary Figure 5
Throughout mitosis the GL2 locus is accessible to FISH probes. (DOC 91 kb)
Rights and permissions
About this article
Cite this article
Costa, S., Shaw, P. Chromatin organization and cell fate switch respond to positional information in Arabidopsis. Nature 439, 493–496 (2006). https://doi.org/10.1038/nature04269
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04269
This article is cited by
-
Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
Nature Communications (2024)
-
Novel whole-mount FISH analysis for intact root of Arabidopsis thaliana with spatial reference to 3D visualization
Journal of Plant Research (2023)
-
Dynamic 3D genome architecture of cotton fiber reveals subgenome-coordinated chromatin topology for 4-staged single-cell differentiation
Genome Biology (2022)
-
Positional cues and cell division dynamics drive meristem development and archegonium formation in Ceratopteris gametophytes
Communications Biology (2022)
-
Microspore embryogenesis in Brassica: calcium signaling, epigenetic modification, and programmed cell death
Planta (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.